diff --git a/Instructions/Install_environment_instructions_mac.pdf b/Instructions/Install_environment_instructions_mac.pdf
deleted file mode 100644
index d6fccb34..00000000
Binary files a/Instructions/Install_environment_instructions_mac.pdf and /dev/null differ
diff --git a/Instructions/Installation_instructions_M1_CellProfiler_CellPose.md b/Instructions/Installation_instructions_M1_CellProfiler_CellPose.md
deleted file mode 100644
index 3ed18f38..00000000
--- a/Instructions/Installation_instructions_M1_CellProfiler_CellPose.md
+++ /dev/null
@@ -1,181 +0,0 @@
-Installation instructions for CellProfiler, CellProfiler plugins and CellPose in a conda environment on Apple silicon (eg M1).
-
-1. **Install brew**
- ```
- /bin/bash -c "$(curl -fsSL https://raw.githubusercontent.com/Homebrew/install/HEAD/install.sh)"
- ```
-
-2. **Update brew**
- ```
- brew update
- ```
-
-3. **Install Java**
- ```
- brew install java
- ```
-
-4. **For the system Java wrappers to find this JDK and symlink it**
- ```
- sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
- ```
-
-5. **Set version in .zshrc**
- ```
- echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
- source ~/.zshrc
- ```
-
-6. **Brew install package requirements**
- ```
- brew install freetype mysql git
- ```
-
-7. **Direct to brew openssl**
- ```
- export LDFLAGS="-L$(brew --prefix)/opt/openssl/lib"
- ```
-
-8. **Install hdf5**
- ```
- brew install hdf5@1.12
- ```
-
-9. **Make sure to get the version directory correct for the version installed. Find with `ls /opt/homebrew/Cellar/hdf5/`**
- ```
- export HDF5_DIR=/opt/homebrew/Cellar/hdf5/1.12.1_1/
- ```
-10. **Create a folder and download cellprofiler-plugins**
-
- ```
- mkdir cp_plugins
- cd cp_plugins
- git clone https://github.com/CellProfiler/CellProfiler-plugins
- ```
-
-11. **Download and install miniconda**
-
- ```
- wget https://repo.anaconda.com/miniconda/Miniconda3-latest-MacOSX-arm64.sh -O ~/miniconda.sh
- ```
-
-
-12. **Create a conda `.yml` file**
- In a text editor, paste the following test and save it as `cellprofiler_plugins_macM1.yml`:
- ```
- name: cpcellpose
-
- channels:
- - conda-forge
- - anaconda
- - bioconda
- - defaults
- - apple
-
- dependencies:
- - python=3.8
- - pip
- - h5py
- - mysqlclient
- - imagecodecs
- - python.app
- - pandas
- - pip:
- - cellpose
- - attrdict
- - sip==5.5.0
- - boto3>=1.12.28
- - centrosome==1.2.1
- - docutils==0.15.2
- - h5py~=3.6.0
- - imageio>=2.5
- - inflect>=2.1
- - Jinja2>=2.11.2
- - joblib>=0.13
- - mahotas>=1.4
- - matplotlib==3.1.3
- - mysqlclient==1.4.6
- - numpy>=1.20.1
- - Pillow>=7.1.0
- - prokaryote==2.4.4
- - python-bioformats==4.0.6
- - python-javabridge==4.0.3
- - pyzmq~=22.3
- - sentry-sdk==0.18.0
- - requests>=2.22
- - scikit-image>=0.17.2
- - scikit-learn>=0.20
- - scipy>=1.4.1
- - six
- - tifffile<2022.4.22
- - wxPython==4.2.0
- ```
-
-
-13. **Create a conda environment using the cellprofiler_plugins_macM1.yml file**
-
- At this stage, your folder/file structure should look like this:
-
- ```
- ├── cp_plugins
- ├── CellProfiler-plugins
- └── cellprofiler_plugins_macM1.yml
- ```
-
- In the terminal, make sure you are in the `cp_plugins` folder mentioned above (run `cd cp_plugins` to get there).
-
- ```
- conda env create -n cp_plugins --file cellprofiler_plugins_macM1.yml
- ```
-
-14. **Activate the conda environment**
-
- ```
- conda activate cp_plugins
- ```
-
-15. **Install CellProfiler and CellProfiler-core**
- ```
- pip install cellprofiler-core
-
- # We install CellProfiler with no dependencies here since we installed them with the conda yml file
- # This allows us to have some dependencies for M1 that are not the default for CellProfiler (eg wxPython)
- git clone https://github.com/CellProfiler/CellProfiler
-
- pip install --no-deps cellprofiler
- ```
-
-16. **Open CellProfiler**
- ```
- pythonw -m cellprofiler
- ```
-
-17. **Connect CellProfiler with the plugins folder**
-
- 1. In CellProfiler, go to `File` then `Preferences...`
- 2. Scroll down and look for `CellProfiler Plugins Directory` on the left.
- 3. Select the Browse button and choose the folder where you extracted the CellProfiler plugins files. It is probably called `CellProfiler-plugins` unless you have renamed it.
- 4. Select Save at the bottom of the Preferences window
- 5. Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` in the command line
-
-
-### Resolving dependencies conflicts
-
- There can be some strange dependency conflicts that can arise with python-javabridge. In the terminal with your environment activate, enter:
- ```
- pip uninstall -y centrosome python-javabridge
- pip install --no-cache-dir --no-deps --no-build-isolation python-javabridge centrosome
- pip uninstall matplotlib -y
- pip install matplotlib==3.2
- pip uninstall mahotas -y
- pip install mahotas
- ```
-
- The latest version of CellProfiler available on github removes the python-javabridge requirement and instead uses ScyJava.
-
-### Test your installation
-
-Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
-In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
diff --git a/Instructions/Windows_batch_file/Install_CellProfiler_with_Plugins.zip b/Instructions/Windows_batch_file/Install_CellProfiler_with_Plugins.zip
deleted file mode 100644
index 68c9a5a3..00000000
Binary files a/Instructions/Windows_batch_file/Install_CellProfiler_with_Plugins.zip and /dev/null differ
diff --git a/Instructions/Windows_batch_file/README.md b/Instructions/Windows_batch_file/README.md
deleted file mode 100644
index 0387e35d..00000000
--- a/Instructions/Windows_batch_file/README.md
+++ /dev/null
@@ -1,62 +0,0 @@
-# How to use the batch file:
-* On a WINDOWS computer, download the zip folder called **Install_CellProfiler_with_plugins.zip**
-* Put the folder on your desktop
-* Extract the contents
-* Right click file called "install_cellprofiler_plugins.bat" and select "Run as Administrator"
- * You may get a security warning when running the batch file. Press More Info, then Run Anyway to proceed.
