From 34976b13f1426969ca346a850a9ac4dc2e2866a0 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 10:17:12 -0700
Subject: [PATCH 1/6] overhaul install docs
---
.../Mac_installation.md | 343 ------------------
.../Windows_installation.md | 208 -----------
.../CP-plugins-documentation/using_plugins.md | 64 ++++
3 files changed, 64 insertions(+), 551 deletions(-)
delete mode 100644 documentation/CP-plugins-documentation/Mac_installation.md
delete mode 100644 documentation/CP-plugins-documentation/Windows_installation.md
create mode 100644 documentation/CP-plugins-documentation/using_plugins.md
diff --git a/documentation/CP-plugins-documentation/Mac_installation.md b/documentation/CP-plugins-documentation/Mac_installation.md
deleted file mode 100644
index 032edd3c..00000000
--- a/documentation/CP-plugins-documentation/Mac_installation.md
+++ /dev/null
@@ -1,343 +0,0 @@
-# How to install CellProfiler from source with all plugins on Mac OS
-
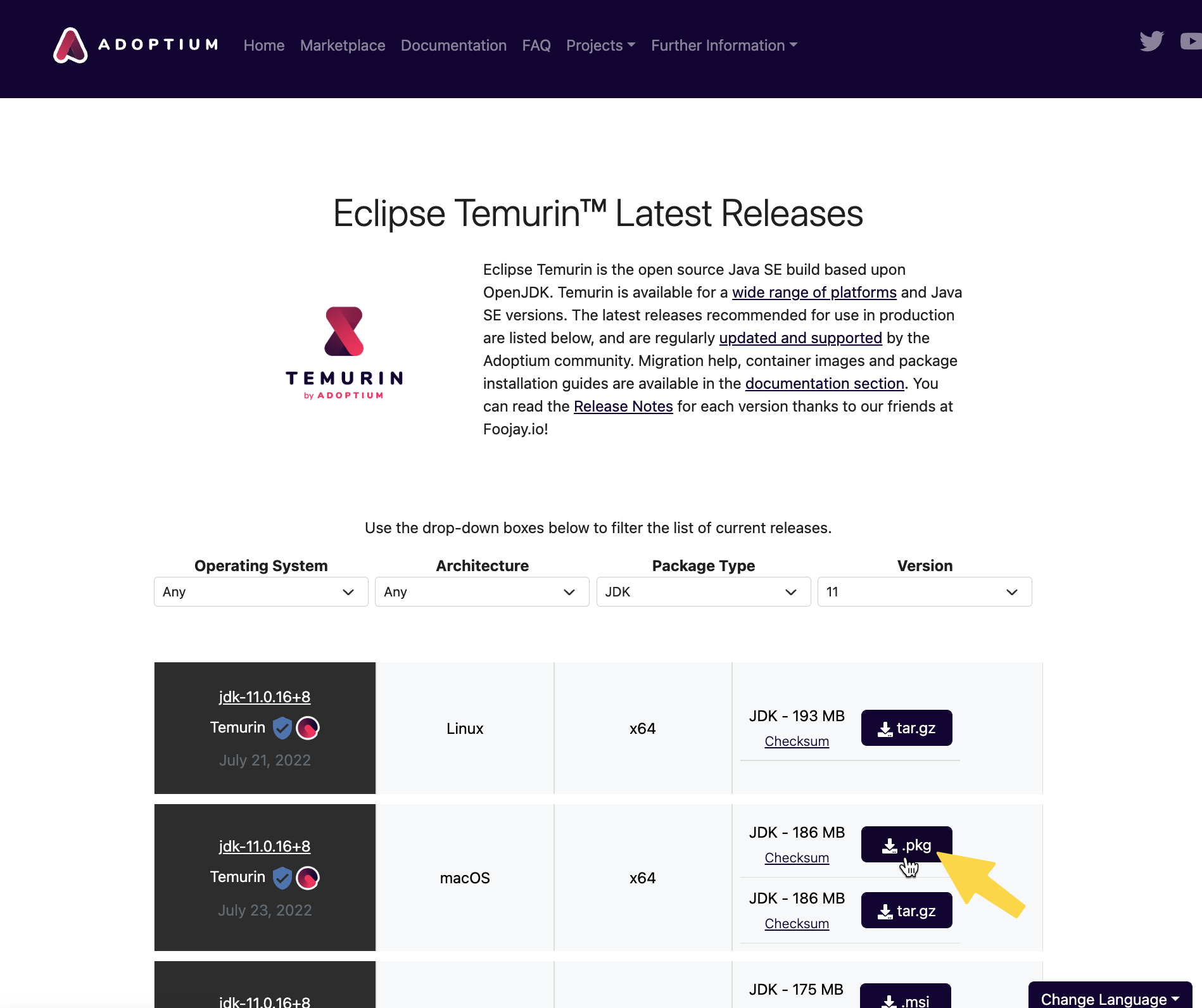
-1. **Download and install [Java JDK 11](https://adoptium.net/temurin/releases/?version=11)**
-
-  -
-
-
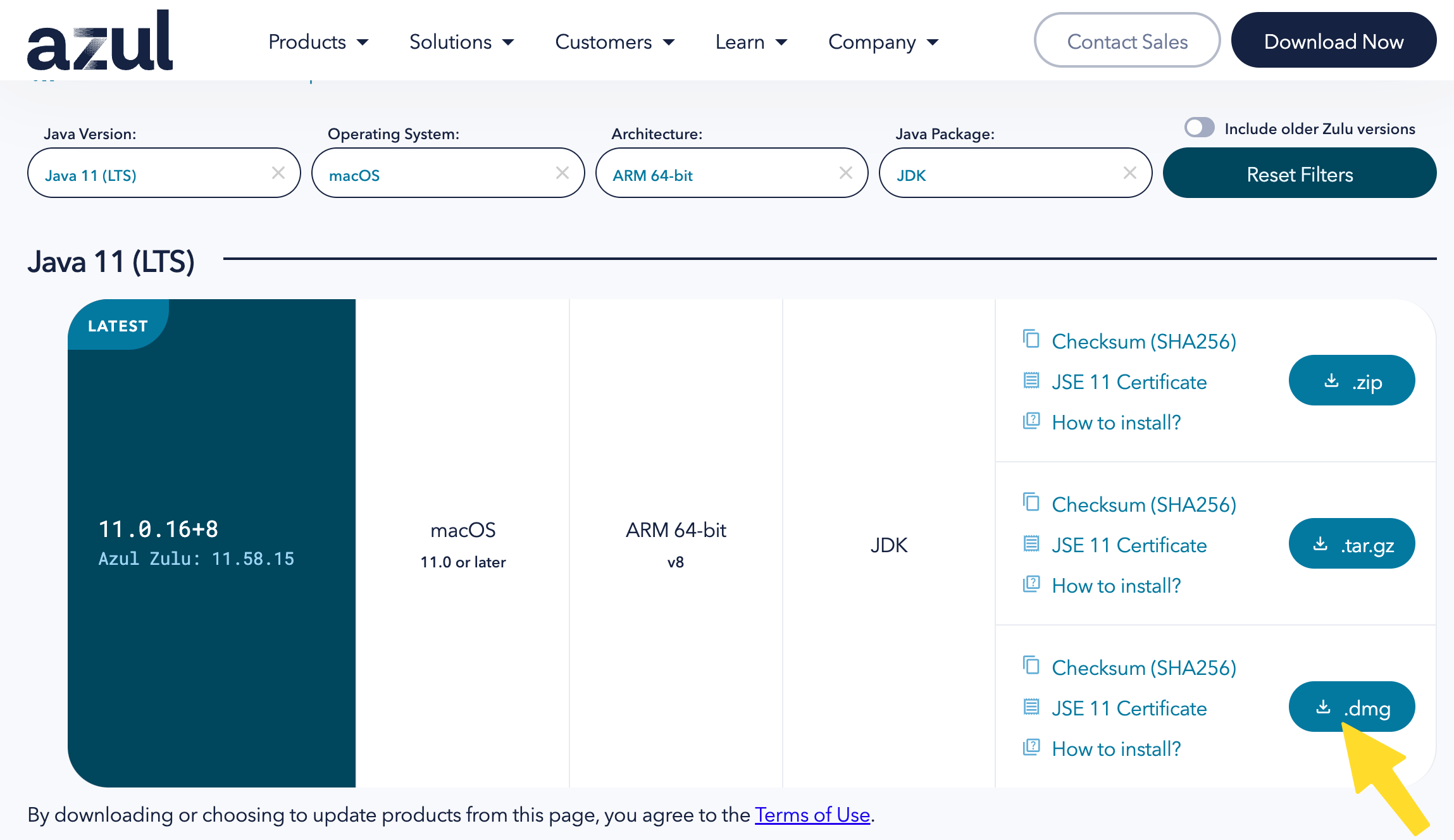
- **NOTE**: For M1/M2 Macs, which use the ARM 64-bit architecture, use [this link](https://www.azul.com/downloads/?version=java-11-lts&os=macos&architecture=arm-64-bit&package=jdk) to get Java 11.
-
-
-
-
-
- **NOTE**: For M1/M2 Macs, which use the ARM 64-bit architecture, use [this link](https://www.azul.com/downloads/?version=java-11-lts&os=macos&architecture=arm-64-bit&package=jdk) to get Java 11.
-
-  -
-
- After you download the file, open the installer and proceed through the steps to install Java 11.
-
-
-
-
-2. **Download and install or update conda**
-
- For beginners, we recommend you use Anaconda Navigator since it is more beginner-friendly, but you can also use miniconda. [Download Anaconda](https://www.anaconda.com/products/distribution) from the website and install.
-
- **NOTE**: if you already have conda, you can get the command to update conda by typing `conda update` on your command line. The command will generally look like:
-
- `conda update --prefix /Users/USERNAME/opt/anaconda3 anaconda`
-
-
-
-3. **Clone the CellProfiler-plugins Repo**
-
- Download a copy of (aka "clone") the CellProfiler-plugins repo from [here](https://github.com/CellProfiler/CellProfiler-plugins.git). You can download the whole repo by cloning it with git or simply clicking the green **Code** button on [the repo page](https://github.com/CellProfiler/CellProfiler-plugins.git) and selecting **Download ZIP** (see below).
-
-
-
-
- After you download the file, open the installer and proceed through the steps to install Java 11.
-
-
-
-
-2. **Download and install or update conda**
-
- For beginners, we recommend you use Anaconda Navigator since it is more beginner-friendly, but you can also use miniconda. [Download Anaconda](https://www.anaconda.com/products/distribution) from the website and install.
-
- **NOTE**: if you already have conda, you can get the command to update conda by typing `conda update` on your command line. The command will generally look like:
-
- `conda update --prefix /Users/USERNAME/opt/anaconda3 anaconda`
-
-
-
-3. **Clone the CellProfiler-plugins Repo**
-
- Download a copy of (aka "clone") the CellProfiler-plugins repo from [here](https://github.com/CellProfiler/CellProfiler-plugins.git). You can download the whole repo by cloning it with git or simply clicking the green **Code** button on [the repo page](https://github.com/CellProfiler/CellProfiler-plugins.git) and selecting **Download ZIP** (see below).
-
-  -
- You can also use git or GitHub Desktop to clone the repo if you prefer.
-
-
-
-4. **Create the environment from the .yml file**
-
- Open Anaconda Navigator and select the **Environments** tab on the left. We recommend you create the environment from the command line. To do this, Select the play button next to your base (root) environment and select **Open Terminal**:
-
-
-
- You can also use git or GitHub Desktop to clone the repo if you prefer.
-
-
-
-4. **Create the environment from the .yml file**
-
- Open Anaconda Navigator and select the **Environments** tab on the left. We recommend you create the environment from the command line. To do this, Select the play button next to your base (root) environment and select **Open Terminal**:
-
-  -
- A black box should pop up with a blinking cursor. This is your terminal. You now need to navigate to where the **cellprofiler_plugins_mac.yml** file is inside of the CellProfiler-plugins folder you downloaded in the last step. This file is in the `Instructions` subfolder. Here is how we recommend you do this:
- 1) In Finder, open the folder you downloaded in the previous step (usually called "CellProfiler-plugins-master")
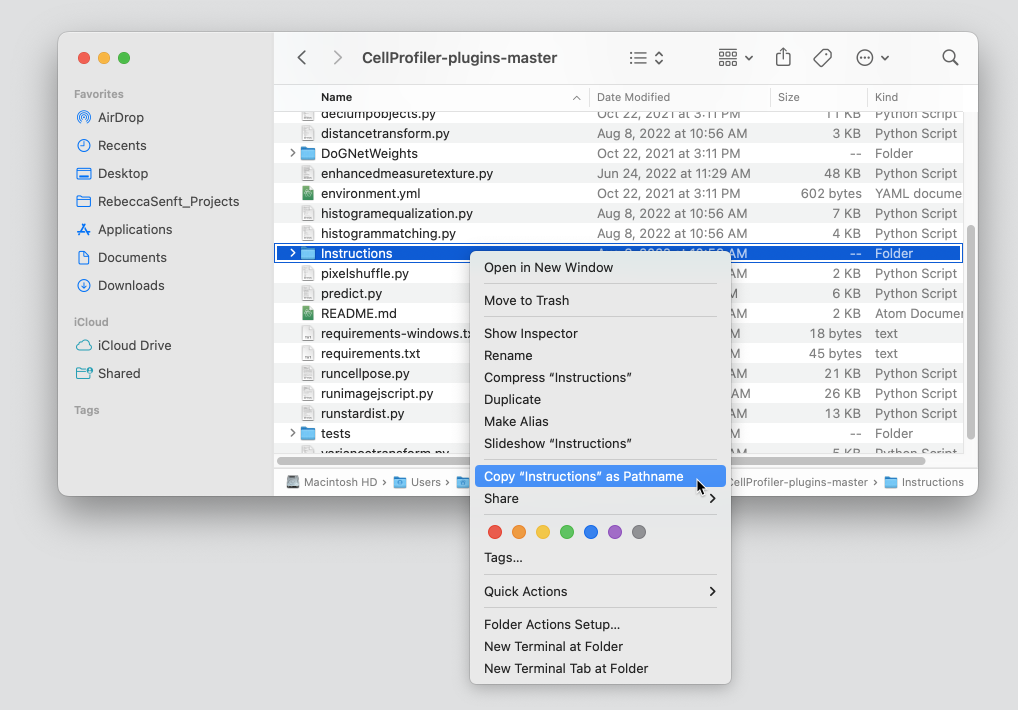
- 2) There should be a folder called **Instructions**. Right click or ctrl+click that folder. Then hold down the option key (or Alt) on your keyboard. An option to **Copy "Instructions" as Pathname** should appear. Select this option. (see below)
-
-
-
- A black box should pop up with a blinking cursor. This is your terminal. You now need to navigate to where the **cellprofiler_plugins_mac.yml** file is inside of the CellProfiler-plugins folder you downloaded in the last step. This file is in the `Instructions` subfolder. Here is how we recommend you do this:
- 1) In Finder, open the folder you downloaded in the previous step (usually called "CellProfiler-plugins-master")
- 2) There should be a folder called **Instructions**. Right click or ctrl+click that folder. Then hold down the option key (or Alt) on your keyboard. An option to **Copy "Instructions" as Pathname** should appear. Select this option. (see below)
-
-  -
-
- 3) Go back to your terminal and type `cd PATH_TO_FOLDER` where `PATH_TO_FOLDER` is the address you copied in the previous step. Press Enter.
-
-
-
- Now that you're in the right place, copy and paste this command into the terminal and press Enter.
- ```
- conda env create -f CellProfiler_plugins_mac.yml
- ```
-
-
-
- 5. **Activate your environment**
-
- In your terminal, enter `conda activate CellProfiler_plugins` to activate your environment
-
-
-
-
- 6. **Verify that cellprofiler is installed correctly by running it from the command line.**
-
- In your terminal, type in `pythonw -m cellprofiler` and hit Enter. this will open CellProfiler or will give you an error message.
-
-
-
- 7. **Install other packages for other plugins**
-
- Certain packages cannot easily installed when importing the environment.
-
- For StarDist: In terminal with your environment activated, enter:
- ```
- pip install stardist csbdeep --no-deps
- ```
-
-
- If you would like to use the omnipose models in cellpose, ensure you have cellpose 1.0.2 (you should by default if you've used the CellProfiler_plugins_mac.yml file to generate the environment) and enter on the command line (in your activated environment):
-
- ```
- pip install omnipose
- ```
-
-
-
-
- 8. **Connect CellProfiler and the plugins repo**
-
- With your environment active, type `pythonw -m cellprofiler` in terminal to open CellProfiler if it is not open already.
-
- * In CellProfiler, go to **File** then **Preferences...**
- * Scroll down and look for "CellProfiler Plugins Directory" on the left.
- * Select the **Browse** button and choose the folder where you extracted the CellProfiler plugins files. It is probably called "CellProfiler-plugins-master" unless you have renamed it.
- * Select **Save** at the bottom of the Preferences window
- * Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` on the command line
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
-
-
-
- 9. **Verify that the installation worked**
-
- Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
- In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
- 
-
- ---
-
-
-## Common errors
-
-1. My wheels are failing to build
-
-- If you get a message like "ERROR: Failed building wheel for pyzmq" this usually means that you do not have pyzmq installed. Try to reinstall pyzmq.
-
-2. Java virtual machine cannot be found
-
-- If you're getting errors about Java, it means that java is not being configured properly on your system.
-- Make sure you have installed The Java Development Kit 11 [here](https://adoptopenjdk.net/). Note that newer versions may not work.
-- Make sure you've added environment variables at the **System** level and not at the **User** level.
-```
-brew install java
-
-# For the system Java wrappers to find this JDK, symlink it with
-sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
-
-# Set version in zshrc
-echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
-source ~/.zshrc
-```
-
-3. Installing pyzmq failed
-
-- You might get an error when trying to install pyzmq. Something like
-```
-ERROR: Command errored out with exit status 1:
-```
-And earlier in the traceback:
-```
-Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
- Cython >= 0.20 is required to compile pyzmq from a development branch.
- Please install Cython or download a release package of pyzmq.
-```
-
- To fix this, `conda install cython`
-
-
-## Installation instructions for CellProfiler, CellProfiler plugins and CellPose in a conda environment on Apple silicon (eg M1).
-
-1. **Install brew**
- ```
- /bin/bash -c "$(curl -fsSL https://raw.githubusercontent.com/Homebrew/install/HEAD/install.sh)"
- ```
-
-2. **Update brew**
- ```
- brew update
- ```
-
-3. **Install Java**
- ```
- brew install java
- ```
-
-4. **For the system Java wrappers to find this JDK and symlink it**
- ```
- sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
- ```
-
-5. **Set version in .zshrc**
- ```
- echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
- source ~/.zshrc
- ```
-
-6. **Brew install package requirements**
- ```
- brew install freetype mysql git
- ```
-
-7. **Direct to brew openssl**
- ```
- export LDFLAGS="-L$(brew --prefix)/opt/openssl/lib"
- ```
-
-8. **Install hdf5**
- ```
- brew install hdf5@1.12
- ```
-
-9. **Make sure to get the version directory correct for the version installed. Find with `ls /opt/homebrew/Cellar/hdf5/`**
- ```
- export HDF5_DIR=/opt/homebrew/Cellar/hdf5/1.12.1_1/
- ```
-10. **Create a folder and download cellprofiler-plugins**
-
- ```
- mkdir cp_plugins
- cd cp_plugins
- git clone https://github.com/CellProfiler/CellProfiler-plugins
- ```
-
-11. **Download and install miniconda**
-
- ```
- wget https://repo.anaconda.com/miniconda/Miniconda3-latest-MacOSX-arm64.sh -O ~/miniconda.sh
- ```
-
-
-12. **Create a conda `.yml` file**
- In a text editor, paste the following test and save it as `cellprofiler_plugins_macM1.yml`:
- ```
- name: cpcellpose
-
- channels:
- - conda-forge
- - anaconda
- - bioconda
- - defaults
- - apple
-
- dependencies:
- - python=3.8
- - pip
- - h5py
- - mysqlclient
- - imagecodecs
- - python.app
- - pandas
- - pip:
- - cellpose
- - attrdict
- - sip==5.5.0
- - boto3>=1.12.28
- - centrosome==1.2.1
- - docutils==0.15.2
- - h5py~=3.6.0
- - imageio>=2.5
- - inflect>=2.1
- - Jinja2>=2.11.2
- - joblib>=0.13
- - mahotas>=1.4
- - matplotlib==3.1.3
- - mysqlclient==1.4.6
- - numpy>=1.20.1
- - Pillow>=7.1.0
- - prokaryote==2.4.4
- - python-bioformats==4.0.6
- - python-javabridge==4.0.3
- - pyzmq~=22.3
- - sentry-sdk==0.18.0
- - requests>=2.22
- - scikit-image>=0.17.2
- - scikit-learn>=0.20
- - scipy>=1.4.1
- - six
- - tifffile<2022.4.22
- - wxPython==4.2.0
- ```
-
-
-13. **Create a conda environment using the cellprofiler_plugins_macM1.yml file**
-
- At this stage, your folder/file structure should look like this:
-
- ```
- ├── cp_plugins
- ├── CellProfiler-plugins
- └── cellprofiler_plugins_macM1.yml
- ```
-
- In the terminal, make sure you are in the `cp_plugins` folder mentioned above (run `cd cp_plugins` to get there).
-
- ```
- conda env create -n cp_plugins --file cellprofiler_plugins_macM1.yml
- ```
-
-14. **Activate the conda environment**
-
- ```
- conda activate cp_plugins
- ```
-
-15. **Install CellProfiler and CellProfiler-core**
- ```
- pip install cellprofiler-core
-
- # We install CellProfiler with no dependencies here since we installed them with the conda yml file
- # This allows us to have some dependencies for M1 that are not the default for CellProfiler (eg wxPython)
- git clone https://github.com/CellProfiler/CellProfiler
-
- pip install --no-deps cellprofiler
- ```
-
-16. **Open CellProfiler**
- ```
- pythonw -m cellprofiler
- ```
-
-17. **Connect CellProfiler with the plugins folder**
-
- 1. In CellProfiler, go to `File` then `Preferences...`
- 2. Scroll down and look for `CellProfiler Plugins Directory` on the left.
- 3. Select the Browse button and choose the folder where you extracted the CellProfiler plugins files. It is probably called `CellProfiler-plugins` unless you have renamed it.
- 4. Select Save at the bottom of the Preferences window
- 5. Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` in the command line
-
-
-### Resolving dependencies conflicts
-
- There can be some strange dependency conflicts that can arise with python-javabridge. In the terminal with your environment activate, enter:
- ```
- pip uninstall -y centrosome python-javabridge
- pip install --no-cache-dir --no-deps --no-build-isolation python-javabridge centrosome
- pip uninstall matplotlib -y
- pip install matplotlib==3.2
- pip uninstall mahotas -y
- pip install mahotas
- ```
-
- The latest version of CellProfiler available on github removes the python-javabridge requirement and instead uses ScyJava.
-
-### Test your installation
-
-Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
-In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
diff --git a/documentation/CP-plugins-documentation/Windows_installation.md b/documentation/CP-plugins-documentation/Windows_installation.md
deleted file mode 100644
index 60bb701c..00000000
--- a/documentation/CP-plugins-documentation/Windows_installation.md
+++ /dev/null
@@ -1,208 +0,0 @@
-# How to install CellProfiler from source with all plugins on Windows
-
-Outlined here is the manual process for installing CellProfiler and plugins from source. Please close all open programs and save your work before starting this, as your computer may require a restart. Note that some of these download steps can take a while; the first 3 steps (as well as downloading Anaconda) can all be done simultaneously to save you time. Also note that for steps like changing the system-level environment variables, you may need admin permissions on your computer.
-
-**NEW:** We now have an automated install available for this workflow (requires administrator permissions on your computer) that will perform these steps for you. To get the necessary files and see instructions for the automatic install, look [here](https://github.com/CellProfiler/CellProfiler-plugins/tree/master/Instructions/Windows_batch_file).
-
-1. **Download and install [Microsoft Visual Studio C++ build tools](https://visualstudio.microsoft.com/visual-cpp-build-tools/)**
-
-
-
-
- 3) Go back to your terminal and type `cd PATH_TO_FOLDER` where `PATH_TO_FOLDER` is the address you copied in the previous step. Press Enter.
-
-
-
- Now that you're in the right place, copy and paste this command into the terminal and press Enter.
- ```
- conda env create -f CellProfiler_plugins_mac.yml
- ```
-
-
-
- 5. **Activate your environment**
-
- In your terminal, enter `conda activate CellProfiler_plugins` to activate your environment
-
-
-
-
- 6. **Verify that cellprofiler is installed correctly by running it from the command line.**
-
- In your terminal, type in `pythonw -m cellprofiler` and hit Enter. this will open CellProfiler or will give you an error message.