- * The script will run through a series of checks and steps. After it finishes each step, it will prompt you to press a key to continue. Before you press a key, please look at the text on the screen and see if there are any errors or warnings present. In general, the bat file will try to continue through most warnings and non-fatal errors and it's up to you to observe if one of these steps did not work. Also, if the script quits before reaching the end, the script has encountered an error. The steps are summarized below in "How the batch file works"
-* After installation finishes, you can open CellProfiler by double clicking the batch file run_cellprofiler.bat. If this does nothing, it also means that somewhere along the way, your install failed.
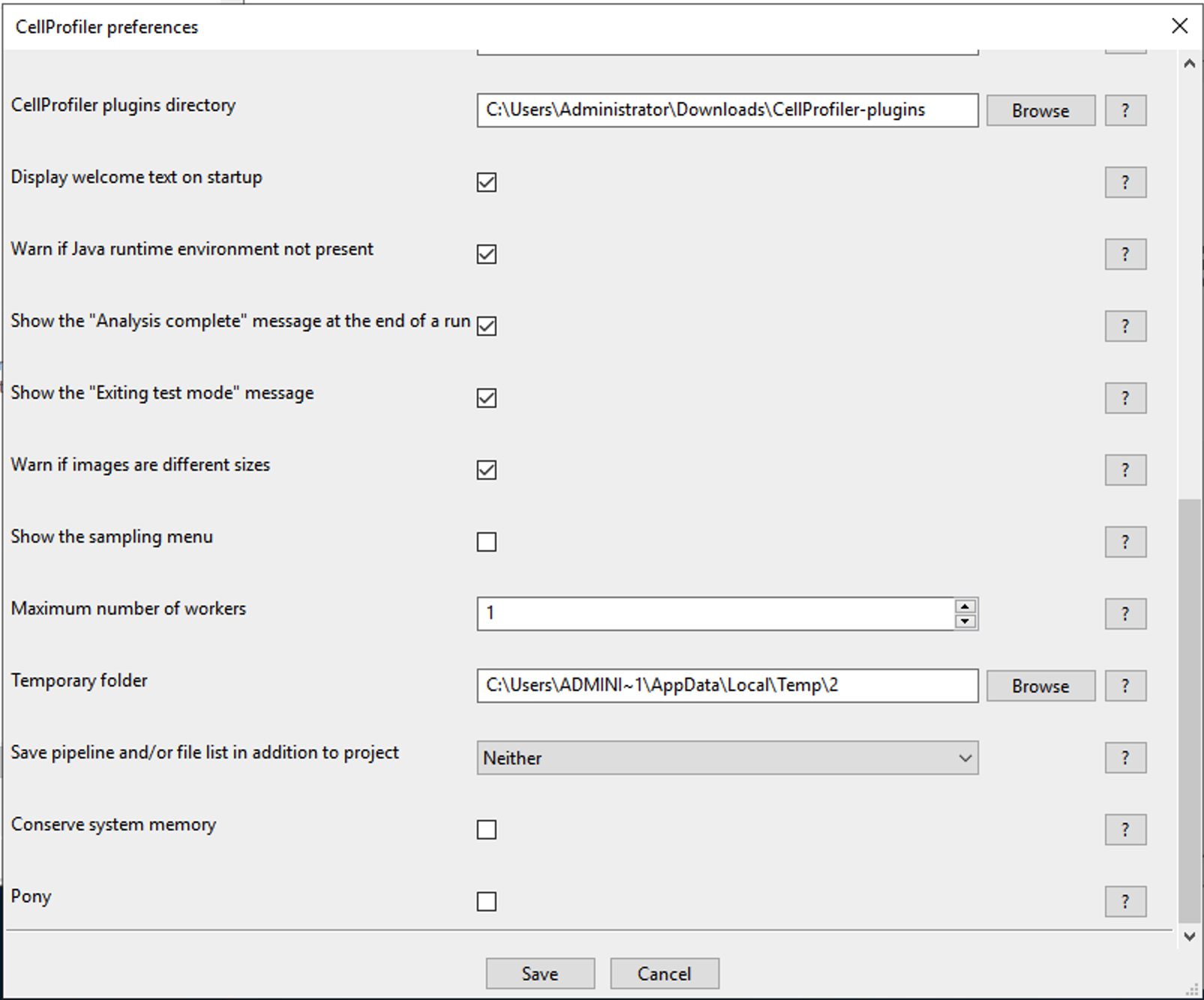
-* The only thing you should have to configure is in CellProfiler, go to **File** > **Preferences** and scroll down to the **CellProfiler plugins folder** option and set this to the location of your CellProflier-plugins folder (it should be in your Desktop or your Downloads folder). See below:
-
- -
-
-
-* Close and re-open CellProfiler and you should have plugins available. If you can add runCellpose or runStarDist to your pipeline, then the installation was a success!
-
-
-# How the batch file works:
-* The beginning checks if you're in administrator mode (necessary for chocolatey)
-* The next part uses powershell to download chocolatey and install it (if it isn't already installed)
-* Then packages are installed the same way as from the command line with `choco install package-name`
-* Next we check if the system has conda available and if not, add miniconda paths to the system-level PATH
-* Next we use powershell to download OpenJDK 11 from adoptium. This step takes some minutes (~5). After downloading, this .ps1 file will install java to C:/Program Files/Java-jdk-11 and then add JAVA_HOME and JDK_HOME system environment variables
-* Next we move into the Downloads folder and if a CellProfiler-plugins folder isn't there, it will download a copy of the repo to the Downloads folder
- * The folder is extracted and the original .zip file removed
-* Finally we use miniconda to create a new CP_plugins environment from the .yml file in the CellProfiler-plugins folder and perform necessary additional installs
-
-There is a separate batch file that just activates the conda environment and opens CellProfiler (so the user doesn't have to type anything into the command line)
-You can also always open CellProfiler by going to your start menu or list of programs and searching for "Anaconda Prompt (miniconda3)." Select this program to open a terminal and then type in `conda activate CP_plugins` then `pip install javabridge` and this will perform the installation. Then you can type in `cellprofiler` and it will open CellProfiler.
-
-
-# Potential errors
-When you try to run CellProfiler, you might see an error. Here are common errors and solutions:
-
-## Can't find the Java virtual machine
-```
-Traceback (most recent call last):
- File "C:\tools\miniconda3\envs\CP_plugins\lib\runpy.py", line 194, in _run_module_as_main
- return _run_code(code, main_globals, None,
- File "C:\tools\miniconda3\envs\CP_plugins\lib\runpy.py", line 87, in _run_code
- exec(code, run_globals)
- File "C:\tools\miniconda3\envs\CP_plugins\Scripts\cellprofiler.exe\__main__.py", line 4, in
- File "C:\tools\miniconda3\envs\CP_plugins\lib\site-packages\cellprofiler\__main__.py", line 13, in
- import bioformats.formatreader
- File "C:\tools\miniconda3\envs\CP_plugins\lib\site-packages\bioformats\__init__.py", line 21, in
- import javabridge
- File "C:\tools\miniconda3\envs\CP_plugins\lib\site-packages\javabridge\__init__.py", line 38, in
- from .jutil import start_vm, kill_vm, vm, activate_awt, deactivate_awt
- File "C:\tools\miniconda3\envs\CP_plugins\lib\site-packages\javabridge\jutil.py", line 150, in
- os.environ["PATH"] = os.environ["PATH"] + os.pathsep + _find_jvm() + \
- File "C:\tools\miniconda3\envs\CP_plugins\lib\site-packages\javabridge\jutil.py", line 139, in _find_jvm
- raise JVMNotFoundError()
-javabridge.jutil.JVMNotFoundError: Can't find the Java Virtual Machine
-```
-
-If you see this error, first verify that Java is installed.
-* There should be a folder in C:/Program Files called **Java-jdk-11**. If not, the installation has failed.
-* Also check your Environment Variables and ensure that JAVA_HOME and JDK_HOME are in the System Variables (second box) and both are set to the location of the Java-jdk-11 folder. If they are not set, you can click Edit and then Browse Directory and choose the Java-jdk-11 folder in Program Files.
- * Windows 10 or below, type "Environment variable" into the search bar.
- * Windows 11: Go to Control Panel > System > Advanced system settings > Environment Variables
-* You can always manually download java jdk 11 from [Adoptium's website](https://adoptium.net/temurin/releases/?version=11).
-
-* Assuming you have java installed and your environment variables set and you still get this error, the solution is to install javabridge, which appears to be necessary for some Windows computers. Go to your start menu or list of programs and search for "Anaconda Prompt (miniconda3)" and then type in `conda activate CP_plugins` then press Enter to activate the environment. Then enter `pip install javabridge` and this will perform the installation. Then you can enter `cellprofiler` and it will open CellProfiler.
diff --git a/Instructions/cellprofiler_plugins_mac.yml b/Instructions/cellprofiler_plugins_mac.yml
deleted file mode 100644
index 4153d52b..00000000
--- a/Instructions/cellprofiler_plugins_mac.yml
+++ /dev/null
@@ -1,26 +0,0 @@
-name: CellProfiler_plugins
-channels:
- - defaults
- - conda-forge
-dependencies:
- - cython=0.29.28
- - sqlite=3.38.3
- - openssl
- - certifi
- - pip=21.2.2
- - python=3.8.13
- - ca-certificates
- - wheel=0.37.1
- - setuptools=61.2.0
- - python.app
- - jpype1
- - python-javabridge
- - mysqlclient=1.4.6
- - pip:
- - cellpose==1.0.2
- - cellprofiler==4.2.1
- - imagej==0.3.1
- - pyjnius==1.4.1
- - tensorflow==2.9.1
-
-prefix: /Users/rsenft/opt/anaconda3/envs/CellProfiler_plugins
diff --git a/Instructions/cellprofiler_plugins_macM1.yml b/Instructions/cellprofiler_plugins_macM1.yml
deleted file mode 100644
index c3c164e5..00000000
--- a/Instructions/cellprofiler_plugins_macM1.yml
+++ /dev/null
@@ -1,46 +0,0 @@
-name: cpcellpose
-
-channels:
-- conda-forge
-- anaconda
-- bioconda
-- defaults
-- apple
-
-dependencies:
-- python=3.8
-- pip
-- h5py
-- mysqlclient
-- imagecodecs
-- python.app
-- pandas
-- pip:
- - cellpose
- - attrdict
- - sip==5.5.0
- - boto3>=1.12.28
- - centrosome==1.2.1
- - docutils==0.15.2
- - h5py~=3.6.0
- - imageio>=2.5
- - inflect>=2.1
- - Jinja2>=2.11.2
- - joblib>=0.13
- - mahotas>=1.4
- - matplotlib==3.1.3
- - mysqlclient==1.4.6
- - numpy>=1.20.1
- - Pillow>=7.1.0
- - prokaryote==2.4.4
- - python-bioformats==4.0.6
- - python-javabridge==4.0.3
- - pyzmq~=22.3
- - sentry-sdk==0.18.0
- - requests>=2.22
- - scikit-image>=0.17.2
- - scikit-learn>=0.20
- - scipy>=1.4.1
- - six
- - tifffile<2022.4.22
- - wxPython==4.2.0
diff --git a/Instructions/cellprofiler_plugins_windows.yml b/Instructions/cellprofiler_plugins_windows.yml
deleted file mode 100644
index 42f3ccee..00000000
--- a/Instructions/cellprofiler_plugins_windows.yml
+++ /dev/null
@@ -1,37 +0,0 @@
-name: CP_plugins
-channels:
- - defaults
-dependencies:
- - blas=1.0=mkl
- - ca-certificates=2022.4.26=haa95532_0
- - certifi=2022.6.15=py38haa95532_0
- - cython=0.29.28=py38hd77b12b_0
- - intel-openmp=2021.4.0=haa95532_3556
- - jpype1=1.2.1=py38h59b6b97_0
- - mkl=2021.4.0=haa95532_640
- - mkl-service=2.4.0=py38h2bbff1b_0
- - mkl_fft=1.3.1=py38h277e83a_0
- - mkl_random=1.2.2=py38hf11a4ad_0
- - numba=0.53.1=py38hf11a4ad_0
- - openssl=1.1.1p=h2bbff1b_0
- - pip=21.2.2=py38haa95532_0
- - python=3.8.13=h6244533_0
- - setuptools=61.2.0=py38haa95532_0
- - six=1.16.0=pyhd3eb1b0_1
- - sqlite=3.38.5=h2bbff1b_0
- - tbb=2020.3=h74a9793_0
- - vc=14.2=h21ff451_1
- - vs2015_runtime=14.27.29016=h5e58377_2
- - wheel=0.37.1=pyhd3eb1b0_0
- - wincertstore=0.2=py38haa95532_2
- - zlib=1.2.12=h8cc25b3_2
- - pip:
- - cellpose==1.0.2
- - cellprofiler==4.2.1
- - cellprofiler-core==4.2.1
- - imagej==0.3.1
- - numpy==1.23.0
- - pyjnius==1.4.1
- - tensorflow==2.9.1
-
-prefix: C:\tools\miniconda3\envs\CP_plugins
diff --git a/README.md b/README.md
index bbad5f1a..1d46292d 100644
--- a/README.md
+++ b/README.md
@@ -1,45 +1,14 @@
-CellProfiler-plugins
-====================
-[](https://travis-ci.org/CellProfiler/CellProfiler-plugins)
+# CellProfiler-plugins
-A home for community-contributed and experimental CellProfiler modules.
+A home for community-contributed, experimental, and dependency-heavy CellProfiler modules.
-## Beginner-level instructions
-[Beginner guidance for using plugins in CellProfiler](https://github.com/CellProfiler/CellProfiler/blob/master/cellprofiler/data/help/other_plugins.rst)
+Plugins advance the capabilities of CellProfiler but are not officially supported in the same way as modules.