-
-
-
- 7. **Install other packages for other plugins**
-
- Certain packages cannot easily installed when importing the environment.
-
- For StarDist: In terminal with your environment activated, enter:
- ```
- pip install stardist csbdeep --no-deps
- ```
-
-
- If you would like to use the omnipose models in cellpose, ensure you have cellpose 1.0.2 (you should by default if you've used the CellProfiler_plugins_mac.yml file to generate the environment) and enter on the command line (in your activated environment):
-
- ```
- pip install omnipose
- ```
-
-
-
-
- 8. **Connect CellProfiler and the plugins repo**
-
- With your environment active, type `pythonw -m cellprofiler` in terminal to open CellProfiler if it is not open already.
-
- * In CellProfiler, go to **File** then **Preferences...**
- * Scroll down and look for "CellProfiler Plugins Directory" on the left.
- * Select the **Browse** button and choose the folder where you extracted the CellProfiler plugins files. It is probably called "CellProfiler-plugins-master" unless you have renamed it.
- * Select **Save** at the bottom of the Preferences window
- * Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` on the command line
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
-
-
-
- 9. **Verify that the installation worked**
-
- Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
- In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
- 
-
- ---
-
-
-## Common errors
-
-1. My wheels are failing to build
-
-- If you get a message like "ERROR: Failed building wheel for pyzmq" this usually means that you do not have pyzmq installed. Try to reinstall pyzmq.
-
-2. Java virtual machine cannot be found
-
-- If you're getting errors about Java, it means that java is not being configured properly on your system.
-- Make sure you have installed The Java Development Kit 11 [here](https://adoptopenjdk.net/). Note that newer versions may not work.
-- Make sure you've added environment variables at the **System** level and not at the **User** level.
-```
-brew install java
-
-# For the system Java wrappers to find this JDK, symlink it with
-sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
-
-# Set version in zshrc
-echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
-source ~/.zshrc
-```
-
-3. Installing pyzmq failed
-
-- You might get an error when trying to install pyzmq. Something like
-```
-ERROR: Command errored out with exit status 1:
-```
-And earlier in the traceback:
-```
-Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
- Cython >= 0.20 is required to compile pyzmq from a development branch.
- Please install Cython or download a release package of pyzmq.
-```
-
- To fix this, `conda install cython`
-
-
-## Installation instructions for CellProfiler, CellProfiler plugins and CellPose in a conda environment on Apple silicon (eg M1).
-
-1. **Install brew**
- ```
- /bin/bash -c "$(curl -fsSL https://raw.githubusercontent.com/Homebrew/install/HEAD/install.sh)"
- ```
-
-2. **Update brew**
- ```
- brew update
- ```
-
-3. **Install Java**
- ```
- brew install java
- ```
-
-4. **For the system Java wrappers to find this JDK and symlink it**
- ```
- sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
- ```
-
-5. **Set version in .zshrc**
- ```
- echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
- source ~/.zshrc
- ```
-
-6. **Brew install package requirements**
- ```
- brew install freetype mysql git
- ```
-
-7. **Direct to brew openssl**
- ```
- export LDFLAGS="-L$(brew --prefix)/opt/openssl/lib"
- ```
-
-8. **Install hdf5**
- ```
- brew install hdf5@1.12
- ```
-
-9. **Make sure to get the version directory correct for the version installed. Find with `ls /opt/homebrew/Cellar/hdf5/`**
- ```
- export HDF5_DIR=/opt/homebrew/Cellar/hdf5/1.12.1_1/
- ```
-10. **Create a folder and download cellprofiler-plugins**
-
- ```
- mkdir cp_plugins
- cd cp_plugins
- git clone https://github.com/CellProfiler/CellProfiler-plugins
- ```
-
-11. **Download and install miniconda**
-
- ```
- wget https://repo.anaconda.com/miniconda/Miniconda3-latest-MacOSX-arm64.sh -O ~/miniconda.sh
- ```
-
-
-12. **Create a conda `.yml` file**
- In a text editor, paste the following test and save it as `cellprofiler_plugins_macM1.yml`:
- ```
- name: cpcellpose
-
- channels:
- - conda-forge
- - anaconda
- - bioconda
- - defaults
- - apple
-
- dependencies:
- - python=3.8
- - pip
- - h5py
- - mysqlclient
- - imagecodecs
- - python.app
- - pandas
- - pip:
- - cellpose
- - attrdict
- - sip==5.5.0
- - boto3>=1.12.28
- - centrosome==1.2.1
- - docutils==0.15.2
- - h5py~=3.6.0
- - imageio>=2.5
- - inflect>=2.1
- - Jinja2>=2.11.2
- - joblib>=0.13
- - mahotas>=1.4
- - matplotlib==3.1.3
- - mysqlclient==1.4.6
- - numpy>=1.20.1
- - Pillow>=7.1.0
- - prokaryote==2.4.4
- - python-bioformats==4.0.6
- - python-javabridge==4.0.3
- - pyzmq~=22.3
- - sentry-sdk==0.18.0
- - requests>=2.22
- - scikit-image>=0.17.2
- - scikit-learn>=0.20
- - scipy>=1.4.1
- - six
- - tifffile<2022.4.22
- - wxPython==4.2.0
- ```
-
-
-13. **Create a conda environment using the cellprofiler_plugins_macM1.yml file**
-
- At this stage, your folder/file structure should look like this:
-
- ```
- ├── cp_plugins
- ├── CellProfiler-plugins
- └── cellprofiler_plugins_macM1.yml
- ```
-
- In the terminal, make sure you are in the `cp_plugins` folder mentioned above (run `cd cp_plugins` to get there).
-
- ```
- conda env create -n cp_plugins --file cellprofiler_plugins_macM1.yml
- ```
-
-14. **Activate the conda environment**
-
- ```
- conda activate cp_plugins
- ```
-
-15. **Install CellProfiler and CellProfiler-core**
- ```
- pip install cellprofiler-core
-
- # We install CellProfiler with no dependencies here since we installed them with the conda yml file
- # This allows us to have some dependencies for M1 that are not the default for CellProfiler (eg wxPython)
- git clone https://github.com/CellProfiler/CellProfiler
-
- pip install --no-deps cellprofiler
- ```
-
-16. **Open CellProfiler**
- ```
- pythonw -m cellprofiler
- ```
-
-17. **Connect CellProfiler with the plugins folder**
-
- 1. In CellProfiler, go to `File` then `Preferences...`
- 2. Scroll down and look for `CellProfiler Plugins Directory` on the left.
- 3. Select the Browse button and choose the folder where you extracted the CellProfiler plugins files. It is probably called `CellProfiler-plugins` unless you have renamed it.
- 4. Select Save at the bottom of the Preferences window
- 5. Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` in the command line
-
-
-### Resolving dependencies conflicts
-
- There can be some strange dependency conflicts that can arise with python-javabridge. In the terminal with your environment activate, enter:
- ```
- pip uninstall -y centrosome python-javabridge
- pip install --no-cache-dir --no-deps --no-build-isolation python-javabridge centrosome
- pip uninstall matplotlib -y
- pip install matplotlib==3.2
- pip uninstall mahotas -y
- pip install mahotas
- ```
-
- The latest version of CellProfiler available on github removes the python-javabridge requirement and instead uses ScyJava.
-
-### Test your installation
-
-Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
-In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
diff --git a/documentation/CP-plugins-documentation/Windows_installation.md b/documentation/CP-plugins-documentation/Windows_installation.md
deleted file mode 100644
index 60bb701c..00000000
--- a/documentation/CP-plugins-documentation/Windows_installation.md
+++ /dev/null
@@ -1,208 +0,0 @@
-# How to install CellProfiler from source with all plugins on Windows
-
-Outlined here is the manual process for installing CellProfiler and plugins from source. Please close all open programs and save your work before starting this, as your computer may require a restart. Note that some of these download steps can take a while; the first 3 steps (as well as downloading Anaconda) can all be done simultaneously to save you time. Also note that for steps like changing the system-level environment variables, you may need admin permissions on your computer.
-
-**NEW:** We now have an automated install available for this workflow (requires administrator permissions on your computer) that will perform these steps for you. To get the necessary files and see instructions for the automatic install, look [here](https://github.com/CellProfiler/CellProfiler-plugins/tree/master/Instructions/Windows_batch_file).
-
-1. **Download and install [Microsoft Visual Studio C++ build tools](https://visualstudio.microsoft.com/visual-cpp-build-tools/)**
-
- 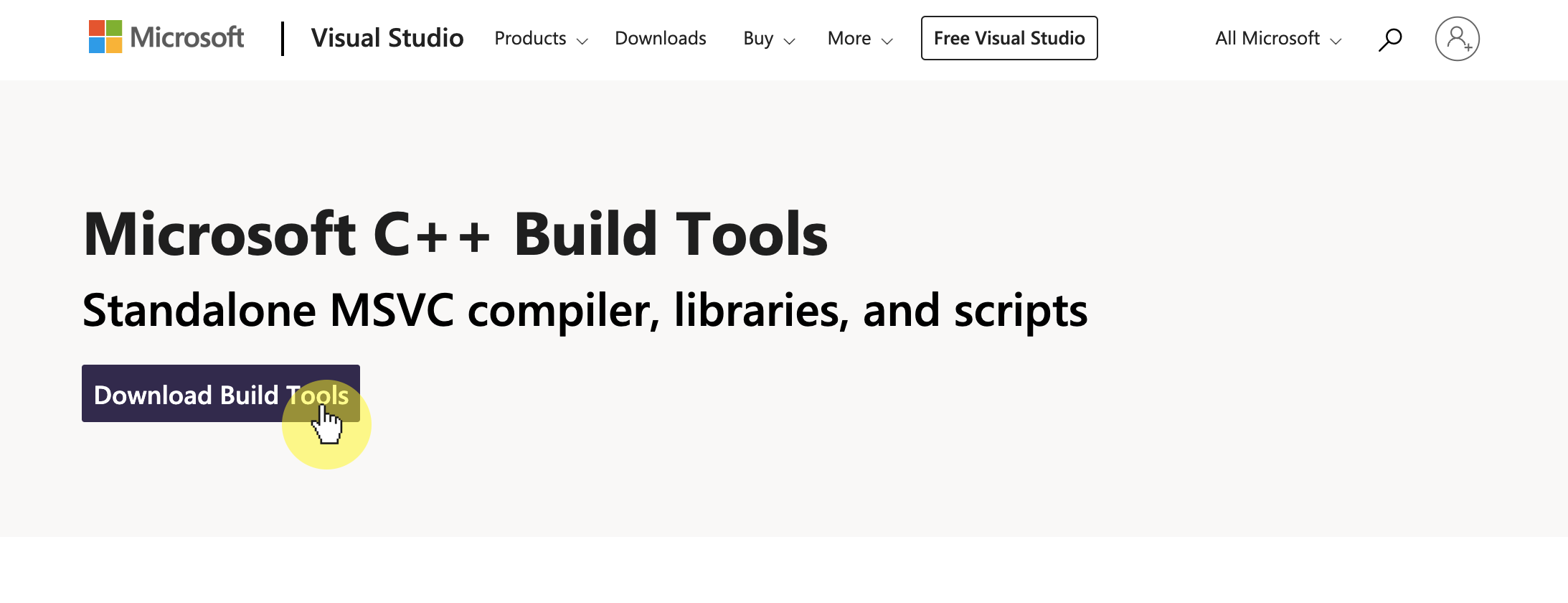 -
- **NOTE**: Make sure to check 'Desktop development with C++' under Desktop and Mobile in the installer:
-
-
-
- **NOTE**: Make sure to check 'Desktop development with C++' under Desktop and Mobile in the installer:
-
-  -
-
-
-2. **Download and install [Microsoft Visual C++ Redistributable 2015-2022]( https://docs.microsoft.com/en-us/cpp/windows/latest-supported-vc-redist?view=msvc-170)**
-
- You may already have this installed on your computer.
-
- Select the version appropriate for your architecture. On windows, you can determine this by going to **Control Panel** then searching for **System** and looking next to "System type:" for your processor architecture. Below is an example image:
-
-
-
-
-
-2. **Download and install [Microsoft Visual C++ Redistributable 2015-2022]( https://docs.microsoft.com/en-us/cpp/windows/latest-supported-vc-redist?view=msvc-170)**
-
- You may already have this installed on your computer.
-
- Select the version appropriate for your architecture. On windows, you can determine this by going to **Control Panel** then searching for **System** and looking next to "System type:" for your processor architecture. Below is an example image:
-
-  -
-
-
-
-3. **Download and install [Java JDK 11](https://adoptium.net/temurin/releases/?version=11)**
-
- Make sure you select the right version for your processor and operating system (see previous step). You can filter by operating system and processor Architecture.
-
-
-
-
-
-
-3. **Download and install [Java JDK 11](https://adoptium.net/temurin/releases/?version=11)**
-
- Make sure you select the right version for your processor and operating system (see previous step). You can filter by operating system and processor Architecture.
-
- 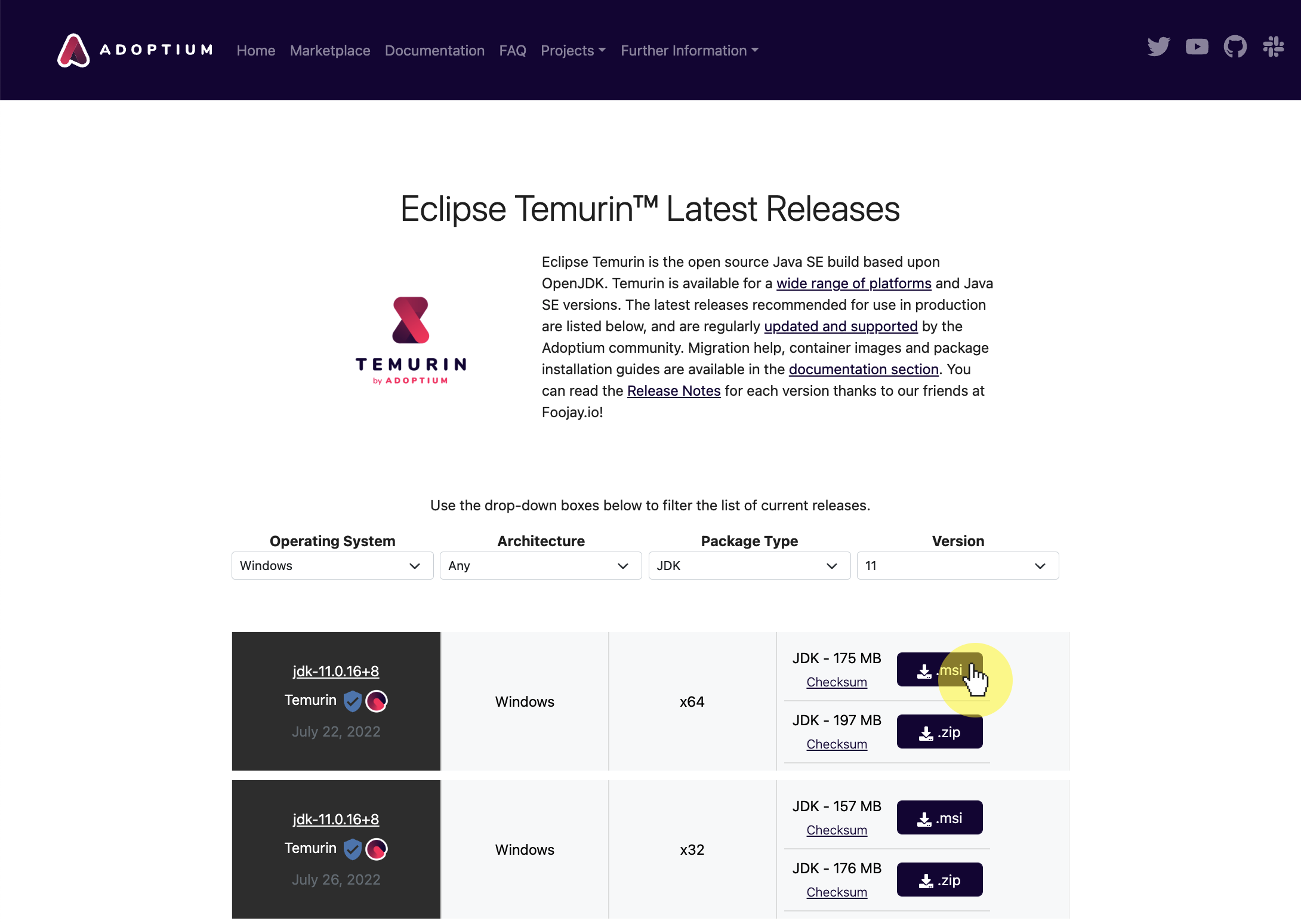 -
- You can alternatively install from [oracle.com](https://www.oracle.com/java/technologies/javase-jdk11-downloads.html) if you'd like, though you will need to make an Oracle account.
-
-
-
-
-4. **Update your Windows Environment Variables for Java**
-
- Go to **Control Panel** then search for **System**, then click on **Advanced System Settings**. Alternatively, for some versions of Windows, you can use the search bar to search for "Environment" and "Edit the system environment variables" will come up as an option.
-
-
-
- You can alternatively install from [oracle.com](https://www.oracle.com/java/technologies/javase-jdk11-downloads.html) if you'd like, though you will need to make an Oracle account.
-
-
-
-
-4. **Update your Windows Environment Variables for Java**
-
- Go to **Control Panel** then search for **System**, then click on **Advanced System Settings**. Alternatively, for some versions of Windows, you can use the search bar to search for "Environment" and "Edit the system environment variables" will come up as an option.
-
-  -
- From the System Properties window that pops up, select **Environment Variables...** Then add a new System variable by selecting **New...** under 'System Variables':
-
-
-
- From the System Properties window that pops up, select **Environment Variables...** Then add a new System variable by selecting **New...** under 'System Variables':
-
-  -
- Add (or if you already have it, modify) two system variables: `JAVA_HOME` and `JDK_HOME`. For each new variable, set its value to the location of your JDK installation (i.e., the location of the folder beginning with 'jdk11'). You can do this by clicking the **Browse Directory...** button. Usually this is in your 'Program Files' in a folder called 'Java'. Here is an example path for `JAVA_HOME`:
-
-
-
- Add (or if you already have it, modify) two system variables: `JAVA_HOME` and `JDK_HOME`. For each new variable, set its value to the location of your JDK installation (i.e., the location of the folder beginning with 'jdk11'). You can do this by clicking the **Browse Directory...** button. Usually this is in your 'Program Files' in a folder called 'Java'. Here is an example path for `JAVA_HOME`:
-
- 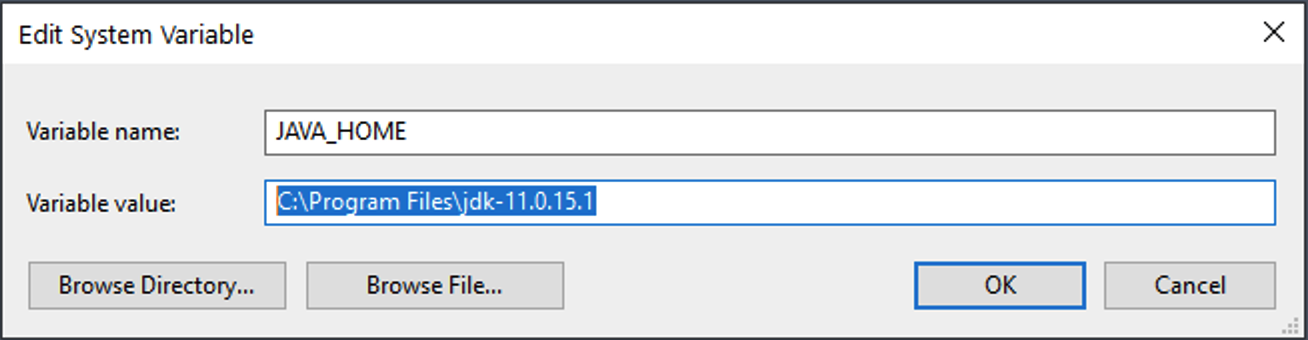 -
- Repeat the same for `JDK_HOME`:
-
-
-
- Repeat the same for `JDK_HOME`:
-
-  -
-
-
-
-
-5. **Download and install or update conda**
-
- For beginners, we recommend you use Anaconda Navigator since it is more beginner-friendly, but you can also use miniconda. [Download Anaconda](https://www.anaconda.com/products/distribution) from the website and install.
-
- **NOTE**: if you already have conda, you can get the command to update conda by typing `conda update` on your command line. The command will generally look like:
-
- `conda update --prefix /Users/USERNAME/opt/anaconda3 anaconda`
-
-
-
-
-6. **Clone the CellProfiler-plugins Repo**
-
- Download a copy of (aka "clone") the CellProfiler-plugins repo from [here](https://github.com/CellProfiler/CellProfiler-plugins.git). You can download the whole repo by cloning it with git or simply clicking the green **Code** button on [the repo page](https://github.com/CellProfiler/CellProfiler-plugins.git) and selecting **Download ZIP** (see below).
-
-
-
-
-
-
-
-5. **Download and install or update conda**
-
- For beginners, we recommend you use Anaconda Navigator since it is more beginner-friendly, but you can also use miniconda. [Download Anaconda](https://www.anaconda.com/products/distribution) from the website and install.
-
- **NOTE**: if you already have conda, you can get the command to update conda by typing `conda update` on your command line. The command will generally look like:
-
- `conda update --prefix /Users/USERNAME/opt/anaconda3 anaconda`
-
-
-
-
-6. **Clone the CellProfiler-plugins Repo**
-
- Download a copy of (aka "clone") the CellProfiler-plugins repo from [here](https://github.com/CellProfiler/CellProfiler-plugins.git). You can download the whole repo by cloning it with git or simply clicking the green **Code** button on [the repo page](https://github.com/CellProfiler/CellProfiler-plugins.git) and selecting **Download ZIP** (see below).
-
-  -
- If you download the ZIP file, be sure to extract the file contents by selecting **Extract All** in File Explorer:
-
-
-
- If you download the ZIP file, be sure to extract the file contents by selecting **Extract All** in File Explorer:
-
-  -
- You can also use git or GitHub Desktop to clone the repo if you prefer.
-
-
-
-7. **Create the environment from the .yml file**
-
- Open Anaconda Navigator and select the **Environments** tab on the left. We recommend you create the environment from the command line. To do this, Select the play button next to your base (root) environment and select **Open Terminal**:
-
-
-
- You can also use git or GitHub Desktop to clone the repo if you prefer.
-
-
-
-7. **Create the environment from the .yml file**
-
- Open Anaconda Navigator and select the **Environments** tab on the left. We recommend you create the environment from the command line. To do this, Select the play button next to your base (root) environment and select **Open Terminal**:
-
-  -
- A black box should pop up with a blinking cursor. This is your terminal. You now need to navigate to where the **cellprofiler_plugins_windows.yml** file is inside of the CellProfiler-plugins folder you downloaded in the last step. This file is in the `Instructions` subfolder. Here is how we recommend you do this:
-
- 1) In File Explorer, open the folder you downloaded in the previous step (usually called "CellProfiler-plugins-master") and then open the **Instructions** subfolder).
- 2) Click in the address bar area and this should highlight an address to where the folder is on your computer. **Copy this address**:
-
-
-
- A black box should pop up with a blinking cursor. This is your terminal. You now need to navigate to where the **cellprofiler_plugins_windows.yml** file is inside of the CellProfiler-plugins folder you downloaded in the last step. This file is in the `Instructions` subfolder. Here is how we recommend you do this:
-
- 1) In File Explorer, open the folder you downloaded in the previous step (usually called "CellProfiler-plugins-master") and then open the **Instructions** subfolder).