+A module may be in CellProfiler-plugins instead of CellProfiler itself because:
+- it is under active development
+- it has a niche audience
+- it is not documented to CellProfiler's standards
+- it only works with certain version of CellProfiler
+- it requires extra libraries or other dependencies we are unable or unwilling to require for CellProfiler
+- it has been contributed by a community member
-Beginner instructions for installing runCellpose, runStarDist, and all other plugins with CellProfiler from source:
-* [Windows](https://github.com/CellProfiler/CellProfiler-plugins/blob/master/Instructions/Install_environment_instructions_windows.md)
-* [Mac_Intel_version](https://github.com/CellProfiler/CellProfiler-plugins/blob/master/Instructions/Install_environment_instructions_mac.pdf)
-
-
-## Use
-1. Clone this repository into a folder (PLUGIN_DIRECTORY) on your local computer:
- ```
- cd PLUGIN_DIRECTORY
- git clone https://github.com/CellProfiler/CellProfiler-plugins.git
- ```
-
- Alternatively download zip and manually extract to PLUGIN_DIRECTORY.
-
-1. Install required dependencies:
- ```
- cd CellProfiler-plugins
- pip install -r requirements.txt
- ```
-
- To install CellProfiler-plugins on a windows machine with support for the deep learning module ClassifyPixels-UNet make sure you have [Visual Studio](https://support.microsoft.com/help/2977003/the-latest-supported-visual-c-downloads) 2017 installed then use
- ```
- cd CellProfiler-plugins
- pip install -r requirements-windows.txt
- ```
-
-1. Configure CellProfiler plugins directory in the GUI via `Preferences > CellProfiler plugins directory` (you will need to restart CellProfiler for the change to take effect). When running CellProfiler via the command line, use the `--plugins-directory` flag to specify the plugins directory, for example:
- ```
- cellprofiler --run --run-headless --project PROJECT_FILE --plugins-directory PLUGIN_DIRECTORY/CellProfiler-plugins
- ```
-
-## ImageJ requirements
-
- If using the `RunImageJScript` module, please note:
- * You will also need to [install Maven](https://github.com/imagej/pyimagej/blob/master/doc/Install.md#installing-via-pip)
- * CellProfiler will need to be [built from source](https://github.com/CellProfiler/CellProfiler/blob/master/cellprofiler/data/help/other_plugins.rst) due to the requirement of additional libraries
+Please see our [CellProfiler-plugins documentation](https://cellprofiler.github.io/CellProfiler-plugins) for more information about installation, currently supported plugins, and how to contribute.
\ No newline at end of file
diff --git a/documentation/.github/workflows/deploy.yml b/documentation/.github/workflows/deploy.yml
new file mode 100644
index 00000000..9e12290f
--- /dev/null
+++ b/documentation/.github/workflows/deploy.yml
@@ -0,0 +1,40 @@
+name: deploy-documentation
+
+# Only run this when the master branch changes
+on:
+ push:
+ branches:
+ - master
+ # Only run if edits in DS-documentation or
+ paths:
+ - documentation/CP-plugins-documentation/**
+ - .github/workflows/deploy.yml
+
+# This job installs dependencies, builds the book, and pushes it to `gh-pages`
+jobs:
+ deploy-book:
+ runs-on: ubuntu-latest
+ steps:
+ - uses: actions/checkout@v2
+
+ # Install dependencies
+ - name: Set up Python 3.8
+ uses: actions/setup-python@v2
+ with:
+ python-version: 3.8
+
+ - name: Install dependencies
+ run: |
+ pip install jupyter-book
+
+ # Build the book
+ - name: Build the book
+ run: |
+ jupyter-book build CP-plugins-documentation/
+
+ # Push the book's HTML to github-pages
+ - name: GitHub Pages action
+ uses: peaceiris/actions-gh-pages@v3.6.1
+ with:
+ github_token: ${{ secrets.GITHUB_TOKEN }}
+ publish_dir: ./CP-plugins-documentation/_build/html
diff --git a/Instructions/Install_environment_instructions_mac.md b/documentation/CP-plugins-documentation/Mac_installation.md
similarity index 59%
rename from Instructions/Install_environment_instructions_mac.md
rename to documentation/CP-plugins-documentation/Mac_installation.md
index c4f1636b..032edd3c 100644
--- a/Instructions/Install_environment_instructions_mac.md
+++ b/documentation/CP-plugins-documentation/Mac_installation.md
@@ -160,4 +160,184 @@ Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
To fix this, `conda install cython`
----
+## Installation instructions for CellProfiler, CellProfiler plugins and CellPose in a conda environment on Apple silicon (eg M1).
+
+1. **Install brew**
+ ```
+ /bin/bash -c "$(curl -fsSL https://raw.githubusercontent.com/Homebrew/install/HEAD/install.sh)"
+ ```
+
+2. **Update brew**
+ ```
+ brew update
+ ```
+
+3. **Install Java**
+ ```
+ brew install java
+ ```
+
+4. **For the system Java wrappers to find this JDK and symlink it**
+ ```
+ sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
+ ```
+
+5. **Set version in .zshrc**
+ ```
+ echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
+ source ~/.zshrc
+ ```
+
+6. **Brew install package requirements**
+ ```
+ brew install freetype mysql git
+ ```
+
+7. **Direct to brew openssl**
+ ```
+ export LDFLAGS="-L$(brew --prefix)/opt/openssl/lib"
+ ```
+
+8. **Install hdf5**
+ ```
+ brew install hdf5@1.12
+ ```
+
+9. **Make sure to get the version directory correct for the version installed. Find with `ls /opt/homebrew/Cellar/hdf5/`**
+ ```
+ export HDF5_DIR=/opt/homebrew/Cellar/hdf5/1.12.1_1/
+ ```
+10. **Create a folder and download cellprofiler-plugins**
+
+ ```
+ mkdir cp_plugins
+ cd cp_plugins
+ git clone https://github.com/CellProfiler/CellProfiler-plugins
+ ```
+
+11. **Download and install miniconda**
+
+ ```
+ wget https://repo.anaconda.com/miniconda/Miniconda3-latest-MacOSX-arm64.sh -O ~/miniconda.sh
+ ```
+
+
+12. **Create a conda `.yml` file**
+ In a text editor, paste the following test and save it as `cellprofiler_plugins_macM1.yml`:
+ ```
+ name: cpcellpose
+
+ channels:
+ - conda-forge
+ - anaconda
+ - bioconda
+ - defaults
+ - apple
+
+ dependencies:
+ - python=3.8
+ - pip
+ - h5py
+ - mysqlclient
+ - imagecodecs
+ - python.app
+ - pandas
+ - pip:
+ - cellpose
+ - attrdict
+ - sip==5.5.0
+ - boto3>=1.12.28
+ - centrosome==1.2.1
+ - docutils==0.15.2
+ - h5py~=3.6.0
+ - imageio>=2.5
+ - inflect>=2.1
+ - Jinja2>=2.11.2
+ - joblib>=0.13
+ - mahotas>=1.4
+ - matplotlib==3.1.3
+ - mysqlclient==1.4.6
+ - numpy>=1.20.1
+ - Pillow>=7.1.0
+ - prokaryote==2.4.4
+ - python-bioformats==4.0.6
+ - python-javabridge==4.0.3
+ - pyzmq~=22.3
+ - sentry-sdk==0.18.0
+ - requests>=2.22
+ - scikit-image>=0.17.2
+ - scikit-learn>=0.20
+ - scipy>=1.4.1
+ - six
+ - tifffile<2022.4.22
+ - wxPython==4.2.0
+ ```
+
+
+13. **Create a conda environment using the cellprofiler_plugins_macM1.yml file**
+
+ At this stage, your folder/file structure should look like this:
+
+ ```
+ ├── cp_plugins
+ ├── CellProfiler-plugins
+ └── cellprofiler_plugins_macM1.yml
+ ```
+
+ In the terminal, make sure you are in the `cp_plugins` folder mentioned above (run `cd cp_plugins` to get there).