- 2) Click in the address bar area and this should highlight an address to where the folder is on your computer. **Copy this address**:
-
- 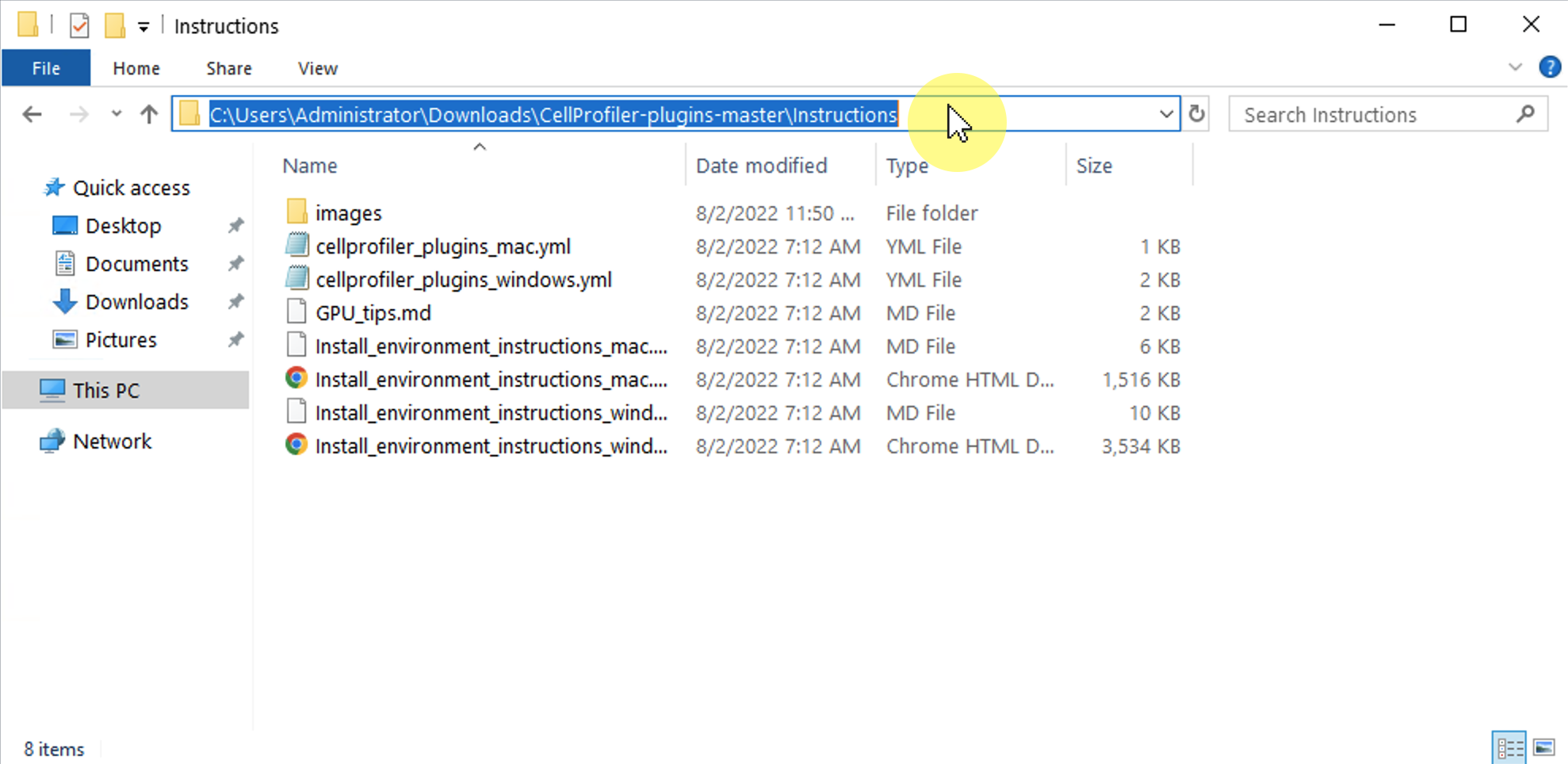 -
- 3) Go back to your terminal and type `cd PATH_TO_FOLDER` where `PATH_TO_FOLDER` is the address you copied in the previous step. Press Enter.
-
-
-
- Now that you're in the right place, copy and paste this command into the terminal and press Enter.
- ```
- conda env create -f CellProfiler_plugins_windows.yml
- ```
-
-
-8. **Activate your environment**
-
- In your terminal, enter `conda activate CP_plugins` to activate your environment
-
-
-
-9. **Verify that cellprofiler is installed correctly by running it from the command line.**
-
- In your terminal, type in `cellprofiler` and hit Enter. this will open CellProfiler or will give you an error message.
-
-
-
-10. **Install other packages for other plugins (just for RunStarDist)**
-
- In terminal with your environment activated, enter:
- ```
- pip install stardist csbdeep --no-deps
- ```
- If you would like to use the omnipose models in cellpose, ensure you have cellpose 1.0.2 (you should by default if you've used our environment yml) and enter on the command line (in your activated environment):
- ```
- pip install omnipose
- ```
-
-
-
-
-
-12. **Connect CellProfiler and the plugins folder**
-
- With your environment active, type `cellprofiler` in terminal to open CellProfiler if it is not open already.
-
- * In CellProfiler, go to **File** then **Preferences...**
- * Scroll down and look for "CellProfiler Plugins Directory" on the left.
- * Select the **Browse** button and choose the folder where you extracted the CellProfiler plugins files. It is probably called "CellProfiler-plugins-master" unless you have renamed it.
- * Select **Save** at the bottom of the Preferences window
- * Close CellProfiler and reopen it by typing `cellprofiler` on the command line
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
-
-
-
-13. **Verify that the installation worked**
-
- Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
- In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
-
-
- 3) Go back to your terminal and type `cd PATH_TO_FOLDER` where `PATH_TO_FOLDER` is the address you copied in the previous step. Press Enter.
-
-
-
- Now that you're in the right place, copy and paste this command into the terminal and press Enter.
- ```
- conda env create -f CellProfiler_plugins_windows.yml
- ```
-
-
-8. **Activate your environment**
-
- In your terminal, enter `conda activate CP_plugins` to activate your environment
-
-
-
-9. **Verify that cellprofiler is installed correctly by running it from the command line.**
-
- In your terminal, type in `cellprofiler` and hit Enter. this will open CellProfiler or will give you an error message.
-
-
-
-10. **Install other packages for other plugins (just for RunStarDist)**
-
- In terminal with your environment activated, enter:
- ```
- pip install stardist csbdeep --no-deps
- ```
- If you would like to use the omnipose models in cellpose, ensure you have cellpose 1.0.2 (you should by default if you've used our environment yml) and enter on the command line (in your activated environment):
- ```
- pip install omnipose
- ```
-
-
-
-
-
-12. **Connect CellProfiler and the plugins folder**
-
- With your environment active, type `cellprofiler` in terminal to open CellProfiler if it is not open already.
-
- * In CellProfiler, go to **File** then **Preferences...**
- * Scroll down and look for "CellProfiler Plugins Directory" on the left.
- * Select the **Browse** button and choose the folder where you extracted the CellProfiler plugins files. It is probably called "CellProfiler-plugins-master" unless you have renamed it.
- * Select **Save** at the bottom of the Preferences window
- * Close CellProfiler and reopen it by typing `cellprofiler` on the command line
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
-
-
-
-13. **Verify that the installation worked**
-
- Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
- In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
-  -
-
-
----
-
-## Common errors
-
-1. My wheels are failing to build
-
- - If you get a message like "ERROR: Failed building wheel for pyzmq" this usually means that you do not have the Microsoft Visual Studio tools installed properly or the wrong version of pyzmq is being installed (e.g., you have a previous version already installed in your environment). See Step #1-2 above and ensure that you have "Desktop development with C++" selected under the install configuration options
-
-
-2. Java virtual machine cannot be found. This error might look something like `javabridge.jutil.JVMNotFoundError`
-
- - If you're getting errors about Java, it means that java is not being configured properly on your system.
- - Make sure you have installed The Java Development Kit 11. Note that newer versions of Java may not work.
- - Make sure you've added environment variables at the **System** level and not at the **User** level. You need to add both `JAVA_HOME` and `JDK_HOME` and both need to point to the folder that holds your jdk installation. Typically this path would look something like `C:\Program Files\Java\jdk-11.0.15.1` but it might be different on your machine depending on where you've installed Java.
- - If you have done these things and you still get this error, then try `pip install javabridge` (inside your activated cellprofiler-plugins conda environment) and then opening cellprofiler with `cellprofiler`
-
-
-3. Installing pyzmq failed
-
- You might get an error when trying to install pyzmq. Something like
- ```
- ERROR: Command errored out with exit status 1:
- ```
- And earlier in the traceback:
- ```
- Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
- Cython >= 0.20 is required to compile pyzmq from a development branch.
- Please install Cython or download a release package of pyzmq.
- ```
-
- To fix this, activate your environment and then `conda install cython`
-
- - You might also get an error like:
- ```
- AttributeError: 'MSVCCompiler' object has no attribute '_vcruntime_redist'
- ```
-
- Generally, this error means that you don't have the right Microsoft Visual Studio C compiler. You can try two things:
-
- 1. Look below in the code. Even if the wheel fails, pyzmq will still attempt to install. Look below for `Running setup.py install for pyzmq ... done` Also make sure you don't have any additional versions of pyzmq installed already.
- 2. You can try reinstalling Microsoft Visual C++ Build tools (#1-2). Versions 2008, 2015-2022 should work. Make sure you have the version of the redistributable compatible with your system processor (see step #2).
-
----
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
new file mode 100644
index 00000000..0020d62b
--- /dev/null
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -0,0 +1,64 @@
+# Using plugins
+
+Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all plugins in the plugin folder that you have set.
+The plugins will appear in the "Add Modules" panel like all standard modules and you can use the plugins as you would any other CellProfiler module.
+
+Please note that, as CellProfiler-plugins are considered experimental, they may not be as well documented as standard modules and they may not have a window that shows on run.
+Please report any installation issues or bugs related to plugins in the [CellProfiler-plugins repository](https://github.com/CellProfiler/CellProfiler-plugins) and not in the main CellProfiler repository.
+
+## Installation
+
+### Installing plugins without dependencies
+
+If your plugin doesn't require additional dependencies, all you need to do is
+
+1. **Install CellProfiler.**
+Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
+
+Or, you can install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
+
+2. **Clone the CellProfiler-plugins repository.**
+This will download all of the plugins in the CellProfiler-plugins repository.
+
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+Alternatively, if you have code for plugins that are not in the CellProfiler-plugins repository, you can place them in any folder that you'll be able to find again.
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler and re-open it.
+
+You are now ready to use any CellProfiler plugin that does not have additional dependencies.
+This is most CellProfiler plugins.
+
+### Installing plugins with dependencies
+
+1. **Install CellProfiler.**
+Install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
+
+2. **Clone the CellProfiler-plugins repository.**
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler and re-open it.
+
+4. **Install the dependencies for your plugin.**
+In your terminal, type the following, where FLAG is the flag specific to the module you would like to run, as noted in [Supported Plugins](supported_plugins.md).
+```bash
+cd CellProfiler-plugins
+pip install -e .[FLAG]
+```
+
+Please note that plugins that have separate install flags may have conflicting dependencies so we recommend making a separate python environment in which to run separate installations.
+(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
\ No newline at end of file
From 4c41db2f05cabb63f634e1ff9c07f459ca39dd72 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 14:03:04 -0700
Subject: [PATCH 2/6] deprecate declumpobjects, add info about unsupported
plugins
---
.../unsupported_plugins.md | 14 +++++++++++++-
.../CellProfiler4/declumpobjects.py | 0
2 files changed, 13 insertions(+), 1 deletion(-)
rename declumpobjects.py => unmaintained_plugins/CellProfiler4/declumpobjects.py (100%)
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index 7e0343da..4ca10a6c 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -8,5 +8,17 @@ We welcome requests for updating particular unsupported plugins, but please note
We welcome community contributed plugin updates.
Likewise, we welcome contributions of community developed plugins.
-Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team or community contributed.
+Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
+## Where are unsupported plugins?
+
+Unsupported plugins can be found in the `unmaintained_plugins` folder in the CellProfiler-plugins repository.
+Those plugins in the `CellProfiler2`, `CellProfiler3`, and `CellProfiler4` folders were, at one point, supported for those versions of CellProfiler.
+Those plugins in the `CellProfiler4_autoconverted` folder were automatically converted from Python2 to Python3 (to support the transition from Python2 in CellProfiler3 to Python3 in CellProfiler4) but were never fully supported and may or may not run.
+
+## What plugins are unsupported?
+
+We cannot provide comprehensive information about why we are not supporting a given plugin.
+Information about select plugins is as follows:
+
+**DeclumpObjects**: DeclumpObjects will split objects based on a seeded watershed method. Functionality from this module was [added into CellProfiler](https://github.com/CellProfiler/CellProfiler/pull/4397) in the Watershed module as of CellProfiler 4.2.0.
diff --git a/declumpobjects.py b/unmaintained_plugins/CellProfiler4/declumpobjects.py
similarity index 100%
rename from declumpobjects.py
rename to unmaintained_plugins/CellProfiler4/declumpobjects.py
From d47666e9fcefc1f98227e02aa4692ed83a022e0c Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 14:20:25 -0700
Subject: [PATCH 3/6] more documentation
---
.../contributing_plugins.md | 17 +++-
.../CP-plugins-documentation/overview.md | 11 +++
.../supported_plugins.md | 77 +++++--------------
.../CP-plugins-documentation/using_GPU.md | 8 ++
4 files changed, 51 insertions(+), 62 deletions(-)
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index 204ec8f8..c3ae78be 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -1,6 +1,8 @@
-## Contributing New Plugins
+# Contributing New Plugins
-Refer to CellProfiler repository and within cellprofiler/modules/plugins you can find two different templates to use for creating your own plugin.
+Within our CellProfiler wiki, you can find an [orientation to CellProfiler code](https://github.com/CellProfiler/CellProfiler/wiki/Orientation-to-CellProfiler-code).
+
+In the CellProfiler repository, within [cellprofiler/modules/plugins](https://github.com/CellProfiler/CellProfiler/tree/master/cellprofiler/modules/plugins) you can find two different templates to use for creating your own plugin.
`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
@@ -8,10 +10,17 @@ In you plugin, you must include:
In your plugin, we appreciate if you also include:
- display functionality
+- extensive module documentation
+- references and citation information in your module documentation
+Please create a Pull Request to the CellProfiler-plugins repository to submit your plugin for inclusion in the repository.
-In your PR, you must include:
+In your PR, you must:
- add your plugin to the [supported_plugins](supported_plugins.md) documentation page
In your PR, we appreciate if you also include:
-- unit tests for your plugin
\ No newline at end of file
+- unit tests for your plugin
+
+## Having your plugin cited
+
+While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/overview.md b/documentation/CP-plugins-documentation/overview.md
index 5ece4c8d..1616e1a3 100644
--- a/documentation/CP-plugins-documentation/overview.md
+++ b/documentation/CP-plugins-documentation/overview.md
@@ -9,6 +9,17 @@ A module may be in CellProfiler-plugins instead of CellProfiler itself because:
- it requires extra libraries or other dependencies we are unable or unwilling to require for CellProfiler
- it has been contributed by a community member
+## How do I use CellProfiler Plugins?
+
+See our [Using Plugins](using_plugins.md) page for information on how to install and use plugins.
+
+## What plugins are available?
+
+See our [Supported Plugins](supported_plugins.md) page for information on all currently supported plugins.
+
+## How do I contribute a plugin?
+
+See our [Contributing Plugins](contributing_plugins.md) page for information on contributing a plugin to CellPainting-plugins.
## Who made this?
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index 5850822a..deb60d61 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -3,61 +3,22 @@
Below is a brief overview of our currently supported plugins.
For details about using any particular plugin, please read the module documentation inside the plugin in CellProfiler.
-## CalculateMoments
-CalculateMoments extracts moments statistics from a given distribution of pixel values.
-
-## CallBarcodes
-CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles.
-It is used for optical sequencing by synthesis (SBS).
-
-## CompensateColors
-CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through.
-It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched.
-
-## DeclumpObjects
-DeclumpObjects will split objects based on a seeded watershed method.
-
-## DistanceTransform
-DistanceTransform computes the distance transform of a binary image.
-The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1.
-
-## EnhancedMeasureTexture
-EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler.
-
-## HistogramEqualization
-HistogramEqualization increases the global contrast of a low-contrast image or volume.
-Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct.
-This module can perform either global or local histogram equalization.
-
-## HistogramMatching
-HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image.
-It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image.
-It allows you to choose which frame to use as the reference.
-
-## PixelShuffle
-PixelShuffle takes the intensity of each pixel in an image and it randomly shuffles its position.
-
-## Predict
-Predict allows you to use an ilastik pixel classifier to generate a probability image.
-CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage).
-
-## RunCellpose
-RunCellpose allows you to run Cellpose within CellProfiler.
-Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells.
-You can use pre-trained Cellpose models or your custom model with this plugin.
-You can use a GPU with this module to dramatically increase your speed/efficiency.
-
-## RunImageJScript
-
-## RunOmnipose
-
-## RunStarDist
-RunStarDist allows you to run StarDist within CellProfiler.
-StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells.
-You can use pre-trained StarDist models or your custom model with this plugin.
-You can use a GPU with this module to dramatically increase your speed/efficiency.
-RunStarDist is generally faster than RunCellpose.
-
-## VarianceTransform
-This module allows you to calculate the variance of an image, using a determined window size.
-It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image.
\ No newline at end of file
+Most plugins will run without any special installation of either CellProfiler or the plugins.
+See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin use as well as for installation information for those plugins that do require installation of dependencies.
+
+| Plugin | Description | Requires installation of dependencies? | Install flag |
+|--------|-------------|----------------------------------------|--------------|
+| CalculateMoments | CalculateMoments extracts moments statistics from a given distribution of pixel values. | No | |
+| CallBarcodes | CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles. It is used for optical sequencing by synthesis (SBS). | No | |
+| CompensateColors | CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through. It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched. | No | |
+| DistanceTransform | DistanceTransform computes the distance transform of a binary image. The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1. | No | |
+| EnhancedMeasureTexture| EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler. | No | |
+| HistogramEqualization | HistogramEqualization increases the global contrast of a low-contrast image or volume. Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct. This module can perform either global or local histogram equalization. | No | |
+| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
+| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
+| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | Yes | XXXXX |
+| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
+| RunImageJScript | | Yes | XXXXX |
+| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
+| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
+| VarianceTransform | This module allows you to calculate the variance of an image, using a determined window size. It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image. | No | |
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/using_GPU.md b/documentation/CP-plugins-documentation/using_GPU.md
index 945d869b..b5dfc4d1 100644
--- a/documentation/CP-plugins-documentation/using_GPU.md
+++ b/documentation/CP-plugins-documentation/using_GPU.md
@@ -24,3 +24,11 @@ If you want to use a GPU to run the model (this is recommended for speed), you'l
pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
```
If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
+
+
+ **NOTE**: You might get a warning like this:
+ ```
+ W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
+ 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
+ ```
+ If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
From 12c52c77ec80879b399743160d6accbfe2049d26 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 12:35:02 -0700
Subject: [PATCH 4/6] more docs
---
.../contributing_plugins.md | 3 +-
.../supported_plugins.md | 4 +-
.../unsupported_plugins.md | 4 +-
.../CP-plugins-documentation/using_plugins.md | 106 ++++++++++++++++--
.../CP-plugins-documentation/versions.md | 49 +-------
5 files changed, 104 insertions(+), 62 deletions(-)
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index c3ae78be..f0084668 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -6,8 +6,6 @@ In the CellProfiler repository, within [cellprofiler/modules/plugins](https://gi
`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
-In you plugin, you must include:
-
In your plugin, we appreciate if you also include:
- display functionality
- extensive module documentation
@@ -20,6 +18,7 @@ In your PR, you must:
In your PR, we appreciate if you also include:
- unit tests for your plugin
+- customized installation in setup.py, if your plugin has dependencies
## Having your plugin cited
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index deb60d61..a17866c0 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -16,9 +16,9 @@ See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin
| HistogramEqualization | HistogramEqualization increases the global contrast of a low-contrast image or volume. Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct. This module can perform either global or local histogram equalization. | No | |
| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
-| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | Yes | XXXXX |
+| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | No | |
| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
-| RunImageJScript | | Yes | XXXXX |
+| RunImageJScript | RunImageJScript allows you to run any supported ImageJ script directly within CellProfiler. It is significantly more performant than RunImageJMacro, and is also less likely to leave behind temporary files. | Yes | XXXXX |
| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
| VarianceTransform | This module allows you to calculate the variance of an image, using a determined window size. It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image. | No | |
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index 4ca10a6c..b7cf3fe8 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -5,11 +5,9 @@ Unsupported plugins are primarily unsupported for either of two reasons:
- Their functions have been integrated into the current version of CellProfiler and therefore a plugin is no longer necessary
We welcome requests for updating particular unsupported plugins, but please note that we have limited bandwidth for working on plugins and may be unable to complete the update.
+Additionally, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
We welcome community contributed plugin updates.
-Likewise, we welcome contributions of community developed plugins.
-Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
-
## Where are unsupported plugins?
Unsupported plugins can be found in the `unmaintained_plugins` folder in the CellProfiler-plugins repository.
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index 0020d62b..bce4bc70 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -8,9 +8,16 @@ Please report any installation issues or bugs related to plugins in the [CellPro
## Installation
-### Installing plugins without dependencies
+If the plugin you would like to use does not have any additional dependencies outside of those required for running CellProfiler (this is most plugins), using plugins is very simple.
+See [Installing plugins without dependencies](#installing-plugins-without-dependencies).
+
+If the plugin you would like to use has dependencies, you have two separate options for installation.
+The first option requires building CellProfiler from source, but plugin installation is simpler.
+See [Installing plugins with dependencies, using CellProfiler from source](#installing-plugins-with-dependencies-using-cellprofiler-from-source).
+The second option allows you to use pre-built CellProfiler, but plugin installation is more complex.
+See [Installing plugins with dependencies, using pre-built CellProfiler](#installing-plugins-with-dependencies-using-pre-built-cellprofiler).
-If your plugin doesn't require additional dependencies, all you need to do is
+### Installing plugins without dependencies
1. **Install CellProfiler.**
Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
@@ -31,12 +38,12 @@ Alternatively, if you have code for plugins that are not in the CellProfiler-plu
- Open CellProfiler.
- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
- Select `Save` at the bottom of the Preferences window
-- Close CellProfiler and re-open it.
+- Close CellProfiler and re-open it
You are now ready to use any CellProfiler plugin that does not have additional dependencies.
This is most CellProfiler plugins.
-### Installing plugins with dependencies
+### Installing plugins with dependencies, using CellProfiler from source
1. **Install CellProfiler.**
Install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
@@ -49,9 +56,9 @@ git clone https://github.com/CellProfiler/CellProfiler-plugins.git
3. **Set the plugins path in CellProfiler.**
- Open CellProfiler.
-- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned.
- Select `Save` at the bottom of the Preferences window
-- Close CellProfiler and re-open it.
+- Close CellProfiler
4. **Install the dependencies for your plugin.**
In your terminal, type the following, where FLAG is the flag specific to the module you would like to run, as noted in [Supported Plugins](supported_plugins.md).
@@ -59,6 +66,91 @@ In your terminal, type the following, where FLAG is the flag specific to the mod
cd CellProfiler-plugins
pip install -e .[FLAG]
```
+e.g. To install Cellpose the pip install command would be `pip install -e .[cellpose]`
+
+5. **Open and use CellProfiler.**
Please note that plugins that have separate install flags may have conflicting dependencies so we recommend making a separate python environment in which to run separate installations.
-(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
\ No newline at end of file
+(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
+
+### Installing plugins with dependencies, using pre-built CellProfiler
+
+1. **Install CellProfiler.**
+Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
+
+2. **Clone the CellProfiler-plugins repository.**
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned.
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler
+
+4. **Identify the Python dependencies of the plugins you want to use.**
+You can find this information for most plugins by looking in `setup.py`.
+```{admonition} e.g. using RunCellpose plugin
+If you would like to install the RunCellpose plugin, in `setup.py` you can see that `cellpose_deps = ["cellpose>=1.0.2"]` so the only dependency is `cellpose`.
+```
+
+Alternatively, you can find this information directly in the plugin code itself by looking at what is imported at the beginning of the plugin.
+Note that you will want to compare any imports to what is already required by CellProfiler with that caveat that dependencies often have dependencies.
+```{admonition} e.g. using RunImageJScript plugin
+If you would like to install the RunImageJScript plugin, you'll notice a number of imports in the beginning of the plugin code.
+You can ignore all those from `cellprofiler_core` as they are required for CellProfiler installation.
+You can ignore all those that are native to Python such as `sys`, `time`, and `threading`.
+A comparison with CellProfiler dependencies will show you that `skimage` is already installed (you can see this in CellProfiler's [environment.yml](https://github.com/CellProfiler/CellProfiler/blob/master/environment.yml) which lists `scikit-image` as a `cellprofiler-core` dependency or in CellProfiler's [setup.py](https://github.com/CellProfiler/CellProfiler/blob/master/setup.py) which lists `scikit-image` as an installation requirement).
+What remains is `pyimagej`
+```
+
+5. **Create a conda environment and install requirements in it**
+Create a conda (or other virtual) environment with a Python version matching that of CellProfiler.
+CellProfiler 4 uses Python 3.8 (which you can see in CellProfiler's [setup.py](https://github.com/CellProfiler/CellProfiler/blob/master/setup.py)).
+Activate the environment and install the plugin requirement/s into it.