+
+ ```
+ conda env create -n cp_plugins --file cellprofiler_plugins_macM1.yml
+ ```
+
+14. **Activate the conda environment**
+
+ ```
+ conda activate cp_plugins
+ ```
+
+15. **Install CellProfiler and CellProfiler-core**
+ ```
+ pip install cellprofiler-core
+
+ # We install CellProfiler with no dependencies here since we installed them with the conda yml file
+ # This allows us to have some dependencies for M1 that are not the default for CellProfiler (eg wxPython)
+ git clone https://github.com/CellProfiler/CellProfiler
+
+ pip install --no-deps cellprofiler
+ ```
+
+16. **Open CellProfiler**
+ ```
+ pythonw -m cellprofiler
+ ```
+
+17. **Connect CellProfiler with the plugins folder**
+
+ 1. In CellProfiler, go to `File` then `Preferences...`
+ 2. Scroll down and look for `CellProfiler Plugins Directory` on the left.
+ 3. Select the Browse button and choose the folder where you extracted the CellProfiler plugins files. It is probably called `CellProfiler-plugins` unless you have renamed it.
+ 4. Select Save at the bottom of the Preferences window
+ 5. Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` in the command line
+
+
+### Resolving dependencies conflicts
+
+ There can be some strange dependency conflicts that can arise with python-javabridge. In the terminal with your environment activate, enter:
+ ```
+ pip uninstall -y centrosome python-javabridge
+ pip install --no-cache-dir --no-deps --no-build-isolation python-javabridge centrosome
+ pip uninstall matplotlib -y
+ pip install matplotlib==3.2
+ pip uninstall mahotas -y
+ pip install mahotas
+ ```
+
+ The latest version of CellProfiler available on github removes the python-javabridge requirement and instead uses ScyJava.
+
+### Test your installation
+
+Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
+
+In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
+
diff --git a/Instructions/Install_environment_instructions_windows.md b/documentation/CP-plugins-documentation/Windows_installation.md
similarity index 100%
rename from Instructions/Install_environment_instructions_windows.md
rename to documentation/CP-plugins-documentation/Windows_installation.md
diff --git a/documentation/CP-plugins-documentation/_config.yml b/documentation/CP-plugins-documentation/_config.yml
new file mode 100644
index 00000000..702fc3aa
--- /dev/null
+++ b/documentation/CP-plugins-documentation/_config.yml
@@ -0,0 +1,34 @@
+# Book settings
+
+# Learn more at https://jupyterbook.org/customize/config.html
+title: CP Plugins Documentation
+author: Broad Institute
+copyright: "2023"
+#logo: img/logo.svg
+
+# Only build files that are in the ToC
+only_build_toc_files: true
+
+# Force re-execution of notebooks on each build.
+# See https://jupyterbook.org/content/execute.html
+execute:
+ execute_notebooks: force
+
+# Information about where the book exists on the web
+repository:
+ url: https://github.com/cellprofiler/cellprofiler-plugins
+ branch: main # Which branch of the repository should be used when creating links (optional)
+ path_to_book: CP-plugins-documentation
+
+html:
+ baseurl: cellprofiler.github.io
+ use_repository_button: true
+ use_issues_button: true
+ use_edit_page_button: true
+ comments:
+ hypothesis: true
+
+parse:
+ myst_enable_extensions:
+ # Only required if you use html ![]() + - html_image
diff --git a/documentation/CP-plugins-documentation/_toc.yml b/documentation/CP-plugins-documentation/_toc.yml
new file mode 100644
index 00000000..311a0a8c
--- /dev/null
+++ b/documentation/CP-plugins-documentation/_toc.yml
@@ -0,0 +1,14 @@
+# Table of contents
+
+format: jb-book
+root: overview
+parts:
+- caption: Overview
+ chapters:
+ - file: overview
+ - file: supported_plugins
+ - file: unsupported_plugins
+- caption: Technical Guides
+ chapters:
+ - file: Windows
+ - file: Mac
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/citing.md b/documentation/CP-plugins-documentation/citing.md
new file mode 100644
index 00000000..fd38d6c6
--- /dev/null
+++ b/documentation/CP-plugins-documentation/citing.md
@@ -0,0 +1,15 @@
+# Citing CellProfiler Plugins
+
+If using CellProfiler Plugins in a pipeline used in a publication, please cite our CellProfiler Plugins paper:
+
+ENTER CITATION
+
+Additionally, some plugins require citation themselves if used.
+You can determine which plugins require citation by reading the Help information for each plugin, accessible through CellProfiler, or by using the Citation generator built into CellProfiler (since VERSION).
+
+To use CellProfiler's citation generator, load your pipeline into CellProfiler.
+Navigate to File => Export => Citation.
+
+Please note that many CellProfiler modules (including plugins) provide references that offer further information/background about the processing happening within the module.
+You may wish to read the references and cite any that upon which your analyses are particularly dependent.
+However, please do note that `References` and `Citations` are separate and citing module references is not required.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
new file mode 100644
index 00000000..204ec8f8
--- /dev/null
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -0,0 +1,17 @@
+## Contributing New Plugins
+
+Refer to CellProfiler repository and within cellprofiler/modules/plugins you can find two different templates to use for creating your own plugin.