+
+In your terminal, type
+```
+# Create a conda environment
+conda create --name ENV_NAME python=3.8
+# Activate the conda environment
+conda activate ENV_NAME
+# Install the plugin requirement/s
+pip install REQUIREMENT
+```
+```{admonition} e.g. using RunImageJScript plugin
+conda create --name cp-ij python=3.8
+conda activate cp-ij
+pip install pyimagej
+```
+
+6. **Copy plugin requirements into CellProfiler installation**
+Find the folder that you installed into in the previous step (where to find these libraries in your local environment) by entering `pip show REQUIREMENT` into your terminal e.g. `pip show pyimagej`.
+In the information it returns, under `Location` you will find a path that will look something like `/Users/eweisbar/mambaforge/envs/cp-ij/lib/python3.8/site-packages`.
+
+Find the folder in which you have installed CellProfiler.
+On Windows this is likely to be `C:\Program Files\CellProfiler` and on Mac it is likely to be `/Applications/CellProfiler.app/Contents/MacOS/`.
+In your terminal, if you `ls` (for Mac) or `dir` (for Windows) that folder (e.g. `ls /Applications/CellProfiler.app/Contents/MacOS/`), you should see a list of all the dependencies that were packaged with CellProfiler.
+
+Copy the folders and their corresponding `.dist-info` folders (e.g. `pytz` and `pytz-2023.3.dist-info`) for any dependencies that were installed with the installation of `pyimagej` that are not already in the CellProfiler folder.
+
+```{admonition} e.g. using RunImageJScript plugin
+These are all the folders you need to copy over:
+`pytz`
+`pytz-2023.3.dist-info`
+`xarray`
+`xarray-2023.1.0.dist-info`
+`imglyb`
+`imglyb-2.1.0.dist-info`
+`jgo`
+`jgo-1.0.5.dist-info`
+`jpype`
+`_jpype.cpython-38-darwin.so`
+`imagej`
+`pyimagej-1.4.1.dist-info`
+```
+
+7. **Open and use CellProfiler.**
+When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/versions.md b/documentation/CP-plugins-documentation/versions.md
index 3755e75b..0f8db604 100644
--- a/documentation/CP-plugins-documentation/versions.md
+++ b/documentation/CP-plugins-documentation/versions.md
@@ -1,55 +1,8 @@
# Versions
-The most current release can always be found on DockerHub at `cellprofiler/distributed-cellprofiler`.
-Current version is 2.0.0.
-Our current tag system is dcpversion_CellProfilerversion, e.g. 1.2.0_3.1.8
-
-Previous release versions can be accessed at `bethcimini/distributed-cellprofiler:versionnumber`
+Current version is
---
# Version History
-## 2.1.0 (forthcoming)
-* Increase support for using plugins
-* Improve role handling and allow file download/upload from/to S3 buckets in different accounts
-* Improve file download to S3FS can be completely bypassed
-* Name EBS volumes
-
-## 2.0.0rc2 - Released 20201110
-* Add optional ability to download files to EBS rather than reading from S3 (helpful for pipelines that access many files/image sets)
-
-## 2.0.0rc1 - Released 20201105
-
-* Remove requirement for boto and fabric, using only boto3
-* Add support for Python 3 and CellProfiler 4
-* Move cluster size, machine type, and machine price to the config file from the fleet file, eliminating mismatches between the two
-* Add the ability to filter only files with certain names when running CHECK_IF_DONE
-* Don't cancel a fleet over capacity errors
-* Add "cheapest" mode to the monitor, allowing you to run more cheaply (at possible expense of running more slowly)
-
-## 1.2.2 - Released 20201103
-
-* Allow pipelines using batch files to also designate an input output_top_directory
-* Add support for multiple LaunchData specifications
-* Add CellProfiler-plugins
-* Additional way to create job submissions with run_batch_general.py
-
-## 1.2.1 - Released 20200109, Updated through 20191002
-
-* Allow monitor to downscale machines when number of jobs < number of machines
-* Add a parameter to discount files when running CHECK_IF_DONE checks if less than a certain size
-
-## 1.2.0 - Released 20181108, Updated through 20200109
-
-* Improved compatibility with CellProfiler 2 and 3
-* Better handling of logging when using output_structure
-
-## 1.1.0 - Released 20170217, Updated 20170221 (bugfixes) - 20181018
-
-* Changes in this release:
-
- * Added the `output_structure` variable to the job file, which allows you to structure the output folders created by DCP (ie `Plate/Well-Site` rather than `Plate-Well-Site`). Job files lacking this variable will still default to the previous settings (hyphenating all Metadata items in order they are presented in the Metadata grouping).
- * Added support for creating the list of groups via `cellprofiler --print-groups`- see [this issue](https://github.com/CellProfiler/Distributed-CellProfiler/issues/52) for example and discussion. Groups listed in this way MUST use the `output_structure` variable to state their desired folder structure or an error will be returned.
-
-## 1.0.0 - Version as of 20170213
From fffb5566c3b4d2892c75c30b14469e66e94bca6a Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 15:04:09 -0700
Subject: [PATCH 5/6] more doc changes
---
.../CP-plugins-documentation/RunCellPose.md | 34 +++++++++++++++++++
.../contributing_plugins.md | 9 ++++-
.../supported_plugins.md | 5 ++-
.../unsupported_plugins.md | 1 +
.../CP-plugins-documentation/using_GPU.md | 34 -------------------
.../CP-plugins-documentation/using_plugins.md | 3 +-
6 files changed, 49 insertions(+), 37 deletions(-)
create mode 100644 documentation/CP-plugins-documentation/RunCellPose.md
delete mode 100644 documentation/CP-plugins-documentation/using_GPU.md
diff --git a/documentation/CP-plugins-documentation/RunCellPose.md b/documentation/CP-plugins-documentation/RunCellPose.md
new file mode 100644
index 00000000..168fa7cc
--- /dev/null
+++ b/documentation/CP-plugins-documentation/RunCellPose.md
@@ -0,0 +1,34 @@
+# RunCellPose
+
+## Using RunCellPose with a GPU
+
+If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU.
+General instructions are available at this [link](https://pytorch.org/get-started/locally/).
+
+1. Your GPU should be visible in Device Manager under Display Adaptors.
+If your GPU isn't there, you likely need to install drivers.
+[Here](https://www.nvidia.com/Download/Find.aspx) is where you can find NVIDIA GPU drivers if you need to install them.
+
+
+2. To test whether the GPU is configured correctly:
+ * Run `python` on the command line (i.e., in Command Prompt or Terminal) to start an interactive session
+ * Then run the following
+ ```
+ import torch
+ torch.cuda.is_available()
+ ```
+ * If this returns `True`, you're all set
+ * If this returns `False`, you likely need to install/reinstall torch. See [here](https://pytorch.org/get-started/locally/) for your exact command.
+ * Exit the session with `exit()` then install torch if necessary
+ ```
+ pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
+ ```
+ If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
+
+
+**NOTE**: You might get a warning like this:
+```
+W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
+2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
+```
+If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index f0084668..8907b250 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -20,6 +20,13 @@ In your PR, we appreciate if you also include:
- unit tests for your plugin
- customized installation in setup.py, if your plugin has dependencies
+## Contributing bug fixes or updating deprecated plugins
+
+Please create a Pull Request to the CellProfiler-plugins repository to submit a bug fix or plugin un-deprecation.
+If you would like to update a currently deprecated plugin, we encourage you to read closed Issues in GitHub relevant to the plugin as they may contain helpful information about bugs present at the time of deprecation.
+
## Having your plugin cited
-While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
\ No newline at end of file
+While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
+
+See [Citing Plugins](citing.md) for information on citing CellProfiler Plugins.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index a17866c0..5ccdda93 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -6,6 +6,9 @@ For details about using any particular plugin, please read the module documentat
Most plugins will run without any special installation of either CellProfiler or the plugins.
See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin use as well as for installation information for those plugins that do require installation of dependencies.
+Most plugin documentation can be found within the plugin itself and can be accessed through CellProfiler help.
+Those plugins that do have extra documentation contain links below.
+
| Plugin | Description | Requires installation of dependencies? | Install flag |
|--------|-------------|----------------------------------------|--------------|
| CalculateMoments | CalculateMoments extracts moments statistics from a given distribution of pixel values. | No | |
@@ -17,7 +20,7 @@ See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin
| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | No | |
-| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
+| [RunCellpose](RunCellPose.md) | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
| RunImageJScript | RunImageJScript allows you to run any supported ImageJ script directly within CellProfiler. It is significantly more performant than RunImageJMacro, and is also less likely to leave behind temporary files. | Yes | XXXXX |
| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index b7cf3fe8..5e277b61 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -19,4 +19,5 @@ Those plugins in the `CellProfiler4_autoconverted` folder were automatically con
We cannot provide comprehensive information about why we are not supporting a given plugin.
Information about select plugins is as follows:
+**ClassifyPixelsUNET**: ClassifyPixelsUNET is a pixel classifier for background/object edge/object body. As far as we are aware, other deep learning based plugins that we do currently support (such as RunCellpose) work better.
**DeclumpObjects**: DeclumpObjects will split objects based on a seeded watershed method. Functionality from this module was [added into CellProfiler](https://github.com/CellProfiler/CellProfiler/pull/4397) in the Watershed module as of CellProfiler 4.2.0.
diff --git a/documentation/CP-plugins-documentation/using_GPU.md b/documentation/CP-plugins-documentation/using_GPU.md
deleted file mode 100644
index b5dfc4d1..00000000
--- a/documentation/CP-plugins-documentation/using_GPU.md
+++ /dev/null
@@ -1,34 +0,0 @@
-# Using a GPU with CellProfiler Plugins
-
-These CellProfiler modules currently support using a GPU: LIST MODULES
-
-## Tips for Setting Up Your GPU (if you have one)
-
-If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU. General instructions are available at this [link](https://pytorch.org/get-started/locally/).
-
-1. Your GPU should be visible in Device Manager under Display Adaptors. If your GPU isn't there, you likely need to install drivers.
- * [Here](https://www.nvidia.com/Download/Find.aspx) is where you can find NVIDIA GPU drivers if you need to install them.
-
-
-2. To test whether the GPU is configured correctly:
- * Run `python` on the command line (i.e., in Command Prompt or Terminal) to start an interactive session
- * Then run the following
- ```
- import torch
- torch.cuda.is_available()
- ```
- * If this returns `True`, you're all set
- * If this returns `False`, you likely need to install/reinstall torch. See [here](https://pytorch.org/get-started/locally/) for your exact command.
- * Exit the session with `exit()` then install torch if necessary
- ```
- pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
- ```
- If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index bce4bc70..946b9ab5 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -153,4 +153,5 @@ These are all the folders you need to copy over:
```
7. **Open and use CellProfiler.**
-When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing.
\ No newline at end of file
+When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing in the terminal window that opens with CellProfiler on Windows machines.
+This information is not available in Mac machines.
\ No newline at end of file
From f99b97e8c94b878f6e7a6059bd5f926f9901b41c Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 15:09:02 -0700
Subject: [PATCH 6/6] docs
---
documentation/CP-plugins-documentation/using_plugins.md | 5 ++++-
1 file changed, 4 insertions(+), 1 deletion(-)
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index 946b9ab5..bab82fe8 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -1,7 +1,10 @@
# Using plugins
-Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all plugins in the plugin folder that you have set.
+Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all useable plugins in the plugin folder that you have set.
The plugins will appear in the "Add Modules" panel like all standard modules and you can use the plugins as you would any other CellProfiler module.
+Typically, if a plugin's dependencies are not installed, it will not be visible in the "Add Modules" panel.
+If you cannot find any plugins in the "Add Modules" panel then you have not properly set the plugins path (see below).
+If you can find some but not all plugins in the "Add Modules" panel then the not-visible plugins have unmet dependencies.
Please note that, as CellProfiler-plugins are considered experimental, they may not be as well documented as standard modules and they may not have a window that shows on run.
Please report any installation issues or bugs related to plugins in the [CellProfiler-plugins repository](https://github.com/CellProfiler/CellProfiler-plugins) and not in the main CellProfiler repository.
-
-
-
----
-
-## Common errors
-
-1. My wheels are failing to build
-
- - If you get a message like "ERROR: Failed building wheel for pyzmq" this usually means that you do not have the Microsoft Visual Studio tools installed properly or the wrong version of pyzmq is being installed (e.g., you have a previous version already installed in your environment). See Step #1-2 above and ensure that you have "Desktop development with C++" selected under the install configuration options
-
-
-2. Java virtual machine cannot be found. This error might look something like `javabridge.jutil.JVMNotFoundError`
-
- - If you're getting errors about Java, it means that java is not being configured properly on your system.
- - Make sure you have installed The Java Development Kit 11. Note that newer versions of Java may not work.
- - Make sure you've added environment variables at the **System** level and not at the **User** level. You need to add both `JAVA_HOME` and `JDK_HOME` and both need to point to the folder that holds your jdk installation. Typically this path would look something like `C:\Program Files\Java\jdk-11.0.15.1` but it might be different on your machine depending on where you've installed Java.
- - If you have done these things and you still get this error, then try `pip install javabridge` (inside your activated cellprofiler-plugins conda environment) and then opening cellprofiler with `cellprofiler`
-
-
-3. Installing pyzmq failed
-
- You might get an error when trying to install pyzmq. Something like
- ```
- ERROR: Command errored out with exit status 1:
- ```
- And earlier in the traceback:
- ```
- Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
- Cython >= 0.20 is required to compile pyzmq from a development branch.
- Please install Cython or download a release package of pyzmq.
- ```
-
- To fix this, activate your environment and then `conda install cython`
-
- - You might also get an error like:
- ```
- AttributeError: 'MSVCCompiler' object has no attribute '_vcruntime_redist'
- ```
-
- Generally, this error means that you don't have the right Microsoft Visual Studio C compiler. You can try two things:
-
- 1. Look below in the code. Even if the wheel fails, pyzmq will still attempt to install. Look below for `Running setup.py install for pyzmq ... done` Also make sure you don't have any additional versions of pyzmq installed already.
- 2. You can try reinstalling Microsoft Visual C++ Build tools (#1-2). Versions 2008, 2015-2022 should work. Make sure you have the version of the redistributable compatible with your system processor (see step #2).
-
----
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
new file mode 100644
index 00000000..0020d62b
--- /dev/null
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -0,0 +1,64 @@
+# Using plugins
+
+Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all plugins in the plugin folder that you have set.
+The plugins will appear in the "Add Modules" panel like all standard modules and you can use the plugins as you would any other CellProfiler module.
+
+Please note that, as CellProfiler-plugins are considered experimental, they may not be as well documented as standard modules and they may not have a window that shows on run.
+Please report any installation issues or bugs related to plugins in the [CellProfiler-plugins repository](https://github.com/CellProfiler/CellProfiler-plugins) and not in the main CellProfiler repository.
+
+## Installation
+
+### Installing plugins without dependencies
+
+If your plugin doesn't require additional dependencies, all you need to do is
+
+1. **Install CellProfiler.**
+Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
+
+Or, you can install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
+
+2. **Clone the CellProfiler-plugins repository.**
+This will download all of the plugins in the CellProfiler-plugins repository.
+
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+Alternatively, if you have code for plugins that are not in the CellProfiler-plugins repository, you can place them in any folder that you'll be able to find again.
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler and re-open it.
+
+You are now ready to use any CellProfiler plugin that does not have additional dependencies.
+This is most CellProfiler plugins.
+
+### Installing plugins with dependencies
+
+1. **Install CellProfiler.**
+Install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
+
+2. **Clone the CellProfiler-plugins repository.**
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler and re-open it.
+
+4. **Install the dependencies for your plugin.**
+In your terminal, type the following, where FLAG is the flag specific to the module you would like to run, as noted in [Supported Plugins](supported_plugins.md).
+```bash
+cd CellProfiler-plugins
+pip install -e .[FLAG]
+```
+
+Please note that plugins that have separate install flags may have conflicting dependencies so we recommend making a separate python environment in which to run separate installations.
+(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
\ No newline at end of file
From 4c41db2f05cabb63f634e1ff9c07f459ca39dd72 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 14:03:04 -0700
Subject: [PATCH 2/6] deprecate declumpobjects, add info about unsupported
plugins
---
.../unsupported_plugins.md | 14 +++++++++++++-
.../CellProfiler4/declumpobjects.py | 0
2 files changed, 13 insertions(+), 1 deletion(-)
rename declumpobjects.py => unmaintained_plugins/CellProfiler4/declumpobjects.py (100%)
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index 7e0343da..4ca10a6c 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -8,5 +8,17 @@ We welcome requests for updating particular unsupported plugins, but please note
We welcome community contributed plugin updates.
Likewise, we welcome contributions of community developed plugins.
-Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team or community contributed.
+Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
+## Where are unsupported plugins?
+
+Unsupported plugins can be found in the `unmaintained_plugins` folder in the CellProfiler-plugins repository.
+Those plugins in the `CellProfiler2`, `CellProfiler3`, and `CellProfiler4` folders were, at one point, supported for those versions of CellProfiler.
+Those plugins in the `CellProfiler4_autoconverted` folder were automatically converted from Python2 to Python3 (to support the transition from Python2 in CellProfiler3 to Python3 in CellProfiler4) but were never fully supported and may or may not run.
+
+## What plugins are unsupported?
+
+We cannot provide comprehensive information about why we are not supporting a given plugin.
+Information about select plugins is as follows:
+
+**DeclumpObjects**: DeclumpObjects will split objects based on a seeded watershed method. Functionality from this module was [added into CellProfiler](https://github.com/CellProfiler/CellProfiler/pull/4397) in the Watershed module as of CellProfiler 4.2.0.
diff --git a/declumpobjects.py b/unmaintained_plugins/CellProfiler4/declumpobjects.py
similarity index 100%
rename from declumpobjects.py
rename to unmaintained_plugins/CellProfiler4/declumpobjects.py
From d47666e9fcefc1f98227e02aa4692ed83a022e0c Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 14:20:25 -0700
Subject: [PATCH 3/6] more documentation
---
.../contributing_plugins.md | 17 +++-
.../CP-plugins-documentation/overview.md | 11 +++
.../supported_plugins.md | 77 +++++--------------
.../CP-plugins-documentation/using_GPU.md | 8 ++
4 files changed, 51 insertions(+), 62 deletions(-)
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index 204ec8f8..c3ae78be 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -1,6 +1,8 @@
-## Contributing New Plugins
+# Contributing New Plugins
-Refer to CellProfiler repository and within cellprofiler/modules/plugins you can find two different templates to use for creating your own plugin.
+Within our CellProfiler wiki, you can find an [orientation to CellProfiler code](https://github.com/CellProfiler/CellProfiler/wiki/Orientation-to-CellProfiler-code).
+
+In the CellProfiler repository, within [cellprofiler/modules/plugins](https://github.com/CellProfiler/CellProfiler/tree/master/cellprofiler/modules/plugins) you can find two different templates to use for creating your own plugin.
`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
@@ -8,10 +10,17 @@ In you plugin, you must include:
In your plugin, we appreciate if you also include:
- display functionality
+- extensive module documentation
+- references and citation information in your module documentation
+Please create a Pull Request to the CellProfiler-plugins repository to submit your plugin for inclusion in the repository.
-In your PR, you must include:
+In your PR, you must:
- add your plugin to the [supported_plugins](supported_plugins.md) documentation page
In your PR, we appreciate if you also include:
-- unit tests for your plugin
\ No newline at end of file
+- unit tests for your plugin
+
+## Having your plugin cited
+
+While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/overview.md b/documentation/CP-plugins-documentation/overview.md
index 5ece4c8d..1616e1a3 100644
--- a/documentation/CP-plugins-documentation/overview.md
+++ b/documentation/CP-plugins-documentation/overview.md
@@ -9,6 +9,17 @@ A module may be in CellProfiler-plugins instead of CellProfiler itself because:
- it requires extra libraries or other dependencies we are unable or unwilling to require for CellProfiler
- it has been contributed by a community member
+## How do I use CellProfiler Plugins?
+
+See our [Using Plugins](using_plugins.md) page for information on how to install and use plugins.
+
+## What plugins are available?
+
+See our [Supported Plugins](supported_plugins.md) page for information on all currently supported plugins.
+
+## How do I contribute a plugin?
+
+See our [Contributing Plugins](contributing_plugins.md) page for information on contributing a plugin to CellPainting-plugins.
## Who made this?
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index 5850822a..deb60d61 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -3,61 +3,22 @@
Below is a brief overview of our currently supported plugins.
For details about using any particular plugin, please read the module documentation inside the plugin in CellProfiler.
-## CalculateMoments
-CalculateMoments extracts moments statistics from a given distribution of pixel values.
-
-## CallBarcodes
-CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles.
-It is used for optical sequencing by synthesis (SBS).
-
-## CompensateColors
-CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through.
-It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched.
-
-## DeclumpObjects
-DeclumpObjects will split objects based on a seeded watershed method.
-
-## DistanceTransform
-DistanceTransform computes the distance transform of a binary image.
-The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1.
-
-## EnhancedMeasureTexture
-EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler.
-
-## HistogramEqualization
-HistogramEqualization increases the global contrast of a low-contrast image or volume.
-Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct.
-This module can perform either global or local histogram equalization.
-
-## HistogramMatching
-HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image.
-It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image.
-It allows you to choose which frame to use as the reference.
-
-## PixelShuffle
-PixelShuffle takes the intensity of each pixel in an image and it randomly shuffles its position.
-
-## Predict
-Predict allows you to use an ilastik pixel classifier to generate a probability image.
-CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage).
-
-## RunCellpose
-RunCellpose allows you to run Cellpose within CellProfiler.
-Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells.
-You can use pre-trained Cellpose models or your custom model with this plugin.
-You can use a GPU with this module to dramatically increase your speed/efficiency.
-
-## RunImageJScript
-
-## RunOmnipose
-
-## RunStarDist
-RunStarDist allows you to run StarDist within CellProfiler.
-StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells.
-You can use pre-trained StarDist models or your custom model with this plugin.
-You can use a GPU with this module to dramatically increase your speed/efficiency.
-RunStarDist is generally faster than RunCellpose.
-
-## VarianceTransform
-This module allows you to calculate the variance of an image, using a determined window size.
-It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image.
\ No newline at end of file
+Most plugins will run without any special installation of either CellProfiler or the plugins.
+See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin use as well as for installation information for those plugins that do require installation of dependencies.
+
+| Plugin | Description | Requires installation of dependencies? | Install flag |
+|--------|-------------|----------------------------------------|--------------|
+| CalculateMoments | CalculateMoments extracts moments statistics from a given distribution of pixel values. | No | |
+| CallBarcodes | CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles. It is used for optical sequencing by synthesis (SBS). | No | |
+| CompensateColors | CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through. It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched. | No | |
+| DistanceTransform | DistanceTransform computes the distance transform of a binary image. The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1. | No | |
+| EnhancedMeasureTexture| EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler. | No | |
+| HistogramEqualization | HistogramEqualization increases the global contrast of a low-contrast image or volume. Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct. This module can perform either global or local histogram equalization. | No | |
+| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
+| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
+| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | Yes | XXXXX |
+| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
+| RunImageJScript | | Yes | XXXXX |
+| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
+| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
+| VarianceTransform | This module allows you to calculate the variance of an image, using a determined window size. It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image. | No | |
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/using_GPU.md b/documentation/CP-plugins-documentation/using_GPU.md
index 945d869b..b5dfc4d1 100644
--- a/documentation/CP-plugins-documentation/using_GPU.md
+++ b/documentation/CP-plugins-documentation/using_GPU.md
@@ -24,3 +24,11 @@ If you want to use a GPU to run the model (this is recommended for speed), you'l
pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
```
If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
+
+
+ **NOTE**: You might get a warning like this:
+ ```
+ W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
+ 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
+ ```
+ If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
From 12c52c77ec80879b399743160d6accbfe2049d26 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 12:35:02 -0700
Subject: [PATCH 4/6] more docs
---
.../contributing_plugins.md | 3 +-
.../supported_plugins.md | 4 +-
.../unsupported_plugins.md | 4 +-
.../CP-plugins-documentation/using_plugins.md | 106 ++++++++++++++++--
.../CP-plugins-documentation/versions.md | 49 +-------
5 files changed, 104 insertions(+), 62 deletions(-)
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index c3ae78be..f0084668 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -6,8 +6,6 @@ In the CellProfiler repository, within [cellprofiler/modules/plugins](https://gi
`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
-In you plugin, you must include:
-
In your plugin, we appreciate if you also include:
- display functionality
- extensive module documentation
@@ -20,6 +18,7 @@ In your PR, you must:
In your PR, we appreciate if you also include:
- unit tests for your plugin
+- customized installation in setup.py, if your plugin has dependencies
## Having your plugin cited
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index deb60d61..a17866c0 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -16,9 +16,9 @@ See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin
| HistogramEqualization | HistogramEqualization increases the global contrast of a low-contrast image or volume. Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct. This module can perform either global or local histogram equalization. | No | |
| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
-| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | Yes | XXXXX |
+| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | No | |
| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
-| RunImageJScript | | Yes | XXXXX |
+| RunImageJScript | RunImageJScript allows you to run any supported ImageJ script directly within CellProfiler. It is significantly more performant than RunImageJMacro, and is also less likely to leave behind temporary files. | Yes | XXXXX |
| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
| VarianceTransform | This module allows you to calculate the variance of an image, using a determined window size. It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image. | No | |
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index 4ca10a6c..b7cf3fe8 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -5,11 +5,9 @@ Unsupported plugins are primarily unsupported for either of two reasons:
- Their functions have been integrated into the current version of CellProfiler and therefore a plugin is no longer necessary
We welcome requests for updating particular unsupported plugins, but please note that we have limited bandwidth for working on plugins and may be unable to complete the update.
+Additionally, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
We welcome community contributed plugin updates.
-Likewise, we welcome contributions of community developed plugins.
-Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
-
## Where are unsupported plugins?
Unsupported plugins can be found in the `unmaintained_plugins` folder in the CellProfiler-plugins repository.
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index 0020d62b..bce4bc70 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -8,9 +8,16 @@ Please report any installation issues or bugs related to plugins in the [CellPro
## Installation
-### Installing plugins without dependencies
+If the plugin you would like to use does not have any additional dependencies outside of those required for running CellProfiler (this is most plugins), using plugins is very simple.
+See [Installing plugins without dependencies](#installing-plugins-without-dependencies).
+
+If the plugin you would like to use has dependencies, you have two separate options for installation.
+The first option requires building CellProfiler from source, but plugin installation is simpler.
+See [Installing plugins with dependencies, using CellProfiler from source](#installing-plugins-with-dependencies-using-cellprofiler-from-source).
+The second option allows you to use pre-built CellProfiler, but plugin installation is more complex.
+See [Installing plugins with dependencies, using pre-built CellProfiler](#installing-plugins-with-dependencies-using-pre-built-cellprofiler).
-If your plugin doesn't require additional dependencies, all you need to do is
+### Installing plugins without dependencies
1. **Install CellProfiler.**
Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
@@ -31,12 +38,12 @@ Alternatively, if you have code for plugins that are not in the CellProfiler-plu
- Open CellProfiler.
- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
- Select `Save` at the bottom of the Preferences window
-- Close CellProfiler and re-open it.
+- Close CellProfiler and re-open it
You are now ready to use any CellProfiler plugin that does not have additional dependencies.
This is most CellProfiler plugins.
-### Installing plugins with dependencies
+### Installing plugins with dependencies, using CellProfiler from source
1. **Install CellProfiler.**
Install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
@@ -49,9 +56,9 @@ git clone https://github.com/CellProfiler/CellProfiler-plugins.git
3. **Set the plugins path in CellProfiler.**
- Open CellProfiler.
-- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned.
- Select `Save` at the bottom of the Preferences window
-- Close CellProfiler and re-open it.
+- Close CellProfiler
4. **Install the dependencies for your plugin.**
In your terminal, type the following, where FLAG is the flag specific to the module you would like to run, as noted in [Supported Plugins](supported_plugins.md).
@@ -59,6 +66,91 @@ In your terminal, type the following, where FLAG is the flag specific to the mod
cd CellProfiler-plugins
pip install -e .[FLAG]
```
+e.g. To install Cellpose the pip install command would be `pip install -e .[cellpose]`
+
+5. **Open and use CellProfiler.**
Please note that plugins that have separate install flags may have conflicting dependencies so we recommend making a separate python environment in which to run separate installations.
-(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
\ No newline at end of file
+(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
+
+### Installing plugins with dependencies, using pre-built CellProfiler
+
+1. **Install CellProfiler.**
+Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
+
+2. **Clone the CellProfiler-plugins repository.**
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned.
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler
+
+4. **Identify the Python dependencies of the plugins you want to use.**
+You can find this information for most plugins by looking in `setup.py`.
+```{admonition} e.g. using RunCellpose plugin
+If you would like to install the RunCellpose plugin, in `setup.py` you can see that `cellpose_deps = ["cellpose>=1.0.2"]` so the only dependency is `cellpose`.
+```
+
+Alternatively, you can find this information directly in the plugin code itself by looking at what is imported at the beginning of the plugin.
+Note that you will want to compare any imports to what is already required by CellProfiler with that caveat that dependencies often have dependencies.
+```{admonition} e.g. using RunImageJScript plugin
+If you would like to install the RunImageJScript plugin, you'll notice a number of imports in the beginning of the plugin code.
+You can ignore all those from `cellprofiler_core` as they are required for CellProfiler installation.
+You can ignore all those that are native to Python such as `sys`, `time`, and `threading`.
+A comparison with CellProfiler dependencies will show you that `skimage` is already installed (you can see this in CellProfiler's [environment.yml](https://github.com/CellProfiler/CellProfiler/blob/master/environment.yml) which lists `scikit-image` as a `cellprofiler-core` dependency or in CellProfiler's [setup.py](https://github.com/CellProfiler/CellProfiler/blob/master/setup.py) which lists `scikit-image` as an installation requirement).
+What remains is `pyimagej`
+```
+
+5. **Create a conda environment and install requirements in it**
+Create a conda (or other virtual) environment with a Python version matching that of CellProfiler.
+CellProfiler 4 uses Python 3.8 (which you can see in CellProfiler's [setup.py](https://github.com/CellProfiler/CellProfiler/blob/master/setup.py)).
+Activate the environment and install the plugin requirement/s into it.
+
+In your terminal, type
+```
+# Create a conda environment
+conda create --name ENV_NAME python=3.8
+# Activate the conda environment
+conda activate ENV_NAME
+# Install the plugin requirement/s
+pip install REQUIREMENT
+```
+```{admonition} e.g. using RunImageJScript plugin
+conda create --name cp-ij python=3.8
+conda activate cp-ij
+pip install pyimagej
+```
+
+6. **Copy plugin requirements into CellProfiler installation**
+Find the folder that you installed into in the previous step (where to find these libraries in your local environment) by entering `pip show REQUIREMENT` into your terminal e.g. `pip show pyimagej`.
+In the information it returns, under `Location` you will find a path that will look something like `/Users/eweisbar/mambaforge/envs/cp-ij/lib/python3.8/site-packages`.
+
+Find the folder in which you have installed CellProfiler.
+On Windows this is likely to be `C:\Program Files\CellProfiler` and on Mac it is likely to be `/Applications/CellProfiler.app/Contents/MacOS/`.
+In your terminal, if you `ls` (for Mac) or `dir` (for Windows) that folder (e.g. `ls /Applications/CellProfiler.app/Contents/MacOS/`), you should see a list of all the dependencies that were packaged with CellProfiler.
+
+Copy the folders and their corresponding `.dist-info` folders (e.g. `pytz` and `pytz-2023.3.dist-info`) for any dependencies that were installed with the installation of `pyimagej` that are not already in the CellProfiler folder.
+
+```{admonition} e.g. using RunImageJScript plugin
+These are all the folders you need to copy over:
+`pytz`
+`pytz-2023.3.dist-info`
+`xarray`
+`xarray-2023.1.0.dist-info`
+`imglyb`
+`imglyb-2.1.0.dist-info`
+`jgo`
+`jgo-1.0.5.dist-info`
+`jpype`
+`_jpype.cpython-38-darwin.so`
+`imagej`
+`pyimagej-1.4.1.dist-info`
+```
+
+7. **Open and use CellProfiler.**
+When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/versions.md b/documentation/CP-plugins-documentation/versions.md
index 3755e75b..0f8db604 100644
--- a/documentation/CP-plugins-documentation/versions.md
+++ b/documentation/CP-plugins-documentation/versions.md
@@ -1,55 +1,8 @@
# Versions
-The most current release can always be found on DockerHub at `cellprofiler/distributed-cellprofiler`.
-Current version is 2.0.0.
-Our current tag system is dcpversion_CellProfilerversion, e.g. 1.2.0_3.1.8
-
-Previous release versions can be accessed at `bethcimini/distributed-cellprofiler:versionnumber`
+Current version is
---
# Version History
-## 2.1.0 (forthcoming)
-* Increase support for using plugins
-* Improve role handling and allow file download/upload from/to S3 buckets in different accounts
-* Improve file download to S3FS can be completely bypassed
-* Name EBS volumes
-
-## 2.0.0rc2 - Released 20201110
-* Add optional ability to download files to EBS rather than reading from S3 (helpful for pipelines that access many files/image sets)
-
-## 2.0.0rc1 - Released 20201105
-
-* Remove requirement for boto and fabric, using only boto3
-* Add support for Python 3 and CellProfiler 4
-* Move cluster size, machine type, and machine price to the config file from the fleet file, eliminating mismatches between the two
-* Add the ability to filter only files with certain names when running CHECK_IF_DONE
-* Don't cancel a fleet over capacity errors
-* Add "cheapest" mode to the monitor, allowing you to run more cheaply (at possible expense of running more slowly)
-
-## 1.2.2 - Released 20201103
-
-* Allow pipelines using batch files to also designate an input output_top_directory
-* Add support for multiple LaunchData specifications
-* Add CellProfiler-plugins
-* Additional way to create job submissions with run_batch_general.py
-
-## 1.2.1 - Released 20200109, Updated through 20191002
-
-* Allow monitor to downscale machines when number of jobs < number of machines
-* Add a parameter to discount files when running CHECK_IF_DONE checks if less than a certain size
-
-## 1.2.0 - Released 20181108, Updated through 20200109
-
-* Improved compatibility with CellProfiler 2 and 3
-* Better handling of logging when using output_structure
-
-## 1.1.0 - Released 20170217, Updated 20170221 (bugfixes) - 20181018
-
-* Changes in this release:
-
- * Added the `output_structure` variable to the job file, which allows you to structure the output folders created by DCP (ie `Plate/Well-Site` rather than `Plate-Well-Site`). Job files lacking this variable will still default to the previous settings (hyphenating all Metadata items in order they are presented in the Metadata grouping).
- * Added support for creating the list of groups via `cellprofiler --print-groups`- see [this issue](https://github.com/CellProfiler/Distributed-CellProfiler/issues/52) for example and discussion. Groups listed in this way MUST use the `output_structure` variable to state their desired folder structure or an error will be returned.
-
-## 1.0.0 - Version as of 20170213
From fffb5566c3b4d2892c75c30b14469e66e94bca6a Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 15:04:09 -0700
Subject: [PATCH 5/6] more doc changes
---
.../CP-plugins-documentation/RunCellPose.md | 34 +++++++++++++++++++
.../contributing_plugins.md | 9 ++++-
.../supported_plugins.md | 5 ++-
.../unsupported_plugins.md | 1 +
.../CP-plugins-documentation/using_GPU.md | 34 -------------------
.../CP-plugins-documentation/using_plugins.md | 3 +-
6 files changed, 49 insertions(+), 37 deletions(-)
create mode 100644 documentation/CP-plugins-documentation/RunCellPose.md
delete mode 100644 documentation/CP-plugins-documentation/using_GPU.md
diff --git a/documentation/CP-plugins-documentation/RunCellPose.md b/documentation/CP-plugins-documentation/RunCellPose.md
new file mode 100644
index 00000000..168fa7cc
--- /dev/null
+++ b/documentation/CP-plugins-documentation/RunCellPose.md
@@ -0,0 +1,34 @@
+# RunCellPose
+
+## Using RunCellPose with a GPU
+
+If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU.
+General instructions are available at this [link](https://pytorch.org/get-started/locally/).
+
+1. Your GPU should be visible in Device Manager under Display Adaptors.
+If your GPU isn't there, you likely need to install drivers.
+[Here](https://www.nvidia.com/Download/Find.aspx) is where you can find NVIDIA GPU drivers if you need to install them.
+
+
+2. To test whether the GPU is configured correctly:
+ * Run `python` on the command line (i.e., in Command Prompt or Terminal) to start an interactive session
+ * Then run the following
+ ```
+ import torch
+ torch.cuda.is_available()
+ ```
+ * If this returns `True`, you're all set
+ * If this returns `False`, you likely need to install/reinstall torch. See [here](https://pytorch.org/get-started/locally/) for your exact command.
+ * Exit the session with `exit()` then install torch if necessary
+ ```
+ pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
+ ```
+ If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
+
+
+**NOTE**: You might get a warning like this:
+```
+W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
+2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
+```
+If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index f0084668..8907b250 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -20,6 +20,13 @@ In your PR, we appreciate if you also include:
- unit tests for your plugin
- customized installation in setup.py, if your plugin has dependencies
+## Contributing bug fixes or updating deprecated plugins
+
+Please create a Pull Request to the CellProfiler-plugins repository to submit a bug fix or plugin un-deprecation.
+If you would like to update a currently deprecated plugin, we encourage you to read closed Issues in GitHub relevant to the plugin as they may contain helpful information about bugs present at the time of deprecation.
+
## Having your plugin cited
-While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
\ No newline at end of file
+While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
+
+See [Citing Plugins](citing.md) for information on citing CellProfiler Plugins.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index a17866c0..5ccdda93 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -6,6 +6,9 @@ For details about using any particular plugin, please read the module documentat
Most plugins will run without any special installation of either CellProfiler or the plugins.
See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin use as well as for installation information for those plugins that do require installation of dependencies.
+Most plugin documentation can be found within the plugin itself and can be accessed through CellProfiler help.
+Those plugins that do have extra documentation contain links below.
+
| Plugin | Description | Requires installation of dependencies? | Install flag |
|--------|-------------|----------------------------------------|--------------|
| CalculateMoments | CalculateMoments extracts moments statistics from a given distribution of pixel values. | No | |
@@ -17,7 +20,7 @@ See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin
| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | No | |
-| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
+| [RunCellpose](RunCellPose.md) | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
| RunImageJScript | RunImageJScript allows you to run any supported ImageJ script directly within CellProfiler. It is significantly more performant than RunImageJMacro, and is also less likely to leave behind temporary files. | Yes | XXXXX |
| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index b7cf3fe8..5e277b61 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -19,4 +19,5 @@ Those plugins in the `CellProfiler4_autoconverted` folder were automatically con
We cannot provide comprehensive information about why we are not supporting a given plugin.
Information about select plugins is as follows:
+**ClassifyPixelsUNET**: ClassifyPixelsUNET is a pixel classifier for background/object edge/object body. As far as we are aware, other deep learning based plugins that we do currently support (such as RunCellpose) work better.
**DeclumpObjects**: DeclumpObjects will split objects based on a seeded watershed method. Functionality from this module was [added into CellProfiler](https://github.com/CellProfiler/CellProfiler/pull/4397) in the Watershed module as of CellProfiler 4.2.0.
diff --git a/documentation/CP-plugins-documentation/using_GPU.md b/documentation/CP-plugins-documentation/using_GPU.md
deleted file mode 100644
index b5dfc4d1..00000000
--- a/documentation/CP-plugins-documentation/using_GPU.md
+++ /dev/null
@@ -1,34 +0,0 @@
-# Using a GPU with CellProfiler Plugins
-
-These CellProfiler modules currently support using a GPU: LIST MODULES
-
-## Tips for Setting Up Your GPU (if you have one)
-
-If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU. General instructions are available at this [link](https://pytorch.org/get-started/locally/).
-
-1. Your GPU should be visible in Device Manager under Display Adaptors. If your GPU isn't there, you likely need to install drivers.
- * [Here](https://www.nvidia.com/Download/Find.aspx) is where you can find NVIDIA GPU drivers if you need to install them.
-
-
-2. To test whether the GPU is configured correctly:
- * Run `python` on the command line (i.e., in Command Prompt or Terminal) to start an interactive session
- * Then run the following
- ```
- import torch
- torch.cuda.is_available()
- ```
- * If this returns `True`, you're all set
- * If this returns `False`, you likely need to install/reinstall torch. See [here](https://pytorch.org/get-started/locally/) for your exact command.
- * Exit the session with `exit()` then install torch if necessary
- ```
- pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
- ```
- If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index bce4bc70..946b9ab5 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -153,4 +153,5 @@ These are all the folders you need to copy over:
```
7. **Open and use CellProfiler.**
-When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing.
\ No newline at end of file
+When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing in the terminal window that opens with CellProfiler on Windows machines.
+This information is not available in Mac machines.
\ No newline at end of file
From f99b97e8c94b878f6e7a6059bd5f926f9901b41c Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 15:09:02 -0700
Subject: [PATCH 6/6] docs
---
documentation/CP-plugins-documentation/using_plugins.md | 5 ++++-
1 file changed, 4 insertions(+), 1 deletion(-)
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index 946b9ab5..bab82fe8 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -1,7 +1,10 @@
# Using plugins
-Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all plugins in the plugin folder that you have set.
+Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all useable plugins in the plugin folder that you have set.
The plugins will appear in the "Add Modules" panel like all standard modules and you can use the plugins as you would any other CellProfiler module.
+Typically, if a plugin's dependencies are not installed, it will not be visible in the "Add Modules" panel.
+If you cannot find any plugins in the "Add Modules" panel then you have not properly set the plugins path (see below).
+If you can find some but not all plugins in the "Add Modules" panel then the not-visible plugins have unmet dependencies.
Please note that, as CellProfiler-plugins are considered experimental, they may not be as well documented as standard modules and they may not have a window that shows on run.
Please report any installation issues or bugs related to plugins in the [CellProfiler-plugins repository](https://github.com/CellProfiler/CellProfiler-plugins) and not in the main CellProfiler repository.
 -
-
-
- **NOTE**: For M1/M2 Macs, which use the ARM 64-bit architecture, use [this link](https://www.azul.com/downloads/?version=java-11-lts&os=macos&architecture=arm-64-bit&package=jdk) to get Java 11.
-
-
-
-
-
- **NOTE**: For M1/M2 Macs, which use the ARM 64-bit architecture, use [this link](https://www.azul.com/downloads/?version=java-11-lts&os=macos&architecture=arm-64-bit&package=jdk) to get Java 11.
-
-  -
-
- After you download the file, open the installer and proceed through the steps to install Java 11.
-
-
-
-
-2. **Download and install or update conda**
-
- For beginners, we recommend you use Anaconda Navigator since it is more beginner-friendly, but you can also use miniconda. [Download Anaconda](https://www.anaconda.com/products/distribution) from the website and install.
-
- **NOTE**: if you already have conda, you can get the command to update conda by typing `conda update` on your command line. The command will generally look like:
-
- `conda update --prefix /Users/USERNAME/opt/anaconda3 anaconda`
-
-
-
-3. **Clone the CellProfiler-plugins Repo**
-
- Download a copy of (aka "clone") the CellProfiler-plugins repo from [here](https://github.com/CellProfiler/CellProfiler-plugins.git). You can download the whole repo by cloning it with git or simply clicking the green **Code** button on [the repo page](https://github.com/CellProfiler/CellProfiler-plugins.git) and selecting **Download ZIP** (see below).
-
-
-
-
- After you download the file, open the installer and proceed through the steps to install Java 11.
-
-
-
-
-2. **Download and install or update conda**
-
- For beginners, we recommend you use Anaconda Navigator since it is more beginner-friendly, but you can also use miniconda. [Download Anaconda](https://www.anaconda.com/products/distribution) from the website and install.
-
- **NOTE**: if you already have conda, you can get the command to update conda by typing `conda update` on your command line. The command will generally look like:
-
- `conda update --prefix /Users/USERNAME/opt/anaconda3 anaconda`
-
-
-
-3. **Clone the CellProfiler-plugins Repo**
-
- Download a copy of (aka "clone") the CellProfiler-plugins repo from [here](https://github.com/CellProfiler/CellProfiler-plugins.git). You can download the whole repo by cloning it with git or simply clicking the green **Code** button on [the repo page](https://github.com/CellProfiler/CellProfiler-plugins.git) and selecting **Download ZIP** (see below).
-
-  -
- You can also use git or GitHub Desktop to clone the repo if you prefer.
-
-
-
-4. **Create the environment from the .yml file**
-
- Open Anaconda Navigator and select the **Environments** tab on the left. We recommend you create the environment from the command line. To do this, Select the play button next to your base (root) environment and select **Open Terminal**:
-
-
-
- You can also use git or GitHub Desktop to clone the repo if you prefer.
-
-
-
-4. **Create the environment from the .yml file**
-
- Open Anaconda Navigator and select the **Environments** tab on the left. We recommend you create the environment from the command line. To do this, Select the play button next to your base (root) environment and select **Open Terminal**:
-
-  -
- A black box should pop up with a blinking cursor. This is your terminal. You now need to navigate to where the **cellprofiler_plugins_mac.yml** file is inside of the CellProfiler-plugins folder you downloaded in the last step. This file is in the `Instructions` subfolder. Here is how we recommend you do this:
- 1) In Finder, open the folder you downloaded in the previous step (usually called "CellProfiler-plugins-master")
- 2) There should be a folder called **Instructions**. Right click or ctrl+click that folder. Then hold down the option key (or Alt) on your keyboard. An option to **Copy "Instructions" as Pathname** should appear. Select this option. (see below)
-
-
-
- A black box should pop up with a blinking cursor. This is your terminal. You now need to navigate to where the **cellprofiler_plugins_mac.yml** file is inside of the CellProfiler-plugins folder you downloaded in the last step. This file is in the `Instructions` subfolder. Here is how we recommend you do this:
- 1) In Finder, open the folder you downloaded in the previous step (usually called "CellProfiler-plugins-master")
- 2) There should be a folder called **Instructions**. Right click or ctrl+click that folder. Then hold down the option key (or Alt) on your keyboard. An option to **Copy "Instructions" as Pathname** should appear. Select this option. (see below)
-
-  -
-
- 3) Go back to your terminal and type `cd PATH_TO_FOLDER` where `PATH_TO_FOLDER` is the address you copied in the previous step. Press Enter.
-
-
-
- Now that you're in the right place, copy and paste this command into the terminal and press Enter.
- ```
- conda env create -f CellProfiler_plugins_mac.yml
- ```
-
-
-
- 5. **Activate your environment**
-
- In your terminal, enter `conda activate CellProfiler_plugins` to activate your environment
-
-
-
-
- 6. **Verify that cellprofiler is installed correctly by running it from the command line.**
-
- In your terminal, type in `pythonw -m cellprofiler` and hit Enter. this will open CellProfiler or will give you an error message.
-
-
-
- 7. **Install other packages for other plugins**
-
- Certain packages cannot easily installed when importing the environment.
-
- For StarDist: In terminal with your environment activated, enter:
- ```
- pip install stardist csbdeep --no-deps
- ```
-
-
- If you would like to use the omnipose models in cellpose, ensure you have cellpose 1.0.2 (you should by default if you've used the CellProfiler_plugins_mac.yml file to generate the environment) and enter on the command line (in your activated environment):
-
- ```
- pip install omnipose
- ```
-
-
-
-
- 8. **Connect CellProfiler and the plugins repo**
-
- With your environment active, type `pythonw -m cellprofiler` in terminal to open CellProfiler if it is not open already.
-
- * In CellProfiler, go to **File** then **Preferences...**
- * Scroll down and look for "CellProfiler Plugins Directory" on the left.
- * Select the **Browse** button and choose the folder where you extracted the CellProfiler plugins files. It is probably called "CellProfiler-plugins-master" unless you have renamed it.
- * Select **Save** at the bottom of the Preferences window
- * Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` on the command line
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
-
-
-
- 9. **Verify that the installation worked**
-
- Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
- In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
- 
-
- ---
-
-
-## Common errors
-
-1. My wheels are failing to build
-
-- If you get a message like "ERROR: Failed building wheel for pyzmq" this usually means that you do not have pyzmq installed. Try to reinstall pyzmq.
-
-2. Java virtual machine cannot be found
-
-- If you're getting errors about Java, it means that java is not being configured properly on your system.
-- Make sure you have installed The Java Development Kit 11 [here](https://adoptopenjdk.net/). Note that newer versions may not work.
-- Make sure you've added environment variables at the **System** level and not at the **User** level.
-```
-brew install java
-
-# For the system Java wrappers to find this JDK, symlink it with
-sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
-
-# Set version in zshrc
-echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
-source ~/.zshrc
-```
-
-3. Installing pyzmq failed
-
-- You might get an error when trying to install pyzmq. Something like
-```
-ERROR: Command errored out with exit status 1:
-```
-And earlier in the traceback:
-```
-Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
- Cython >= 0.20 is required to compile pyzmq from a development branch.
- Please install Cython or download a release package of pyzmq.
-```
-
- To fix this, `conda install cython`
-
-
-## Installation instructions for CellProfiler, CellProfiler plugins and CellPose in a conda environment on Apple silicon (eg M1).