+`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
+`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
+
+In you plugin, you must include:
+
+In your plugin, we appreciate if you also include:
+- display functionality
+
+
+In your PR, you must include:
+- add your plugin to the [supported_plugins](supported_plugins.md) documentation page
+
+In your PR, we appreciate if you also include:
+- unit tests for your plugin
\ No newline at end of file
diff --git a/Instructions/images/Install_environment_instructions/2022-05-26T15-39-07.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions/2022-05-26T15-39-07.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions/2022-05-26T15-39-07.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions/2022-05-26T15-39-07.png
diff --git a/Instructions/images/Install_environment_instructions/file-import-solid.svg b/documentation/CP-plugins-documentation/images/Install_environment_instructions/file-import-solid.svg
similarity index 100%
rename from Instructions/images/Install_environment_instructions/file-import-solid.svg
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions/file-import-solid.svg
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T20-56-38.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T20-56-38.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T20-56-38.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T20-56-38.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-00-53.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-00-53.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-00-53.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-00-53.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-05-30.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-05-30.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-05-30.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-05-30.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-11-49.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-11-49.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-11-49.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-11-49.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-16-31.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-16-31.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-16-31.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-16-31.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-17-09.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-17-09.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-17-09.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-17-09.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-30-10.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-30-10.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-30-10.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-30-10.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-39-05.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-39-05.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-39-05.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-39-05.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-43-56.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-43-56.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-43-56.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-43-56.png
diff --git a/Instructions/images/Install_environment_instructions_windows/Control_panel.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Control_panel.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/Control_panel.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Control_panel.png
diff --git a/Instructions/images/Install_environment_instructions_windows/Control_panel_ev.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Control_panel_ev.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/Control_panel_ev.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Control_panel_ev.png
diff --git a/Instructions/images/Install_environment_instructions_windows/Java-download.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Java-download.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/Java-download.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Java-download.png
diff --git a/Instructions/images/Install_environment_instructions_windows/build-tools-download.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/build-tools-download.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/build-tools-download.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/build-tools-download.png
diff --git a/Instructions/images/Install_environment_instructions_windows/windows-file-explorer.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/windows-file-explorer.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/windows-file-explorer.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/windows-file-explorer.png
diff --git a/documentation/CP-plugins-documentation/overview.md b/documentation/CP-plugins-documentation/overview.md
new file mode 100644
index 00000000..5ece4c8d
--- /dev/null
+++ b/documentation/CP-plugins-documentation/overview.md
@@ -0,0 +1,15 @@
+# What are CellProfiler Plugins?
+
+Plugins advance the capabilities of CellProfiler but are not officially supported in the same way as modules.
+A module may be in CellProfiler-plugins instead of CellProfiler itself because:
+- it is under active development
+- it has a niche audience
+- it is not documented to CellProfiler's standards
+- it only works with certain version of CellProfiler
+- it requires extra libraries or other dependencies we are unable or unwilling to require for CellProfiler
+- it has been contributed by a community member
+
+
+## Who made this?
+
+CellProfiler and CellProfiler-plugins are maintained and developed in the [Cimini Lab](https://cimini-lab.broadinstitute.org) in the Imaging Platform at the Broad Institute in Cambridge, MA, USA.
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
new file mode 100644
index 00000000..5850822a
--- /dev/null
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -0,0 +1,63 @@
+# Supported Plugins
+
+Below is a brief overview of our currently supported plugins.
+For details about using any particular plugin, please read the module documentation inside the plugin in CellProfiler.
+
+## CalculateMoments
+CalculateMoments extracts moments statistics from a given distribution of pixel values.
+
+## CallBarcodes
+CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles.
+It is used for optical sequencing by synthesis (SBS).
+
+## CompensateColors
+CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through.
+It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched.
+
+## DeclumpObjects
+DeclumpObjects will split objects based on a seeded watershed method.
+
+## DistanceTransform
+DistanceTransform computes the distance transform of a binary image.
+The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1.
+
+## EnhancedMeasureTexture
+EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler.
+
+## HistogramEqualization
+HistogramEqualization increases the global contrast of a low-contrast image or volume.
+Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct.
+This module can perform either global or local histogram equalization.
+
+## HistogramMatching
+HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image.
+It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image.
+It allows you to choose which frame to use as the reference.
+
+## PixelShuffle
+PixelShuffle takes the intensity of each pixel in an image and it randomly shuffles its position.
+
+## Predict
+Predict allows you to use an ilastik pixel classifier to generate a probability image.
+CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage).
+
+## RunCellpose
+RunCellpose allows you to run Cellpose within CellProfiler.
+Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells.
+You can use pre-trained Cellpose models or your custom model with this plugin.
+You can use a GPU with this module to dramatically increase your speed/efficiency.
+
+## RunImageJScript
+
+## RunOmnipose
+
+## RunStarDist
+RunStarDist allows you to run StarDist within CellProfiler.
+StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells.
+You can use pre-trained StarDist models or your custom model with this plugin.
+You can use a GPU with this module to dramatically increase your speed/efficiency.
+RunStarDist is generally faster than RunCellpose.
+
+## VarianceTransform
+This module allows you to calculate the variance of an image, using a determined window size.
+It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
new file mode 100644
index 00000000..7e0343da
--- /dev/null
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -0,0 +1,12 @@
+# Currently Unsupported Plugins
+
+Unsupported plugins are primarily unsupported for either of two reasons:
+- They were made for a previous major version of CellProfiler and have not been updated to be compatible with the current version
+- Their functions have been integrated into the current version of CellProfiler and therefore a plugin is no longer necessary
+
+We welcome requests for updating particular unsupported plugins, but please note that we have limited bandwidth for working on plugins and may be unable to complete the update.
+We welcome community contributed plugin updates.
+
+Likewise, we welcome contributions of community developed plugins.
+Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team or community contributed.
+
diff --git a/Instructions/GPU_tips.md b/documentation/CP-plugins-documentation/using_GPU.md
similarity index 88%
rename from Instructions/GPU_tips.md
rename to documentation/CP-plugins-documentation/using_GPU.md
index 686c1ed7..945d869b 100644
--- a/Instructions/GPU_tips.md
+++ b/documentation/CP-plugins-documentation/using_GPU.md
@@ -1,8 +1,8 @@
-# Tips on setting up your GPU
+# Using a GPU with CellProfiler Plugins
-Updated June 28, 2022
+These CellProfiler modules currently support using a GPU: LIST MODULES
-## Setup your GPU (if you have one)
+## Tips for Setting Up Your GPU (if you have one)
If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU. General instructions are available at this [link](https://pytorch.org/get-started/locally/).
diff --git a/documentation/CP-plugins-documentation/versions.md b/documentation/CP-plugins-documentation/versions.md
new file mode 100644
index 00000000..3755e75b
--- /dev/null
+++ b/documentation/CP-plugins-documentation/versions.md
@@ -0,0 +1,55 @@
+# Versions
+
+The most current release can always be found on DockerHub at `cellprofiler/distributed-cellprofiler`.
+Current version is 2.0.0.
+Our current tag system is dcpversion_CellProfilerversion, e.g. 1.2.0_3.1.8
+
+Previous release versions can be accessed at `bethcimini/distributed-cellprofiler:versionnumber`
+
+---
+
+# Version History
+
+## 2.1.0 (forthcoming)
+* Increase support for using plugins
+* Improve role handling and allow file download/upload from/to S3 buckets in different accounts
+* Improve file download to S3FS can be completely bypassed
+* Name EBS volumes
+
+## 2.0.0rc2 - Released 20201110
+* Add optional ability to download files to EBS rather than reading from S3 (helpful for pipelines that access many files/image sets)
+
+## 2.0.0rc1 - Released 20201105
+
+* Remove requirement for boto and fabric, using only boto3
+* Add support for Python 3 and CellProfiler 4
+* Move cluster size, machine type, and machine price to the config file from the fleet file, eliminating mismatches between the two
+* Add the ability to filter only files with certain names when running CHECK_IF_DONE
+* Don't cancel a fleet over capacity errors
+* Add "cheapest" mode to the monitor, allowing you to run more cheaply (at possible expense of running more slowly)
+
+## 1.2.2 - Released 20201103
+
+* Allow pipelines using batch files to also designate an input output_top_directory
+* Add support for multiple LaunchData specifications
+* Add CellProfiler-plugins
+* Additional way to create job submissions with run_batch_general.py
+
+## 1.2.1 - Released 20200109, Updated through 20191002
+
+* Allow monitor to downscale machines when number of jobs < number of machines
+* Add a parameter to discount files when running CHECK_IF_DONE checks if less than a certain size
+
+## 1.2.0 - Released 20181108, Updated through 20200109
+
+* Improved compatibility with CellProfiler 2 and 3
+* Better handling of logging when using output_structure
+
+## 1.1.0 - Released 20170217, Updated 20170221 (bugfixes) - 20181018
+
+* Changes in this release:
+
+ * Added the `output_structure` variable to the job file, which allows you to structure the output folders created by DCP (ie `Plate/Well-Site` rather than `Plate-Well-Site`). Job files lacking this variable will still default to the previous settings (hyphenating all Metadata items in order they are presented in the Metadata grouping).
+ * Added support for creating the list of groups via `cellprofiler --print-groups`- see [this issue](https://github.com/CellProfiler/Distributed-CellProfiler/issues/52) for example and discussion. Groups listed in this way MUST use the `output_structure` variable to state their desired folder structure or an error will be returned.
+
+## 1.0.0 - Version as of 20170213
+ - html_image
diff --git a/documentation/CP-plugins-documentation/_toc.yml b/documentation/CP-plugins-documentation/_toc.yml
new file mode 100644
index 00000000..311a0a8c
--- /dev/null
+++ b/documentation/CP-plugins-documentation/_toc.yml
@@ -0,0 +1,14 @@
+# Table of contents
+
+format: jb-book
+root: overview
+parts:
+- caption: Overview
+ chapters:
+ - file: overview
+ - file: supported_plugins
+ - file: unsupported_plugins
+- caption: Technical Guides
+ chapters:
+ - file: Windows
+ - file: Mac
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/citing.md b/documentation/CP-plugins-documentation/citing.md
new file mode 100644
index 00000000..fd38d6c6
--- /dev/null
+++ b/documentation/CP-plugins-documentation/citing.md
@@ -0,0 +1,15 @@
+# Citing CellProfiler Plugins
+
+If using CellProfiler Plugins in a pipeline used in a publication, please cite our CellProfiler Plugins paper:
+
+ENTER CITATION
+
+Additionally, some plugins require citation themselves if used.
+You can determine which plugins require citation by reading the Help information for each plugin, accessible through CellProfiler, or by using the Citation generator built into CellProfiler (since VERSION).
+
+To use CellProfiler's citation generator, load your pipeline into CellProfiler.
+Navigate to File => Export => Citation.
+
+Please note that many CellProfiler modules (including plugins) provide references that offer further information/background about the processing happening within the module.
+You may wish to read the references and cite any that upon which your analyses are particularly dependent.
+However, please do note that `References` and `Citations` are separate and citing module references is not required.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
new file mode 100644
index 00000000..204ec8f8
--- /dev/null
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -0,0 +1,17 @@
+## Contributing New Plugins
+
+Refer to CellProfiler repository and within cellprofiler/modules/plugins you can find two different templates to use for creating your own plugin.
+`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
+`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
+
+In you plugin, you must include:
+
+In your plugin, we appreciate if you also include:
+- display functionality
+
+
+In your PR, you must include:
+- add your plugin to the [supported_plugins](supported_plugins.md) documentation page
+
+In your PR, we appreciate if you also include:
+- unit tests for your plugin
\ No newline at end of file
diff --git a/Instructions/images/Install_environment_instructions/2022-05-26T15-39-07.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions/2022-05-26T15-39-07.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions/2022-05-26T15-39-07.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions/2022-05-26T15-39-07.png
diff --git a/Instructions/images/Install_environment_instructions/file-import-solid.svg b/documentation/CP-plugins-documentation/images/Install_environment_instructions/file-import-solid.svg
similarity index 100%
rename from Instructions/images/Install_environment_instructions/file-import-solid.svg
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions/file-import-solid.svg
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T20-56-38.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T20-56-38.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T20-56-38.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T20-56-38.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-00-53.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-00-53.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-00-53.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-00-53.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-05-30.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-05-30.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-05-30.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-05-30.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-11-49.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-11-49.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-11-49.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-11-49.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-16-31.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-16-31.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-16-31.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-16-31.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-17-09.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-17-09.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-17-09.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-17-09.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-30-10.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-30-10.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-30-10.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-30-10.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-39-05.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-39-05.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-39-05.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-39-05.png
diff --git a/Instructions/images/Install_environment_instructions_windows/2022-06-02T21-43-56.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-43-56.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/2022-06-02T21-43-56.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/2022-06-02T21-43-56.png
diff --git a/Instructions/images/Install_environment_instructions_windows/Control_panel.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Control_panel.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/Control_panel.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Control_panel.png
diff --git a/Instructions/images/Install_environment_instructions_windows/Control_panel_ev.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Control_panel_ev.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/Control_panel_ev.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Control_panel_ev.png
diff --git a/Instructions/images/Install_environment_instructions_windows/Java-download.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Java-download.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/Java-download.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/Java-download.png
diff --git a/Instructions/images/Install_environment_instructions_windows/build-tools-download.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/build-tools-download.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/build-tools-download.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/build-tools-download.png
diff --git a/Instructions/images/Install_environment_instructions_windows/windows-file-explorer.png b/documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/windows-file-explorer.png
similarity index 100%
rename from Instructions/images/Install_environment_instructions_windows/windows-file-explorer.png
rename to documentation/CP-plugins-documentation/images/Install_environment_instructions_windows/windows-file-explorer.png
diff --git a/documentation/CP-plugins-documentation/overview.md b/documentation/CP-plugins-documentation/overview.md
new file mode 100644
index 00000000..5ece4c8d
--- /dev/null
+++ b/documentation/CP-plugins-documentation/overview.md
@@ -0,0 +1,15 @@
+# What are CellProfiler Plugins?
+
+Plugins advance the capabilities of CellProfiler but are not officially supported in the same way as modules.