-
-1. **Install brew**
- ```
- /bin/bash -c "$(curl -fsSL https://raw.githubusercontent.com/Homebrew/install/HEAD/install.sh)"
- ```
-
-2. **Update brew**
- ```
- brew update
- ```
-
-3. **Install Java**
- ```
- brew install java
- ```
-
-4. **For the system Java wrappers to find this JDK and symlink it**
- ```
- sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
- ```
-
-5. **Set version in .zshrc**
- ```
- echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
- source ~/.zshrc
- ```
-
-6. **Brew install package requirements**
- ```
- brew install freetype mysql git
- ```
-
-7. **Direct to brew openssl**
- ```
- export LDFLAGS="-L$(brew --prefix)/opt/openssl/lib"
- ```
-
-8. **Install hdf5**
- ```
- brew install hdf5@1.12
- ```
-
-9. **Make sure to get the version directory correct for the version installed. Find with `ls /opt/homebrew/Cellar/hdf5/`**
- ```
- export HDF5_DIR=/opt/homebrew/Cellar/hdf5/1.12.1_1/
- ```
-10. **Create a folder and download cellprofiler-plugins**
-
- ```
- mkdir cp_plugins
- cd cp_plugins
- git clone https://github.com/CellProfiler/CellProfiler-plugins
- ```
-
-11. **Download and install miniconda**
-
- ```
- wget https://repo.anaconda.com/miniconda/Miniconda3-latest-MacOSX-arm64.sh -O ~/miniconda.sh
- ```
-
-
-12. **Create a conda `.yml` file**
- In a text editor, paste the following test and save it as `cellprofiler_plugins_macM1.yml`:
- ```
- name: cpcellpose
-
- channels:
- - conda-forge
- - anaconda
- - bioconda
- - defaults
- - apple
-
- dependencies:
- - python=3.8
- - pip
- - h5py
- - mysqlclient
- - imagecodecs
- - python.app
- - pandas
- - pip:
- - cellpose
- - attrdict
- - sip==5.5.0
- - boto3>=1.12.28
- - centrosome==1.2.1
- - docutils==0.15.2
- - h5py~=3.6.0
- - imageio>=2.5
- - inflect>=2.1
- - Jinja2>=2.11.2
- - joblib>=0.13
- - mahotas>=1.4
- - matplotlib==3.1.3
- - mysqlclient==1.4.6
- - numpy>=1.20.1
- - Pillow>=7.1.0
- - prokaryote==2.4.4
- - python-bioformats==4.0.6
- - python-javabridge==4.0.3
- - pyzmq~=22.3
- - sentry-sdk==0.18.0
- - requests>=2.22
- - scikit-image>=0.17.2
- - scikit-learn>=0.20
- - scipy>=1.4.1
- - six
- - tifffile<2022.4.22
- - wxPython==4.2.0
- ```
-
-
-13. **Create a conda environment using the cellprofiler_plugins_macM1.yml file**
-
- At this stage, your folder/file structure should look like this:
-
- ```
- ├── cp_plugins
- ├── CellProfiler-plugins
- └── cellprofiler_plugins_macM1.yml
- ```
-
- In the terminal, make sure you are in the `cp_plugins` folder mentioned above (run `cd cp_plugins` to get there).
-
- ```
- conda env create -n cp_plugins --file cellprofiler_plugins_macM1.yml
- ```
-
-14. **Activate the conda environment**
-
- ```
- conda activate cp_plugins
- ```
-
-15. **Install CellProfiler and CellProfiler-core**
- ```
- pip install cellprofiler-core
-
- # We install CellProfiler with no dependencies here since we installed them with the conda yml file
- # This allows us to have some dependencies for M1 that are not the default for CellProfiler (eg wxPython)
- git clone https://github.com/CellProfiler/CellProfiler
-
- pip install --no-deps cellprofiler
- ```
-
-16. **Open CellProfiler**
- ```
- pythonw -m cellprofiler
- ```
-
-17. **Connect CellProfiler with the plugins folder**
-
- 1. In CellProfiler, go to `File` then `Preferences...`
- 2. Scroll down and look for `CellProfiler Plugins Directory` on the left.
- 3. Select the Browse button and choose the folder where you extracted the CellProfiler plugins files. It is probably called `CellProfiler-plugins` unless you have renamed it.
- 4. Select Save at the bottom of the Preferences window
- 5. Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` in the command line
-
-
-### Resolving dependencies conflicts
-
- There can be some strange dependency conflicts that can arise with python-javabridge. In the terminal with your environment activate, enter:
- ```
- pip uninstall -y centrosome python-javabridge
- pip install --no-cache-dir --no-deps --no-build-isolation python-javabridge centrosome
- pip uninstall matplotlib -y
- pip install matplotlib==3.2
- pip uninstall mahotas -y
- pip install mahotas
- ```
-
- The latest version of CellProfiler available on github removes the python-javabridge requirement and instead uses ScyJava.
-
-### Test your installation
-
-Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
-In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
diff --git a/documentation/CP-plugins-documentation/Windows_installation.md b/documentation/CP-plugins-documentation/Windows_installation.md
deleted file mode 100644
index 60bb701c..00000000
--- a/documentation/CP-plugins-documentation/Windows_installation.md
+++ /dev/null
@@ -1,208 +0,0 @@
-# How to install CellProfiler from source with all plugins on Windows
-
-Outlined here is the manual process for installing CellProfiler and plugins from source. Please close all open programs and save your work before starting this, as your computer may require a restart. Note that some of these download steps can take a while; the first 3 steps (as well as downloading Anaconda) can all be done simultaneously to save you time. Also note that for steps like changing the system-level environment variables, you may need admin permissions on your computer.
-
-**NEW:** We now have an automated install available for this workflow (requires administrator permissions on your computer) that will perform these steps for you. To get the necessary files and see instructions for the automatic install, look [here](https://github.com/CellProfiler/CellProfiler-plugins/tree/master/Instructions/Windows_batch_file).
-
-1. **Download and install [Microsoft Visual Studio C++ build tools](https://visualstudio.microsoft.com/visual-cpp-build-tools/)**
-
-
-
-
- 3) Go back to your terminal and type `cd PATH_TO_FOLDER` where `PATH_TO_FOLDER` is the address you copied in the previous step. Press Enter.
-
-
-
- Now that you're in the right place, copy and paste this command into the terminal and press Enter.
- ```
- conda env create -f CellProfiler_plugins_mac.yml
- ```
-
-
-
- 5. **Activate your environment**
-
- In your terminal, enter `conda activate CellProfiler_plugins` to activate your environment
-
-
-
-
- 6. **Verify that cellprofiler is installed correctly by running it from the command line.**
-
- In your terminal, type in `pythonw -m cellprofiler` and hit Enter. this will open CellProfiler or will give you an error message.
-
-
-
- 7. **Install other packages for other plugins**
-
- Certain packages cannot easily installed when importing the environment.
-
- For StarDist: In terminal with your environment activated, enter:
- ```
- pip install stardist csbdeep --no-deps
- ```
-
-
- If you would like to use the omnipose models in cellpose, ensure you have cellpose 1.0.2 (you should by default if you've used the CellProfiler_plugins_mac.yml file to generate the environment) and enter on the command line (in your activated environment):
-
- ```
- pip install omnipose
- ```
-
-
-
-
- 8. **Connect CellProfiler and the plugins repo**
-
- With your environment active, type `pythonw -m cellprofiler` in terminal to open CellProfiler if it is not open already.
-
- * In CellProfiler, go to **File** then **Preferences...**
- * Scroll down and look for "CellProfiler Plugins Directory" on the left.
- * Select the **Browse** button and choose the folder where you extracted the CellProfiler plugins files. It is probably called "CellProfiler-plugins-master" unless you have renamed it.
- * Select **Save** at the bottom of the Preferences window
- * Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` on the command line
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
-
-
-
- 9. **Verify that the installation worked**
-
- Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
- In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
- 
-
- ---
-
-
-## Common errors
-
-1. My wheels are failing to build
-
-- If you get a message like "ERROR: Failed building wheel for pyzmq" this usually means that you do not have pyzmq installed. Try to reinstall pyzmq.
-
-2. Java virtual machine cannot be found
-
-- If you're getting errors about Java, it means that java is not being configured properly on your system.
-- Make sure you have installed The Java Development Kit 11 [here](https://adoptopenjdk.net/). Note that newer versions may not work.
-- Make sure you've added environment variables at the **System** level and not at the **User** level.
-```
-brew install java
-
-# For the system Java wrappers to find this JDK, symlink it with
-sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
-
-# Set version in zshrc
-echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
-source ~/.zshrc
-```
-
-3. Installing pyzmq failed
-
-- You might get an error when trying to install pyzmq. Something like
-```
-ERROR: Command errored out with exit status 1:
-```
-And earlier in the traceback:
-```
-Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
- Cython >= 0.20 is required to compile pyzmq from a development branch.
- Please install Cython or download a release package of pyzmq.
-```
-
- To fix this, `conda install cython`
-
-
-## Installation instructions for CellProfiler, CellProfiler plugins and CellPose in a conda environment on Apple silicon (eg M1).
-
-1. **Install brew**
- ```
- /bin/bash -c "$(curl -fsSL https://raw.githubusercontent.com/Homebrew/install/HEAD/install.sh)"
- ```
-
-2. **Update brew**
- ```
- brew update
- ```
-
-3. **Install Java**
- ```
- brew install java
- ```
-
-4. **For the system Java wrappers to find this JDK and symlink it**
- ```
- sudo ln -sfn /opt/homebrew/opt/openjdk/libexec/openjdk.jdk /Library/Java/JavaVirtualMachines/openjdk.jdk
- ```
-
-5. **Set version in .zshrc**
- ```
- echo export JAVA_HOME=$(/usr/libexec/java_home -v 1.8) >> ~/.zshrc
- source ~/.zshrc
- ```
-
-6. **Brew install package requirements**
- ```
- brew install freetype mysql git
- ```
-
-7. **Direct to brew openssl**
- ```
- export LDFLAGS="-L$(brew --prefix)/opt/openssl/lib"
- ```
-
-8. **Install hdf5**
- ```
- brew install hdf5@1.12
- ```
-
-9. **Make sure to get the version directory correct for the version installed. Find with `ls /opt/homebrew/Cellar/hdf5/`**
- ```
- export HDF5_DIR=/opt/homebrew/Cellar/hdf5/1.12.1_1/
- ```
-10. **Create a folder and download cellprofiler-plugins**
-
- ```
- mkdir cp_plugins
- cd cp_plugins
- git clone https://github.com/CellProfiler/CellProfiler-plugins
- ```
-
-11. **Download and install miniconda**
-
- ```
- wget https://repo.anaconda.com/miniconda/Miniconda3-latest-MacOSX-arm64.sh -O ~/miniconda.sh
- ```
-
-
-12. **Create a conda `.yml` file**
- In a text editor, paste the following test and save it as `cellprofiler_plugins_macM1.yml`:
- ```
- name: cpcellpose
-
- channels:
- - conda-forge
- - anaconda
- - bioconda
- - defaults
- - apple
-
- dependencies:
- - python=3.8
- - pip
- - h5py
- - mysqlclient
- - imagecodecs
- - python.app
- - pandas
- - pip:
- - cellpose
- - attrdict
- - sip==5.5.0
- - boto3>=1.12.28
- - centrosome==1.2.1
- - docutils==0.15.2
- - h5py~=3.6.0
- - imageio>=2.5
- - inflect>=2.1
- - Jinja2>=2.11.2
- - joblib>=0.13
- - mahotas>=1.4
- - matplotlib==3.1.3
- - mysqlclient==1.4.6
- - numpy>=1.20.1
- - Pillow>=7.1.0
- - prokaryote==2.4.4
- - python-bioformats==4.0.6
- - python-javabridge==4.0.3
- - pyzmq~=22.3
- - sentry-sdk==0.18.0
- - requests>=2.22
- - scikit-image>=0.17.2
- - scikit-learn>=0.20
- - scipy>=1.4.1
- - six
- - tifffile<2022.4.22
- - wxPython==4.2.0
- ```
-
-
-13. **Create a conda environment using the cellprofiler_plugins_macM1.yml file**
-
- At this stage, your folder/file structure should look like this:
-
- ```
- ├── cp_plugins
- ├── CellProfiler-plugins
- └── cellprofiler_plugins_macM1.yml
- ```
-
- In the terminal, make sure you are in the `cp_plugins` folder mentioned above (run `cd cp_plugins` to get there).
-
- ```
- conda env create -n cp_plugins --file cellprofiler_plugins_macM1.yml
- ```
-
-14. **Activate the conda environment**
-
- ```
- conda activate cp_plugins
- ```
-
-15. **Install CellProfiler and CellProfiler-core**
- ```
- pip install cellprofiler-core
-
- # We install CellProfiler with no dependencies here since we installed them with the conda yml file
- # This allows us to have some dependencies for M1 that are not the default for CellProfiler (eg wxPython)
- git clone https://github.com/CellProfiler/CellProfiler
-
- pip install --no-deps cellprofiler
- ```
-
-16. **Open CellProfiler**
- ```
- pythonw -m cellprofiler
- ```
-
-17. **Connect CellProfiler with the plugins folder**
-
- 1. In CellProfiler, go to `File` then `Preferences...`
- 2. Scroll down and look for `CellProfiler Plugins Directory` on the left.
- 3. Select the Browse button and choose the folder where you extracted the CellProfiler plugins files. It is probably called `CellProfiler-plugins` unless you have renamed it.
- 4. Select Save at the bottom of the Preferences window
- 5. Close CellProfiler and reopen it by typing `pythonw -m cellprofiler` in the command line
-
-
-### Resolving dependencies conflicts
-
- There can be some strange dependency conflicts that can arise with python-javabridge. In the terminal with your environment activate, enter:
- ```
- pip uninstall -y centrosome python-javabridge
- pip install --no-cache-dir --no-deps --no-build-isolation python-javabridge centrosome
- pip uninstall matplotlib -y
- pip install matplotlib==3.2
- pip uninstall mahotas -y
- pip install mahotas
- ```
-
- The latest version of CellProfiler available on github removes the python-javabridge requirement and instead uses ScyJava.
-
-### Test your installation
-
-Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
-In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
diff --git a/documentation/CP-plugins-documentation/Windows_installation.md b/documentation/CP-plugins-documentation/Windows_installation.md
deleted file mode 100644
index 60bb701c..00000000
--- a/documentation/CP-plugins-documentation/Windows_installation.md
+++ /dev/null
@@ -1,208 +0,0 @@
-# How to install CellProfiler from source with all plugins on Windows
-
-Outlined here is the manual process for installing CellProfiler and plugins from source. Please close all open programs and save your work before starting this, as your computer may require a restart. Note that some of these download steps can take a while; the first 3 steps (as well as downloading Anaconda) can all be done simultaneously to save you time. Also note that for steps like changing the system-level environment variables, you may need admin permissions on your computer.
-
-**NEW:** We now have an automated install available for this workflow (requires administrator permissions on your computer) that will perform these steps for you. To get the necessary files and see instructions for the automatic install, look [here](https://github.com/CellProfiler/CellProfiler-plugins/tree/master/Instructions/Windows_batch_file).
-
-1. **Download and install [Microsoft Visual Studio C++ build tools](https://visualstudio.microsoft.com/visual-cpp-build-tools/)**
-
-  -
- **NOTE**: Make sure to check 'Desktop development with C++' under Desktop and Mobile in the installer:
-
-
-
- **NOTE**: Make sure to check 'Desktop development with C++' under Desktop and Mobile in the installer:
-
-  -
-
-
-2. **Download and install [Microsoft Visual C++ Redistributable 2015-2022]( https://docs.microsoft.com/en-us/cpp/windows/latest-supported-vc-redist?view=msvc-170)**
-
- You may already have this installed on your computer.
-
- Select the version appropriate for your architecture. On windows, you can determine this by going to **Control Panel** then searching for **System** and looking next to "System type:" for your processor architecture. Below is an example image:
-
-
-
-
-
-2. **Download and install [Microsoft Visual C++ Redistributable 2015-2022]( https://docs.microsoft.com/en-us/cpp/windows/latest-supported-vc-redist?view=msvc-170)**
-
- You may already have this installed on your computer.
-
- Select the version appropriate for your architecture. On windows, you can determine this by going to **Control Panel** then searching for **System** and looking next to "System type:" for your processor architecture. Below is an example image:
-
-  -
-
-
-
-3. **Download and install [Java JDK 11](https://adoptium.net/temurin/releases/?version=11)**
-
- Make sure you select the right version for your processor and operating system (see previous step). You can filter by operating system and processor Architecture.
-
-
-
-
-
-
-3. **Download and install [Java JDK 11](https://adoptium.net/temurin/releases/?version=11)**
-
- Make sure you select the right version for your processor and operating system (see previous step). You can filter by operating system and processor Architecture.
-
-  -
- You can alternatively install from [oracle.com](https://www.oracle.com/java/technologies/javase-jdk11-downloads.html) if you'd like, though you will need to make an Oracle account.
-
-
-
-
-4. **Update your Windows Environment Variables for Java**
-
- Go to **Control Panel** then search for **System**, then click on **Advanced System Settings**. Alternatively, for some versions of Windows, you can use the search bar to search for "Environment" and "Edit the system environment variables" will come up as an option.
-
-
-
- You can alternatively install from [oracle.com](https://www.oracle.com/java/technologies/javase-jdk11-downloads.html) if you'd like, though you will need to make an Oracle account.
-
-
-
-
-4. **Update your Windows Environment Variables for Java**
-
- Go to **Control Panel** then search for **System**, then click on **Advanced System Settings**. Alternatively, for some versions of Windows, you can use the search bar to search for "Environment" and "Edit the system environment variables" will come up as an option.
-
-  -
- From the System Properties window that pops up, select **Environment Variables...** Then add a new System variable by selecting **New...** under 'System Variables':
-
-
-
- From the System Properties window that pops up, select **Environment Variables...** Then add a new System variable by selecting **New...** under 'System Variables':
-
-  -
- Add (or if you already have it, modify) two system variables: `JAVA_HOME` and `JDK_HOME`. For each new variable, set its value to the location of your JDK installation (i.e., the location of the folder beginning with 'jdk11'). You can do this by clicking the **Browse Directory...** button. Usually this is in your 'Program Files' in a folder called 'Java'. Here is an example path for `JAVA_HOME`:
-
-
-
- Add (or if you already have it, modify) two system variables: `JAVA_HOME` and `JDK_HOME`. For each new variable, set its value to the location of your JDK installation (i.e., the location of the folder beginning with 'jdk11'). You can do this by clicking the **Browse Directory...** button. Usually this is in your 'Program Files' in a folder called 'Java'. Here is an example path for `JAVA_HOME`:
-
-  -
- Repeat the same for `JDK_HOME`:
-
-
-
- Repeat the same for `JDK_HOME`:
-
-  -
-
-
-
-
-5. **Download and install or update conda**
-
- For beginners, we recommend you use Anaconda Navigator since it is more beginner-friendly, but you can also use miniconda. [Download Anaconda](https://www.anaconda.com/products/distribution) from the website and install.
-
- **NOTE**: if you already have conda, you can get the command to update conda by typing `conda update` on your command line. The command will generally look like:
-
- `conda update --prefix /Users/USERNAME/opt/anaconda3 anaconda`
-
-
-
-
-6. **Clone the CellProfiler-plugins Repo**
-
- Download a copy of (aka "clone") the CellProfiler-plugins repo from [here](https://github.com/CellProfiler/CellProfiler-plugins.git). You can download the whole repo by cloning it with git or simply clicking the green **Code** button on [the repo page](https://github.com/CellProfiler/CellProfiler-plugins.git) and selecting **Download ZIP** (see below).
-
-
-
-
-
-
-
-5. **Download and install or update conda**
-
- For beginners, we recommend you use Anaconda Navigator since it is more beginner-friendly, but you can also use miniconda. [Download Anaconda](https://www.anaconda.com/products/distribution) from the website and install.
-
- **NOTE**: if you already have conda, you can get the command to update conda by typing `conda update` on your command line. The command will generally look like:
-
- `conda update --prefix /Users/USERNAME/opt/anaconda3 anaconda`
-
-
-
-
-6. **Clone the CellProfiler-plugins Repo**
-
- Download a copy of (aka "clone") the CellProfiler-plugins repo from [here](https://github.com/CellProfiler/CellProfiler-plugins.git). You can download the whole repo by cloning it with git or simply clicking the green **Code** button on [the repo page](https://github.com/CellProfiler/CellProfiler-plugins.git) and selecting **Download ZIP** (see below).
-
-  -
- If you download the ZIP file, be sure to extract the file contents by selecting **Extract All** in File Explorer:
-
-
-
- If you download the ZIP file, be sure to extract the file contents by selecting **Extract All** in File Explorer:
-
-  -
- You can also use git or GitHub Desktop to clone the repo if you prefer.
-
-
-
-7. **Create the environment from the .yml file**
-
- Open Anaconda Navigator and select the **Environments** tab on the left. We recommend you create the environment from the command line. To do this, Select the play button next to your base (root) environment and select **Open Terminal**:
-
-
-
- You can also use git or GitHub Desktop to clone the repo if you prefer.
-
-
-
-7. **Create the environment from the .yml file**
-
- Open Anaconda Navigator and select the **Environments** tab on the left. We recommend you create the environment from the command line. To do this, Select the play button next to your base (root) environment and select **Open Terminal**:
-
-  -
- A black box should pop up with a blinking cursor. This is your terminal. You now need to navigate to where the **cellprofiler_plugins_windows.yml** file is inside of the CellProfiler-plugins folder you downloaded in the last step. This file is in the `Instructions` subfolder. Here is how we recommend you do this:
-
- 1) In File Explorer, open the folder you downloaded in the previous step (usually called "CellProfiler-plugins-master") and then open the **Instructions** subfolder).
- 2) Click in the address bar area and this should highlight an address to where the folder is on your computer. **Copy this address**:
-
-
-
- A black box should pop up with a blinking cursor. This is your terminal. You now need to navigate to where the **cellprofiler_plugins_windows.yml** file is inside of the CellProfiler-plugins folder you downloaded in the last step. This file is in the `Instructions` subfolder. Here is how we recommend you do this:
-
- 1) In File Explorer, open the folder you downloaded in the previous step (usually called "CellProfiler-plugins-master") and then open the **Instructions** subfolder).
- 2) Click in the address bar area and this should highlight an address to where the folder is on your computer. **Copy this address**:
-
-  -
- 3) Go back to your terminal and type `cd PATH_TO_FOLDER` where `PATH_TO_FOLDER` is the address you copied in the previous step. Press Enter.
-
-
-
- Now that you're in the right place, copy and paste this command into the terminal and press Enter.
- ```
- conda env create -f CellProfiler_plugins_windows.yml
- ```
-
-
-8. **Activate your environment**
-
- In your terminal, enter `conda activate CP_plugins` to activate your environment
-
-
-
-9. **Verify that cellprofiler is installed correctly by running it from the command line.**
-
- In your terminal, type in `cellprofiler` and hit Enter. this will open CellProfiler or will give you an error message.
-
-
-
-10. **Install other packages for other plugins (just for RunStarDist)**
-
- In terminal with your environment activated, enter:
- ```
- pip install stardist csbdeep --no-deps
- ```
- If you would like to use the omnipose models in cellpose, ensure you have cellpose 1.0.2 (you should by default if you've used our environment yml) and enter on the command line (in your activated environment):
- ```
- pip install omnipose
- ```
-
-
-
-
-
-12. **Connect CellProfiler and the plugins folder**
-
- With your environment active, type `cellprofiler` in terminal to open CellProfiler if it is not open already.
-
- * In CellProfiler, go to **File** then **Preferences...**
- * Scroll down and look for "CellProfiler Plugins Directory" on the left.
- * Select the **Browse** button and choose the folder where you extracted the CellProfiler plugins files. It is probably called "CellProfiler-plugins-master" unless you have renamed it.
- * Select **Save** at the bottom of the Preferences window
- * Close CellProfiler and reopen it by typing `cellprofiler` on the command line
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
-
-
-
-13. **Verify that the installation worked**
-
- Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
- In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
-
-
- 3) Go back to your terminal and type `cd PATH_TO_FOLDER` where `PATH_TO_FOLDER` is the address you copied in the previous step. Press Enter.
-
-
-
- Now that you're in the right place, copy and paste this command into the terminal and press Enter.
- ```
- conda env create -f CellProfiler_plugins_windows.yml
- ```
-
-
-8. **Activate your environment**
-
- In your terminal, enter `conda activate CP_plugins` to activate your environment
-
-
-
-9. **Verify that cellprofiler is installed correctly by running it from the command line.**
-
- In your terminal, type in `cellprofiler` and hit Enter. this will open CellProfiler or will give you an error message.
-
-
-
-10. **Install other packages for other plugins (just for RunStarDist)**
-
- In terminal with your environment activated, enter:
- ```
- pip install stardist csbdeep --no-deps
- ```
- If you would like to use the omnipose models in cellpose, ensure you have cellpose 1.0.2 (you should by default if you've used our environment yml) and enter on the command line (in your activated environment):
- ```
- pip install omnipose
- ```
-
-
-
-
-
-12. **Connect CellProfiler and the plugins folder**
-
- With your environment active, type `cellprofiler` in terminal to open CellProfiler if it is not open already.
-
- * In CellProfiler, go to **File** then **Preferences...**
- * Scroll down and look for "CellProfiler Plugins Directory" on the left.
- * Select the **Browse** button and choose the folder where you extracted the CellProfiler plugins files. It is probably called "CellProfiler-plugins-master" unless you have renamed it.
- * Select **Save** at the bottom of the Preferences window
- * Close CellProfiler and reopen it by typing `cellprofiler` on the command line
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
-
-
-
-13. **Verify that the installation worked**
-
- Add a module to your pipeline by hitting the **+** button in the pipeline panel (bottom left)
-
- In the "Add Modules" window that pops up, type "run" into the search bar. You should be able to see plugins like RunCellpose and RunStarDist if the installation was successful:
-
-  -
-
-
----
-
-## Common errors
-
-1. My wheels are failing to build
-
- - If you get a message like "ERROR: Failed building wheel for pyzmq" this usually means that you do not have the Microsoft Visual Studio tools installed properly or the wrong version of pyzmq is being installed (e.g., you have a previous version already installed in your environment). See Step #1-2 above and ensure that you have "Desktop development with C++" selected under the install configuration options
-
-
-2. Java virtual machine cannot be found. This error might look something like `javabridge.jutil.JVMNotFoundError`
-
- - If you're getting errors about Java, it means that java is not being configured properly on your system.
- - Make sure you have installed The Java Development Kit 11. Note that newer versions of Java may not work.
- - Make sure you've added environment variables at the **System** level and not at the **User** level. You need to add both `JAVA_HOME` and `JDK_HOME` and both need to point to the folder that holds your jdk installation. Typically this path would look something like `C:\Program Files\Java\jdk-11.0.15.1` but it might be different on your machine depending on where you've installed Java.
- - If you have done these things and you still get this error, then try `pip install javabridge` (inside your activated cellprofiler-plugins conda environment) and then opening cellprofiler with `cellprofiler`
-
-
-3. Installing pyzmq failed
-
- You might get an error when trying to install pyzmq. Something like
- ```
- ERROR: Command errored out with exit status 1:
- ```
- And earlier in the traceback:
- ```
- Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
- Cython >= 0.20 is required to compile pyzmq from a development branch.
- Please install Cython or download a release package of pyzmq.
- ```
-
- To fix this, activate your environment and then `conda install cython`
-
- - You might also get an error like:
- ```
- AttributeError: 'MSVCCompiler' object has no attribute '_vcruntime_redist'
- ```
-
- Generally, this error means that you don't have the right Microsoft Visual Studio C compiler. You can try two things:
-
- 1. Look below in the code. Even if the wheel fails, pyzmq will still attempt to install. Look below for `Running setup.py install for pyzmq ... done` Also make sure you don't have any additional versions of pyzmq installed already.
- 2. You can try reinstalling Microsoft Visual C++ Build tools (#1-2). Versions 2008, 2015-2022 should work. Make sure you have the version of the redistributable compatible with your system processor (see step #2).
-
----
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
new file mode 100644
index 00000000..0020d62b
--- /dev/null
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -0,0 +1,64 @@
+# Using plugins
+
+Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all plugins in the plugin folder that you have set.
+The plugins will appear in the "Add Modules" panel like all standard modules and you can use the plugins as you would any other CellProfiler module.
+
+Please note that, as CellProfiler-plugins are considered experimental, they may not be as well documented as standard modules and they may not have a window that shows on run.
+Please report any installation issues or bugs related to plugins in the [CellProfiler-plugins repository](https://github.com/CellProfiler/CellProfiler-plugins) and not in the main CellProfiler repository.
+
+## Installation
+
+### Installing plugins without dependencies
+
+If your plugin doesn't require additional dependencies, all you need to do is
+
+1. **Install CellProfiler.**
+Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
+
+Or, you can install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
+
+2. **Clone the CellProfiler-plugins repository.**
+This will download all of the plugins in the CellProfiler-plugins repository.
+
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+Alternatively, if you have code for plugins that are not in the CellProfiler-plugins repository, you can place them in any folder that you'll be able to find again.
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler and re-open it.
+
+You are now ready to use any CellProfiler plugin that does not have additional dependencies.
+This is most CellProfiler plugins.
+
+### Installing plugins with dependencies
+
+1. **Install CellProfiler.**
+Install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
+
+2. **Clone the CellProfiler-plugins repository.**
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler and re-open it.
+
+4. **Install the dependencies for your plugin.**
+In your terminal, type the following, where FLAG is the flag specific to the module you would like to run, as noted in [Supported Plugins](supported_plugins.md).
+```bash
+cd CellProfiler-plugins
+pip install -e .[FLAG]
+```
+
+Please note that plugins that have separate install flags may have conflicting dependencies so we recommend making a separate python environment in which to run separate installations.
+(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
\ No newline at end of file
From 4c41db2f05cabb63f634e1ff9c07f459ca39dd72 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 14:03:04 -0700
Subject: [PATCH 2/6] deprecate declumpobjects, add info about unsupported
plugins
---
.../unsupported_plugins.md | 14 +++++++++++++-
.../CellProfiler4/declumpobjects.py | 0
2 files changed, 13 insertions(+), 1 deletion(-)
rename declumpobjects.py => unmaintained_plugins/CellProfiler4/declumpobjects.py (100%)
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index 7e0343da..4ca10a6c 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -8,5 +8,17 @@ We welcome requests for updating particular unsupported plugins, but please note
We welcome community contributed plugin updates.
Likewise, we welcome contributions of community developed plugins.
-Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team or community contributed.
+Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
+## Where are unsupported plugins?
+
+Unsupported plugins can be found in the `unmaintained_plugins` folder in the CellProfiler-plugins repository.
+Those plugins in the `CellProfiler2`, `CellProfiler3`, and `CellProfiler4` folders were, at one point, supported for those versions of CellProfiler.
+Those plugins in the `CellProfiler4_autoconverted` folder were automatically converted from Python2 to Python3 (to support the transition from Python2 in CellProfiler3 to Python3 in CellProfiler4) but were never fully supported and may or may not run.
+
+## What plugins are unsupported?
+
+We cannot provide comprehensive information about why we are not supporting a given plugin.
+Information about select plugins is as follows:
+
+**DeclumpObjects**: DeclumpObjects will split objects based on a seeded watershed method. Functionality from this module was [added into CellProfiler](https://github.com/CellProfiler/CellProfiler/pull/4397) in the Watershed module as of CellProfiler 4.2.0.
diff --git a/declumpobjects.py b/unmaintained_plugins/CellProfiler4/declumpobjects.py
similarity index 100%
rename from declumpobjects.py
rename to unmaintained_plugins/CellProfiler4/declumpobjects.py
From d47666e9fcefc1f98227e02aa4692ed83a022e0c Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 14:20:25 -0700
Subject: [PATCH 3/6] more documentation
---
.../contributing_plugins.md | 17 +++-
.../CP-plugins-documentation/overview.md | 11 +++
.../supported_plugins.md | 77 +++++--------------
.../CP-plugins-documentation/using_GPU.md | 8 ++
4 files changed, 51 insertions(+), 62 deletions(-)
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index 204ec8f8..c3ae78be 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -1,6 +1,8 @@
-## Contributing New Plugins
+# Contributing New Plugins
-Refer to CellProfiler repository and within cellprofiler/modules/plugins you can find two different templates to use for creating your own plugin.
+Within our CellProfiler wiki, you can find an [orientation to CellProfiler code](https://github.com/CellProfiler/CellProfiler/wiki/Orientation-to-CellProfiler-code).
+
+In the CellProfiler repository, within [cellprofiler/modules/plugins](https://github.com/CellProfiler/CellProfiler/tree/master/cellprofiler/modules/plugins) you can find two different templates to use for creating your own plugin.
`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
@@ -8,10 +10,17 @@ In you plugin, you must include:
In your plugin, we appreciate if you also include:
- display functionality
+- extensive module documentation
+- references and citation information in your module documentation
+Please create a Pull Request to the CellProfiler-plugins repository to submit your plugin for inclusion in the repository.
-In your PR, you must include:
+In your PR, you must:
- add your plugin to the [supported_plugins](supported_plugins.md) documentation page
In your PR, we appreciate if you also include:
-- unit tests for your plugin
\ No newline at end of file
+- unit tests for your plugin
+
+## Having your plugin cited
+
+While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/overview.md b/documentation/CP-plugins-documentation/overview.md
index 5ece4c8d..1616e1a3 100644
--- a/documentation/CP-plugins-documentation/overview.md
+++ b/documentation/CP-plugins-documentation/overview.md
@@ -9,6 +9,17 @@ A module may be in CellProfiler-plugins instead of CellProfiler itself because:
- it requires extra libraries or other dependencies we are unable or unwilling to require for CellProfiler
- it has been contributed by a community member
+## How do I use CellProfiler Plugins?
+
+See our [Using Plugins](using_plugins.md) page for information on how to install and use plugins.
+
+## What plugins are available?
+
+See our [Supported Plugins](supported_plugins.md) page for information on all currently supported plugins.
+
+## How do I contribute a plugin?
+
+See our [Contributing Plugins](contributing_plugins.md) page for information on contributing a plugin to CellPainting-plugins.
## Who made this?
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index 5850822a..deb60d61 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -3,61 +3,22 @@
Below is a brief overview of our currently supported plugins.
For details about using any particular plugin, please read the module documentation inside the plugin in CellProfiler.
-## CalculateMoments
-CalculateMoments extracts moments statistics from a given distribution of pixel values.
-
-## CallBarcodes
-CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles.
-It is used for optical sequencing by synthesis (SBS).
-
-## CompensateColors
-CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through.
-It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched.
-
-## DeclumpObjects
-DeclumpObjects will split objects based on a seeded watershed method.
-
-## DistanceTransform
-DistanceTransform computes the distance transform of a binary image.
-The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1.
-
-## EnhancedMeasureTexture
-EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler.
-
-## HistogramEqualization
-HistogramEqualization increases the global contrast of a low-contrast image or volume.
-Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct.
-This module can perform either global or local histogram equalization.
-
-## HistogramMatching
-HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image.
-It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image.
-It allows you to choose which frame to use as the reference.
-
-## PixelShuffle
-PixelShuffle takes the intensity of each pixel in an image and it randomly shuffles its position.
-
-## Predict
-Predict allows you to use an ilastik pixel classifier to generate a probability image.
-CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage).
-
-## RunCellpose
-RunCellpose allows you to run Cellpose within CellProfiler.
-Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells.
-You can use pre-trained Cellpose models or your custom model with this plugin.
-You can use a GPU with this module to dramatically increase your speed/efficiency.
-
-## RunImageJScript
-
-## RunOmnipose
-
-## RunStarDist
-RunStarDist allows you to run StarDist within CellProfiler.
-StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells.
-You can use pre-trained StarDist models or your custom model with this plugin.
-You can use a GPU with this module to dramatically increase your speed/efficiency.
-RunStarDist is generally faster than RunCellpose.
-
-## VarianceTransform
-This module allows you to calculate the variance of an image, using a determined window size.
-It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image.
\ No newline at end of file
+Most plugins will run without any special installation of either CellProfiler or the plugins.
+See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin use as well as for installation information for those plugins that do require installation of dependencies.
+
+| Plugin | Description | Requires installation of dependencies? | Install flag |
+|--------|-------------|----------------------------------------|--------------|
+| CalculateMoments | CalculateMoments extracts moments statistics from a given distribution of pixel values. | No | |
+| CallBarcodes | CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles. It is used for optical sequencing by synthesis (SBS). | No | |
+| CompensateColors | CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through. It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched. | No | |
+| DistanceTransform | DistanceTransform computes the distance transform of a binary image. The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1. | No | |
+| EnhancedMeasureTexture| EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler. | No | |
+| HistogramEqualization | HistogramEqualization increases the global contrast of a low-contrast image or volume. Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct. This module can perform either global or local histogram equalization. | No | |
+| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
+| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
+| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | Yes | XXXXX |
+| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
+| RunImageJScript | | Yes | XXXXX |
+| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
+| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
+| VarianceTransform | This module allows you to calculate the variance of an image, using a determined window size. It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image. | No | |
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/using_GPU.md b/documentation/CP-plugins-documentation/using_GPU.md
index 945d869b..b5dfc4d1 100644
--- a/documentation/CP-plugins-documentation/using_GPU.md
+++ b/documentation/CP-plugins-documentation/using_GPU.md
@@ -24,3 +24,11 @@ If you want to use a GPU to run the model (this is recommended for speed), you'l
pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
```
If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
+
+
+ **NOTE**: You might get a warning like this:
+ ```
+ W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
+ 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
+ ```
+ If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
From 12c52c77ec80879b399743160d6accbfe2049d26 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 12:35:02 -0700
Subject: [PATCH 4/6] more docs
---
.../contributing_plugins.md | 3 +-
.../supported_plugins.md | 4 +-
.../unsupported_plugins.md | 4 +-
.../CP-plugins-documentation/using_plugins.md | 106 ++++++++++++++++--
.../CP-plugins-documentation/versions.md | 49 +-------
5 files changed, 104 insertions(+), 62 deletions(-)
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index c3ae78be..f0084668 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -6,8 +6,6 @@ In the CellProfiler repository, within [cellprofiler/modules/plugins](https://gi
`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
-In you plugin, you must include:
-
In your plugin, we appreciate if you also include:
- display functionality
- extensive module documentation
@@ -20,6 +18,7 @@ In your PR, you must:
In your PR, we appreciate if you also include:
- unit tests for your plugin
+- customized installation in setup.py, if your plugin has dependencies
## Having your plugin cited
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index deb60d61..a17866c0 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -16,9 +16,9 @@ See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin
| HistogramEqualization | HistogramEqualization increases the global contrast of a low-contrast image or volume. Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct. This module can perform either global or local histogram equalization. | No | |
| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
-| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | Yes | XXXXX |
+| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | No | |
| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
-| RunImageJScript | | Yes | XXXXX |
+| RunImageJScript | RunImageJScript allows you to run any supported ImageJ script directly within CellProfiler. It is significantly more performant than RunImageJMacro, and is also less likely to leave behind temporary files. | Yes | XXXXX |
| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
| VarianceTransform | This module allows you to calculate the variance of an image, using a determined window size. It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image. | No | |
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index 4ca10a6c..b7cf3fe8 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -5,11 +5,9 @@ Unsupported plugins are primarily unsupported for either of two reasons:
- Their functions have been integrated into the current version of CellProfiler and therefore a plugin is no longer necessary
We welcome requests for updating particular unsupported plugins, but please note that we have limited bandwidth for working on plugins and may be unable to complete the update.
+Additionally, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
We welcome community contributed plugin updates.
-Likewise, we welcome contributions of community developed plugins.
-Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
-
## Where are unsupported plugins?
Unsupported plugins can be found in the `unmaintained_plugins` folder in the CellProfiler-plugins repository.
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index 0020d62b..bce4bc70 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -8,9 +8,16 @@ Please report any installation issues or bugs related to plugins in the [CellPro
## Installation
-### Installing plugins without dependencies
+If the plugin you would like to use does not have any additional dependencies outside of those required for running CellProfiler (this is most plugins), using plugins is very simple.
+See [Installing plugins without dependencies](#installing-plugins-without-dependencies).
+
+If the plugin you would like to use has dependencies, you have two separate options for installation.
+The first option requires building CellProfiler from source, but plugin installation is simpler.
+See [Installing plugins with dependencies, using CellProfiler from source](#installing-plugins-with-dependencies-using-cellprofiler-from-source).
+The second option allows you to use pre-built CellProfiler, but plugin installation is more complex.
+See [Installing plugins with dependencies, using pre-built CellProfiler](#installing-plugins-with-dependencies-using-pre-built-cellprofiler).
-If your plugin doesn't require additional dependencies, all you need to do is
+### Installing plugins without dependencies
1. **Install CellProfiler.**
Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
@@ -31,12 +38,12 @@ Alternatively, if you have code for plugins that are not in the CellProfiler-plu
- Open CellProfiler.
- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
- Select `Save` at the bottom of the Preferences window
-- Close CellProfiler and re-open it.
+- Close CellProfiler and re-open it
You are now ready to use any CellProfiler plugin that does not have additional dependencies.
This is most CellProfiler plugins.
-### Installing plugins with dependencies
+### Installing plugins with dependencies, using CellProfiler from source
1. **Install CellProfiler.**
Install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
@@ -49,9 +56,9 @@ git clone https://github.com/CellProfiler/CellProfiler-plugins.git
3. **Set the plugins path in CellProfiler.**
- Open CellProfiler.
-- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned.
- Select `Save` at the bottom of the Preferences window
-- Close CellProfiler and re-open it.
+- Close CellProfiler
4. **Install the dependencies for your plugin.**
In your terminal, type the following, where FLAG is the flag specific to the module you would like to run, as noted in [Supported Plugins](supported_plugins.md).
@@ -59,6 +66,91 @@ In your terminal, type the following, where FLAG is the flag specific to the mod
cd CellProfiler-plugins
pip install -e .[FLAG]
```
+e.g. To install Cellpose the pip install command would be `pip install -e .[cellpose]`
+
+5. **Open and use CellProfiler.**
Please note that plugins that have separate install flags may have conflicting dependencies so we recommend making a separate python environment in which to run separate installations.
-(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
\ No newline at end of file
+(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
+
+### Installing plugins with dependencies, using pre-built CellProfiler
+
+1. **Install CellProfiler.**
+Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
+
+2. **Clone the CellProfiler-plugins repository.**
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned.
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler
+
+4. **Identify the Python dependencies of the plugins you want to use.**
+You can find this information for most plugins by looking in `setup.py`.
+```{admonition} e.g. using RunCellpose plugin
+If you would like to install the RunCellpose plugin, in `setup.py` you can see that `cellpose_deps = ["cellpose>=1.0.2"]` so the only dependency is `cellpose`.
+```
+
+Alternatively, you can find this information directly in the plugin code itself by looking at what is imported at the beginning of the plugin.
+Note that you will want to compare any imports to what is already required by CellProfiler with that caveat that dependencies often have dependencies.
+```{admonition} e.g. using RunImageJScript plugin
+If you would like to install the RunImageJScript plugin, you'll notice a number of imports in the beginning of the plugin code.
+You can ignore all those from `cellprofiler_core` as they are required for CellProfiler installation.
+You can ignore all those that are native to Python such as `sys`, `time`, and `threading`.
+A comparison with CellProfiler dependencies will show you that `skimage` is already installed (you can see this in CellProfiler's [environment.yml](https://github.com/CellProfiler/CellProfiler/blob/master/environment.yml) which lists `scikit-image` as a `cellprofiler-core` dependency or in CellProfiler's [setup.py](https://github.com/CellProfiler/CellProfiler/blob/master/setup.py) which lists `scikit-image` as an installation requirement).
+What remains is `pyimagej`
+```
+
+5. **Create a conda environment and install requirements in it**
+Create a conda (or other virtual) environment with a Python version matching that of CellProfiler.
+CellProfiler 4 uses Python 3.8 (which you can see in CellProfiler's [setup.py](https://github.com/CellProfiler/CellProfiler/blob/master/setup.py)).
+Activate the environment and install the plugin requirement/s into it.
+
+In your terminal, type
+```
+# Create a conda environment
+conda create --name ENV_NAME python=3.8
+# Activate the conda environment
+conda activate ENV_NAME
+# Install the plugin requirement/s
+pip install REQUIREMENT
+```
+```{admonition} e.g. using RunImageJScript plugin
+conda create --name cp-ij python=3.8
+conda activate cp-ij
+pip install pyimagej
+```
+
+6. **Copy plugin requirements into CellProfiler installation**
+Find the folder that you installed into in the previous step (where to find these libraries in your local environment) by entering `pip show REQUIREMENT` into your terminal e.g. `pip show pyimagej`.
+In the information it returns, under `Location` you will find a path that will look something like `/Users/eweisbar/mambaforge/envs/cp-ij/lib/python3.8/site-packages`.
+
+Find the folder in which you have installed CellProfiler.
+On Windows this is likely to be `C:\Program Files\CellProfiler` and on Mac it is likely to be `/Applications/CellProfiler.app/Contents/MacOS/`.
+In your terminal, if you `ls` (for Mac) or `dir` (for Windows) that folder (e.g. `ls /Applications/CellProfiler.app/Contents/MacOS/`), you should see a list of all the dependencies that were packaged with CellProfiler.
+
+Copy the folders and their corresponding `.dist-info` folders (e.g. `pytz` and `pytz-2023.3.dist-info`) for any dependencies that were installed with the installation of `pyimagej` that are not already in the CellProfiler folder.
+
+```{admonition} e.g. using RunImageJScript plugin
+These are all the folders you need to copy over:
+`pytz`
+`pytz-2023.3.dist-info`
+`xarray`
+`xarray-2023.1.0.dist-info`
+`imglyb`
+`imglyb-2.1.0.dist-info`
+`jgo`
+`jgo-1.0.5.dist-info`
+`jpype`
+`_jpype.cpython-38-darwin.so`
+`imagej`
+`pyimagej-1.4.1.dist-info`
+```
+
+7. **Open and use CellProfiler.**
+When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/versions.md b/documentation/CP-plugins-documentation/versions.md
index 3755e75b..0f8db604 100644
--- a/documentation/CP-plugins-documentation/versions.md
+++ b/documentation/CP-plugins-documentation/versions.md
@@ -1,55 +1,8 @@
# Versions
-The most current release can always be found on DockerHub at `cellprofiler/distributed-cellprofiler`.
-Current version is 2.0.0.
-Our current tag system is dcpversion_CellProfilerversion, e.g. 1.2.0_3.1.8
-
-Previous release versions can be accessed at `bethcimini/distributed-cellprofiler:versionnumber`
+Current version is
---
# Version History
-## 2.1.0 (forthcoming)
-* Increase support for using plugins
-* Improve role handling and allow file download/upload from/to S3 buckets in different accounts
-* Improve file download to S3FS can be completely bypassed
-* Name EBS volumes
-
-## 2.0.0rc2 - Released 20201110
-* Add optional ability to download files to EBS rather than reading from S3 (helpful for pipelines that access many files/image sets)
-
-## 2.0.0rc1 - Released 20201105
-
-* Remove requirement for boto and fabric, using only boto3
-* Add support for Python 3 and CellProfiler 4
-* Move cluster size, machine type, and machine price to the config file from the fleet file, eliminating mismatches between the two
-* Add the ability to filter only files with certain names when running CHECK_IF_DONE
-* Don't cancel a fleet over capacity errors
-* Add "cheapest" mode to the monitor, allowing you to run more cheaply (at possible expense of running more slowly)
-
-## 1.2.2 - Released 20201103
-
-* Allow pipelines using batch files to also designate an input output_top_directory
-* Add support for multiple LaunchData specifications
-* Add CellProfiler-plugins
-* Additional way to create job submissions with run_batch_general.py
-
-## 1.2.1 - Released 20200109, Updated through 20191002
-
-* Allow monitor to downscale machines when number of jobs < number of machines
-* Add a parameter to discount files when running CHECK_IF_DONE checks if less than a certain size
-
-## 1.2.0 - Released 20181108, Updated through 20200109
-
-* Improved compatibility with CellProfiler 2 and 3
-* Better handling of logging when using output_structure
-
-## 1.1.0 - Released 20170217, Updated 20170221 (bugfixes) - 20181018
-
-* Changes in this release:
-
- * Added the `output_structure` variable to the job file, which allows you to structure the output folders created by DCP (ie `Plate/Well-Site` rather than `Plate-Well-Site`). Job files lacking this variable will still default to the previous settings (hyphenating all Metadata items in order they are presented in the Metadata grouping).
- * Added support for creating the list of groups via `cellprofiler --print-groups`- see [this issue](https://github.com/CellProfiler/Distributed-CellProfiler/issues/52) for example and discussion. Groups listed in this way MUST use the `output_structure` variable to state their desired folder structure or an error will be returned.