+A module may be in CellProfiler-plugins instead of CellProfiler itself because:
+- it is under active development
+- it has a niche audience
+- it is not documented to CellProfiler's standards
+- it only works with certain version of CellProfiler
+- it requires extra libraries or other dependencies we are unable or unwilling to require for CellProfiler
+- it has been contributed by a community member
+
+
+## Who made this?
+
+CellProfiler and CellProfiler-plugins are maintained and developed in the [Cimini Lab](https://cimini-lab.broadinstitute.org) in the Imaging Platform at the Broad Institute in Cambridge, MA, USA.
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
new file mode 100644
index 00000000..5850822a
--- /dev/null
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -0,0 +1,63 @@
+# Supported Plugins
+
+Below is a brief overview of our currently supported plugins.
+For details about using any particular plugin, please read the module documentation inside the plugin in CellProfiler.
+
+## CalculateMoments
+CalculateMoments extracts moments statistics from a given distribution of pixel values.
+
+## CallBarcodes
+CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles.
+It is used for optical sequencing by synthesis (SBS).
+
+## CompensateColors
+CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through.
+It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched.
+
+## DeclumpObjects
+DeclumpObjects will split objects based on a seeded watershed method.
+
+## DistanceTransform
+DistanceTransform computes the distance transform of a binary image.
+The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1.
+
+## EnhancedMeasureTexture
+EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler.
+
+## HistogramEqualization
+HistogramEqualization increases the global contrast of a low-contrast image or volume.
+Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct.
+This module can perform either global or local histogram equalization.
+
+## HistogramMatching
+HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image.
+It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image.
+It allows you to choose which frame to use as the reference.
+
+## PixelShuffle
+PixelShuffle takes the intensity of each pixel in an image and it randomly shuffles its position.
+
+## Predict
+Predict allows you to use an ilastik pixel classifier to generate a probability image.
+CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage).
+
+## RunCellpose
+RunCellpose allows you to run Cellpose within CellProfiler.
+Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells.
+You can use pre-trained Cellpose models or your custom model with this plugin.
+You can use a GPU with this module to dramatically increase your speed/efficiency.
+
+## RunImageJScript
+
+## RunOmnipose
+
+## RunStarDist
+RunStarDist allows you to run StarDist within CellProfiler.
+StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells.
+You can use pre-trained StarDist models or your custom model with this plugin.
+You can use a GPU with this module to dramatically increase your speed/efficiency.
+RunStarDist is generally faster than RunCellpose.
+
+## VarianceTransform
+This module allows you to calculate the variance of an image, using a determined window size.
+It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
new file mode 100644
index 00000000..7e0343da
--- /dev/null
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -0,0 +1,12 @@
+# Currently Unsupported Plugins
+
+Unsupported plugins are primarily unsupported for either of two reasons:
+- They were made for a previous major version of CellProfiler and have not been updated to be compatible with the current version
+- Their functions have been integrated into the current version of CellProfiler and therefore a plugin is no longer necessary
+
+We welcome requests for updating particular unsupported plugins, but please note that we have limited bandwidth for working on plugins and may be unable to complete the update.
+We welcome community contributed plugin updates.
+
+Likewise, we welcome contributions of community developed plugins.
+Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team or community contributed.
+
diff --git a/Instructions/GPU_tips.md b/documentation/CP-plugins-documentation/using_GPU.md
similarity index 88%
rename from Instructions/GPU_tips.md
rename to documentation/CP-plugins-documentation/using_GPU.md
index 686c1ed7..945d869b 100644
--- a/Instructions/GPU_tips.md
+++ b/documentation/CP-plugins-documentation/using_GPU.md
@@ -1,8 +1,8 @@
-# Tips on setting up your GPU
+# Using a GPU with CellProfiler Plugins
-Updated June 28, 2022
+These CellProfiler modules currently support using a GPU: LIST MODULES
-## Setup your GPU (if you have one)
+## Tips for Setting Up Your GPU (if you have one)
If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU. General instructions are available at this [link](https://pytorch.org/get-started/locally/).
diff --git a/documentation/CP-plugins-documentation/versions.md b/documentation/CP-plugins-documentation/versions.md
new file mode 100644
index 00000000..3755e75b
--- /dev/null
+++ b/documentation/CP-plugins-documentation/versions.md
@@ -0,0 +1,55 @@
+# Versions
+
+The most current release can always be found on DockerHub at `cellprofiler/distributed-cellprofiler`.
+Current version is 2.0.0.
+Our current tag system is dcpversion_CellProfilerversion, e.g. 1.2.0_3.1.8
+
+Previous release versions can be accessed at `bethcimini/distributed-cellprofiler:versionnumber`
+
+---
+
+# Version History
+
+## 2.1.0 (forthcoming)
+* Increase support for using plugins
+* Improve role handling and allow file download/upload from/to S3 buckets in different accounts
+* Improve file download to S3FS can be completely bypassed
+* Name EBS volumes
+
+## 2.0.0rc2 - Released 20201110
+* Add optional ability to download files to EBS rather than reading from S3 (helpful for pipelines that access many files/image sets)
+
+## 2.0.0rc1 - Released 20201105
+
+* Remove requirement for boto and fabric, using only boto3
+* Add support for Python 3 and CellProfiler 4
+* Move cluster size, machine type, and machine price to the config file from the fleet file, eliminating mismatches between the two
+* Add the ability to filter only files with certain names when running CHECK_IF_DONE
+* Don't cancel a fleet over capacity errors
+* Add "cheapest" mode to the monitor, allowing you to run more cheaply (at possible expense of running more slowly)
+
+## 1.2.2 - Released 20201103
+
+* Allow pipelines using batch files to also designate an input output_top_directory
+* Add support for multiple LaunchData specifications
+* Add CellProfiler-plugins
+* Additional way to create job submissions with run_batch_general.py
+
+## 1.2.1 - Released 20200109, Updated through 20191002
+
+* Allow monitor to downscale machines when number of jobs < number of machines
+* Add a parameter to discount files when running CHECK_IF_DONE checks if less than a certain size
+
+## 1.2.0 - Released 20181108, Updated through 20200109
+
+* Improved compatibility with CellProfiler 2 and 3
+* Better handling of logging when using output_structure
+
+## 1.1.0 - Released 20170217, Updated 20170221 (bugfixes) - 20181018
+
+* Changes in this release:
+
+ * Added the `output_structure` variable to the job file, which allows you to structure the output folders created by DCP (ie `Plate/Well-Site` rather than `Plate-Well-Site`). Job files lacking this variable will still default to the previous settings (hyphenating all Metadata items in order they are presented in the Metadata grouping).
+ * Added support for creating the list of groups via `cellprofiler --print-groups`- see [this issue](https://github.com/CellProfiler/Distributed-CellProfiler/issues/52) for example and discussion. Groups listed in this way MUST use the `output_structure` variable to state their desired folder structure or an error will be returned.
+
+## 1.0.0 - Version as of 20170213
 -
- -
-