-
-## 1.0.0 - Version as of 20170213
From fffb5566c3b4d2892c75c30b14469e66e94bca6a Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 15:04:09 -0700
Subject: [PATCH 5/6] more doc changes
---
.../CP-plugins-documentation/RunCellPose.md | 34 +++++++++++++++++++
.../contributing_plugins.md | 9 ++++-
.../supported_plugins.md | 5 ++-
.../unsupported_plugins.md | 1 +
.../CP-plugins-documentation/using_GPU.md | 34 -------------------
.../CP-plugins-documentation/using_plugins.md | 3 +-
6 files changed, 49 insertions(+), 37 deletions(-)
create mode 100644 documentation/CP-plugins-documentation/RunCellPose.md
delete mode 100644 documentation/CP-plugins-documentation/using_GPU.md
diff --git a/documentation/CP-plugins-documentation/RunCellPose.md b/documentation/CP-plugins-documentation/RunCellPose.md
new file mode 100644
index 00000000..168fa7cc
--- /dev/null
+++ b/documentation/CP-plugins-documentation/RunCellPose.md
@@ -0,0 +1,34 @@
+# RunCellPose
+
+## Using RunCellPose with a GPU
+
+If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU.
+General instructions are available at this [link](https://pytorch.org/get-started/locally/).
+
+1. Your GPU should be visible in Device Manager under Display Adaptors.
+If your GPU isn't there, you likely need to install drivers.
+[Here](https://www.nvidia.com/Download/Find.aspx) is where you can find NVIDIA GPU drivers if you need to install them.
+
+
+2. To test whether the GPU is configured correctly:
+ * Run `python` on the command line (i.e., in Command Prompt or Terminal) to start an interactive session
+ * Then run the following
+ ```
+ import torch
+ torch.cuda.is_available()
+ ```
+ * If this returns `True`, you're all set
+ * If this returns `False`, you likely need to install/reinstall torch. See [here](https://pytorch.org/get-started/locally/) for your exact command.
+ * Exit the session with `exit()` then install torch if necessary
+ ```
+ pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
+ ```
+ If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
+
+
+**NOTE**: You might get a warning like this:
+```
+W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
+2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
+```
+If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index f0084668..8907b250 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -20,6 +20,13 @@ In your PR, we appreciate if you also include:
- unit tests for your plugin
- customized installation in setup.py, if your plugin has dependencies
+## Contributing bug fixes or updating deprecated plugins
+
+Please create a Pull Request to the CellProfiler-plugins repository to submit a bug fix or plugin un-deprecation.
+If you would like to update a currently deprecated plugin, we encourage you to read closed Issues in GitHub relevant to the plugin as they may contain helpful information about bugs present at the time of deprecation.
+
## Having your plugin cited
-While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
\ No newline at end of file
+While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
+
+See [Citing Plugins](citing.md) for information on citing CellProfiler Plugins.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index a17866c0..5ccdda93 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -6,6 +6,9 @@ For details about using any particular plugin, please read the module documentat
Most plugins will run without any special installation of either CellProfiler or the plugins.
See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin use as well as for installation information for those plugins that do require installation of dependencies.
+Most plugin documentation can be found within the plugin itself and can be accessed through CellProfiler help.
+Those plugins that do have extra documentation contain links below.
+
| Plugin | Description | Requires installation of dependencies? | Install flag |
|--------|-------------|----------------------------------------|--------------|
| CalculateMoments | CalculateMoments extracts moments statistics from a given distribution of pixel values. | No | |
@@ -17,7 +20,7 @@ See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin
| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | No | |
-| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
+| [RunCellpose](RunCellPose.md) | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
| RunImageJScript | RunImageJScript allows you to run any supported ImageJ script directly within CellProfiler. It is significantly more performant than RunImageJMacro, and is also less likely to leave behind temporary files. | Yes | XXXXX |
| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index b7cf3fe8..5e277b61 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -19,4 +19,5 @@ Those plugins in the `CellProfiler4_autoconverted` folder were automatically con
We cannot provide comprehensive information about why we are not supporting a given plugin.
Information about select plugins is as follows:
+**ClassifyPixelsUNET**: ClassifyPixelsUNET is a pixel classifier for background/object edge/object body. As far as we are aware, other deep learning based plugins that we do currently support (such as RunCellpose) work better.
**DeclumpObjects**: DeclumpObjects will split objects based on a seeded watershed method. Functionality from this module was [added into CellProfiler](https://github.com/CellProfiler/CellProfiler/pull/4397) in the Watershed module as of CellProfiler 4.2.0.
diff --git a/documentation/CP-plugins-documentation/using_GPU.md b/documentation/CP-plugins-documentation/using_GPU.md
deleted file mode 100644
index b5dfc4d1..00000000
--- a/documentation/CP-plugins-documentation/using_GPU.md
+++ /dev/null
@@ -1,34 +0,0 @@
-# Using a GPU with CellProfiler Plugins
-
-These CellProfiler modules currently support using a GPU: LIST MODULES
-
-## Tips for Setting Up Your GPU (if you have one)
-
-If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU. General instructions are available at this [link](https://pytorch.org/get-started/locally/).
-
-1. Your GPU should be visible in Device Manager under Display Adaptors. If your GPU isn't there, you likely need to install drivers.
- * [Here](https://www.nvidia.com/Download/Find.aspx) is where you can find NVIDIA GPU drivers if you need to install them.
-
-
-2. To test whether the GPU is configured correctly:
- * Run `python` on the command line (i.e., in Command Prompt or Terminal) to start an interactive session
- * Then run the following
- ```
- import torch
- torch.cuda.is_available()
- ```
- * If this returns `True`, you're all set
- * If this returns `False`, you likely need to install/reinstall torch. See [here](https://pytorch.org/get-started/locally/) for your exact command.
- * Exit the session with `exit()` then install torch if necessary
- ```
- pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
- ```
- If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index bce4bc70..946b9ab5 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -153,4 +153,5 @@ These are all the folders you need to copy over:
```
7. **Open and use CellProfiler.**
-When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing.
\ No newline at end of file
+When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing in the terminal window that opens with CellProfiler on Windows machines.
+This information is not available in Mac machines.
\ No newline at end of file
From f99b97e8c94b878f6e7a6059bd5f926f9901b41c Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 15:09:02 -0700
Subject: [PATCH 6/6] docs
---
documentation/CP-plugins-documentation/using_plugins.md | 5 ++++-
1 file changed, 4 insertions(+), 1 deletion(-)
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index 946b9ab5..bab82fe8 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -1,7 +1,10 @@
# Using plugins
-Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all plugins in the plugin folder that you have set.
+Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all useable plugins in the plugin folder that you have set.
The plugins will appear in the "Add Modules" panel like all standard modules and you can use the plugins as you would any other CellProfiler module.
+Typically, if a plugin's dependencies are not installed, it will not be visible in the "Add Modules" panel.
+If you cannot find any plugins in the "Add Modules" panel then you have not properly set the plugins path (see below).
+If you can find some but not all plugins in the "Add Modules" panel then the not-visible plugins have unmet dependencies.
Please note that, as CellProfiler-plugins are considered experimental, they may not be as well documented as standard modules and they may not have a window that shows on run.
Please report any installation issues or bugs related to plugins in the [CellProfiler-plugins repository](https://github.com/CellProfiler/CellProfiler-plugins) and not in the main CellProfiler repository.
-
-
-
----
-
-## Common errors
-
-1. My wheels are failing to build
-
- - If you get a message like "ERROR: Failed building wheel for pyzmq" this usually means that you do not have the Microsoft Visual Studio tools installed properly or the wrong version of pyzmq is being installed (e.g., you have a previous version already installed in your environment). See Step #1-2 above and ensure that you have "Desktop development with C++" selected under the install configuration options
-
-
-2. Java virtual machine cannot be found. This error might look something like `javabridge.jutil.JVMNotFoundError`
-
- - If you're getting errors about Java, it means that java is not being configured properly on your system.
- - Make sure you have installed The Java Development Kit 11. Note that newer versions of Java may not work.
- - Make sure you've added environment variables at the **System** level and not at the **User** level. You need to add both `JAVA_HOME` and `JDK_HOME` and both need to point to the folder that holds your jdk installation. Typically this path would look something like `C:\Program Files\Java\jdk-11.0.15.1` but it might be different on your machine depending on where you've installed Java.
- - If you have done these things and you still get this error, then try `pip install javabridge` (inside your activated cellprofiler-plugins conda environment) and then opening cellprofiler with `cellprofiler`
-
-
-3. Installing pyzmq failed
-
- You might get an error when trying to install pyzmq. Something like
- ```
- ERROR: Command errored out with exit status 1:
- ```
- And earlier in the traceback:
- ```
- Fatal: Cython-generated file 'zmq\backend\cython\_device.c' not found.
- Cython >= 0.20 is required to compile pyzmq from a development branch.
- Please install Cython or download a release package of pyzmq.
- ```
-
- To fix this, activate your environment and then `conda install cython`
-
- - You might also get an error like:
- ```
- AttributeError: 'MSVCCompiler' object has no attribute '_vcruntime_redist'
- ```
-
- Generally, this error means that you don't have the right Microsoft Visual Studio C compiler. You can try two things:
-
- 1. Look below in the code. Even if the wheel fails, pyzmq will still attempt to install. Look below for `Running setup.py install for pyzmq ... done` Also make sure you don't have any additional versions of pyzmq installed already.
- 2. You can try reinstalling Microsoft Visual C++ Build tools (#1-2). Versions 2008, 2015-2022 should work. Make sure you have the version of the redistributable compatible with your system processor (see step #2).
-
----
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
new file mode 100644
index 00000000..0020d62b
--- /dev/null
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -0,0 +1,64 @@
+# Using plugins
+
+Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all plugins in the plugin folder that you have set.
+The plugins will appear in the "Add Modules" panel like all standard modules and you can use the plugins as you would any other CellProfiler module.
+
+Please note that, as CellProfiler-plugins are considered experimental, they may not be as well documented as standard modules and they may not have a window that shows on run.
+Please report any installation issues or bugs related to plugins in the [CellProfiler-plugins repository](https://github.com/CellProfiler/CellProfiler-plugins) and not in the main CellProfiler repository.
+
+## Installation
+
+### Installing plugins without dependencies
+
+If your plugin doesn't require additional dependencies, all you need to do is
+
+1. **Install CellProfiler.**
+Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
+
+Or, you can install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
+
+2. **Clone the CellProfiler-plugins repository.**
+This will download all of the plugins in the CellProfiler-plugins repository.
+
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+Alternatively, if you have code for plugins that are not in the CellProfiler-plugins repository, you can place them in any folder that you'll be able to find again.
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler and re-open it.
+
+You are now ready to use any CellProfiler plugin that does not have additional dependencies.
+This is most CellProfiler plugins.
+
+### Installing plugins with dependencies
+
+1. **Install CellProfiler.**
+Install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
+
+2. **Clone the CellProfiler-plugins repository.**
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler and re-open it.
+
+4. **Install the dependencies for your plugin.**
+In your terminal, type the following, where FLAG is the flag specific to the module you would like to run, as noted in [Supported Plugins](supported_plugins.md).
+```bash
+cd CellProfiler-plugins
+pip install -e .[FLAG]
+```
+
+Please note that plugins that have separate install flags may have conflicting dependencies so we recommend making a separate python environment in which to run separate installations.
+(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
\ No newline at end of file
From 4c41db2f05cabb63f634e1ff9c07f459ca39dd72 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 14:03:04 -0700
Subject: [PATCH 2/6] deprecate declumpobjects, add info about unsupported
plugins
---
.../unsupported_plugins.md | 14 +++++++++++++-
.../CellProfiler4/declumpobjects.py | 0
2 files changed, 13 insertions(+), 1 deletion(-)
rename declumpobjects.py => unmaintained_plugins/CellProfiler4/declumpobjects.py (100%)
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index 7e0343da..4ca10a6c 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -8,5 +8,17 @@ We welcome requests for updating particular unsupported plugins, but please note
We welcome community contributed plugin updates.
Likewise, we welcome contributions of community developed plugins.
-Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team or community contributed.
+Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
+## Where are unsupported plugins?
+
+Unsupported plugins can be found in the `unmaintained_plugins` folder in the CellProfiler-plugins repository.
+Those plugins in the `CellProfiler2`, `CellProfiler3`, and `CellProfiler4` folders were, at one point, supported for those versions of CellProfiler.
+Those plugins in the `CellProfiler4_autoconverted` folder were automatically converted from Python2 to Python3 (to support the transition from Python2 in CellProfiler3 to Python3 in CellProfiler4) but were never fully supported and may or may not run.
+
+## What plugins are unsupported?
+
+We cannot provide comprehensive information about why we are not supporting a given plugin.
+Information about select plugins is as follows:
+
+**DeclumpObjects**: DeclumpObjects will split objects based on a seeded watershed method. Functionality from this module was [added into CellProfiler](https://github.com/CellProfiler/CellProfiler/pull/4397) in the Watershed module as of CellProfiler 4.2.0.
diff --git a/declumpobjects.py b/unmaintained_plugins/CellProfiler4/declumpobjects.py
similarity index 100%
rename from declumpobjects.py
rename to unmaintained_plugins/CellProfiler4/declumpobjects.py
From d47666e9fcefc1f98227e02aa4692ed83a022e0c Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Fri, 26 May 2023 14:20:25 -0700
Subject: [PATCH 3/6] more documentation
---
.../contributing_plugins.md | 17 +++-
.../CP-plugins-documentation/overview.md | 11 +++
.../supported_plugins.md | 77 +++++--------------
.../CP-plugins-documentation/using_GPU.md | 8 ++
4 files changed, 51 insertions(+), 62 deletions(-)
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index 204ec8f8..c3ae78be 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -1,6 +1,8 @@
-## Contributing New Plugins
+# Contributing New Plugins
-Refer to CellProfiler repository and within cellprofiler/modules/plugins you can find two different templates to use for creating your own plugin.
+Within our CellProfiler wiki, you can find an [orientation to CellProfiler code](https://github.com/CellProfiler/CellProfiler/wiki/Orientation-to-CellProfiler-code).
+
+In the CellProfiler repository, within [cellprofiler/modules/plugins](https://github.com/CellProfiler/CellProfiler/tree/master/cellprofiler/modules/plugins) you can find two different templates to use for creating your own plugin.
`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
@@ -8,10 +10,17 @@ In you plugin, you must include:
In your plugin, we appreciate if you also include:
- display functionality
+- extensive module documentation
+- references and citation information in your module documentation
+Please create a Pull Request to the CellProfiler-plugins repository to submit your plugin for inclusion in the repository.
-In your PR, you must include:
+In your PR, you must:
- add your plugin to the [supported_plugins](supported_plugins.md) documentation page
In your PR, we appreciate if you also include:
-- unit tests for your plugin
\ No newline at end of file
+- unit tests for your plugin
+
+## Having your plugin cited
+
+While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/overview.md b/documentation/CP-plugins-documentation/overview.md
index 5ece4c8d..1616e1a3 100644
--- a/documentation/CP-plugins-documentation/overview.md
+++ b/documentation/CP-plugins-documentation/overview.md
@@ -9,6 +9,17 @@ A module may be in CellProfiler-plugins instead of CellProfiler itself because:
- it requires extra libraries or other dependencies we are unable or unwilling to require for CellProfiler
- it has been contributed by a community member
+## How do I use CellProfiler Plugins?
+
+See our [Using Plugins](using_plugins.md) page for information on how to install and use plugins.
+
+## What plugins are available?
+
+See our [Supported Plugins](supported_plugins.md) page for information on all currently supported plugins.
+
+## How do I contribute a plugin?
+
+See our [Contributing Plugins](contributing_plugins.md) page for information on contributing a plugin to CellPainting-plugins.
## Who made this?
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index 5850822a..deb60d61 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -3,61 +3,22 @@
Below is a brief overview of our currently supported plugins.
For details about using any particular plugin, please read the module documentation inside the plugin in CellProfiler.
-## CalculateMoments
-CalculateMoments extracts moments statistics from a given distribution of pixel values.
-
-## CallBarcodes
-CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles.
-It is used for optical sequencing by synthesis (SBS).
-
-## CompensateColors
-CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through.
-It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched.
-
-## DeclumpObjects
-DeclumpObjects will split objects based on a seeded watershed method.
-
-## DistanceTransform
-DistanceTransform computes the distance transform of a binary image.
-The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1.
-
-## EnhancedMeasureTexture
-EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler.
-
-## HistogramEqualization
-HistogramEqualization increases the global contrast of a low-contrast image or volume.
-Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct.
-This module can perform either global or local histogram equalization.
-
-## HistogramMatching
-HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image.
-It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image.
-It allows you to choose which frame to use as the reference.
-
-## PixelShuffle
-PixelShuffle takes the intensity of each pixel in an image and it randomly shuffles its position.
-
-## Predict
-Predict allows you to use an ilastik pixel classifier to generate a probability image.
-CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage).
-
-## RunCellpose
-RunCellpose allows you to run Cellpose within CellProfiler.
-Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells.
-You can use pre-trained Cellpose models or your custom model with this plugin.
-You can use a GPU with this module to dramatically increase your speed/efficiency.
-
-## RunImageJScript
-
-## RunOmnipose
-
-## RunStarDist
-RunStarDist allows you to run StarDist within CellProfiler.
-StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells.
-You can use pre-trained StarDist models or your custom model with this plugin.
-You can use a GPU with this module to dramatically increase your speed/efficiency.
-RunStarDist is generally faster than RunCellpose.
-
-## VarianceTransform
-This module allows you to calculate the variance of an image, using a determined window size.
-It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image.
\ No newline at end of file
+Most plugins will run without any special installation of either CellProfiler or the plugins.
+See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin use as well as for installation information for those plugins that do require installation of dependencies.
+
+| Plugin | Description | Requires installation of dependencies? | Install flag |
+|--------|-------------|----------------------------------------|--------------|
+| CalculateMoments | CalculateMoments extracts moments statistics from a given distribution of pixel values. | No | |
+| CallBarcodes | CallBarcodes is used for assigning a barcode to an object based on the channel with the strongest intensity for a given number of cycles. It is used for optical sequencing by synthesis (SBS). | No | |
+| CompensateColors | CompensateColors determines how much signal in any given channel is because of bleed-through from another channel and removes the bleed-through. It can be performed across an image or masked to objects and provides a number of preprocessing and rescaling options to allow for troubleshooting if input image intensities are not well matched. | No | |
+| DistanceTransform | DistanceTransform computes the distance transform of a binary image. The distance of each foreground pixel is computed to the nearest background pixel and the resulting image is then scaled so that the largest distance is 1. | No | |
+| EnhancedMeasureTexture| EnhancedMeasureTexture measures the degree and nature of textures within an image or objects in a more comprehensive/tuneable manner than the MeasureTexture module native to CellProfiler. | No | |
+| HistogramEqualization | HistogramEqualization increases the global contrast of a low-contrast image or volume. Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct. This module can perform either global or local histogram equalization. | No | |
+| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
+| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
+| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | Yes | XXXXX |
+| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
+| RunImageJScript | | Yes | XXXXX |
+| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
+| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
+| VarianceTransform | This module allows you to calculate the variance of an image, using a determined window size. It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image. | No | |
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/using_GPU.md b/documentation/CP-plugins-documentation/using_GPU.md
index 945d869b..b5dfc4d1 100644
--- a/documentation/CP-plugins-documentation/using_GPU.md
+++ b/documentation/CP-plugins-documentation/using_GPU.md
@@ -24,3 +24,11 @@ If you want to use a GPU to run the model (this is recommended for speed), you'l
pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
```
If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
+
+
+ **NOTE**: You might get a warning like this:
+ ```
+ W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
+ 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
+ ```
+ If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
From 12c52c77ec80879b399743160d6accbfe2049d26 Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 12:35:02 -0700
Subject: [PATCH 4/6] more docs
---
.../contributing_plugins.md | 3 +-
.../supported_plugins.md | 4 +-
.../unsupported_plugins.md | 4 +-
.../CP-plugins-documentation/using_plugins.md | 106 ++++++++++++++++--
.../CP-plugins-documentation/versions.md | 49 +-------
5 files changed, 104 insertions(+), 62 deletions(-)
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index c3ae78be..f0084668 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -6,8 +6,6 @@ In the CellProfiler repository, within [cellprofiler/modules/plugins](https://gi
`imagetemplate.py` provides a template that takes one image as an input and produces a second image for downstream processing.
`measurementtemplate.py` provides a template that measures a property of an image both for the image as a whole and for every object in the image.
-In you plugin, you must include:
-
In your plugin, we appreciate if you also include:
- display functionality
- extensive module documentation
@@ -20,6 +18,7 @@ In your PR, you must:
In your PR, we appreciate if you also include:
- unit tests for your plugin
+- customized installation in setup.py, if your plugin has dependencies
## Having your plugin cited
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index deb60d61..a17866c0 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -16,9 +16,9 @@ See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin
| HistogramEqualization | HistogramEqualization increases the global contrast of a low-contrast image or volume. Histogram equalization redistributes intensities to utilize the full range of intensities, such that the most common frequencies are more distinct. This module can perform either global or local histogram equalization. | No | |
| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
-| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | Yes | XXXXX |
+| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | No | |
| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
-| RunImageJScript | | Yes | XXXXX |
+| RunImageJScript | RunImageJScript allows you to run any supported ImageJ script directly within CellProfiler. It is significantly more performant than RunImageJMacro, and is also less likely to leave behind temporary files. | Yes | XXXXX |
| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
| VarianceTransform | This module allows you to calculate the variance of an image, using a determined window size. It also has the option to find the optimal window size from a predetermined range to obtain the maximum variance of an image. | No | |
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index 4ca10a6c..b7cf3fe8 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -5,11 +5,9 @@ Unsupported plugins are primarily unsupported for either of two reasons:
- Their functions have been integrated into the current version of CellProfiler and therefore a plugin is no longer necessary
We welcome requests for updating particular unsupported plugins, but please note that we have limited bandwidth for working on plugins and may be unable to complete the update.
+Additionally, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
We welcome community contributed plugin updates.
-Likewise, we welcome contributions of community developed plugins.
-Please note that, because of our limited bandwidth, we cannot commit to maintaining any given plugin, CellProfiler team- or community-contributed.
-
## Where are unsupported plugins?
Unsupported plugins can be found in the `unmaintained_plugins` folder in the CellProfiler-plugins repository.
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index 0020d62b..bce4bc70 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -8,9 +8,16 @@ Please report any installation issues or bugs related to plugins in the [CellPro
## Installation
-### Installing plugins without dependencies
+If the plugin you would like to use does not have any additional dependencies outside of those required for running CellProfiler (this is most plugins), using plugins is very simple.
+See [Installing plugins without dependencies](#installing-plugins-without-dependencies).
+
+If the plugin you would like to use has dependencies, you have two separate options for installation.
+The first option requires building CellProfiler from source, but plugin installation is simpler.
+See [Installing plugins with dependencies, using CellProfiler from source](#installing-plugins-with-dependencies-using-cellprofiler-from-source).
+The second option allows you to use pre-built CellProfiler, but plugin installation is more complex.
+See [Installing plugins with dependencies, using pre-built CellProfiler](#installing-plugins-with-dependencies-using-pre-built-cellprofiler).
-If your plugin doesn't require additional dependencies, all you need to do is
+### Installing plugins without dependencies
1. **Install CellProfiler.**
Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
@@ -31,12 +38,12 @@ Alternatively, if you have code for plugins that are not in the CellProfiler-plu
- Open CellProfiler.
- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
- Select `Save` at the bottom of the Preferences window
-- Close CellProfiler and re-open it.
+- Close CellProfiler and re-open it
You are now ready to use any CellProfiler plugin that does not have additional dependencies.
This is most CellProfiler plugins.
-### Installing plugins with dependencies
+### Installing plugins with dependencies, using CellProfiler from source
1. **Install CellProfiler.**
Install CellProfiler from source (See instructions for: [Windows](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Windows%29); [Mac Intel](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28OS-X-and-macOS%29); [Mac Apple Silicon](https://github.com/CellProfiler/CellProfiler/wiki/Installation-of-CellProfiler-4-from-source-on-MacOS-M1); [Linux](https://github.com/CellProfiler/CellProfiler/wiki/Source-installation-%28Linux%29))
@@ -49,9 +56,9 @@ git clone https://github.com/CellProfiler/CellProfiler-plugins.git
3. **Set the plugins path in CellProfiler.**
- Open CellProfiler.
-- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned (or, if you didn't clone the whole repository, whatever location you have saved your plugins into).
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned.
- Select `Save` at the bottom of the Preferences window
-- Close CellProfiler and re-open it.
+- Close CellProfiler
4. **Install the dependencies for your plugin.**
In your terminal, type the following, where FLAG is the flag specific to the module you would like to run, as noted in [Supported Plugins](supported_plugins.md).
@@ -59,6 +66,91 @@ In your terminal, type the following, where FLAG is the flag specific to the mod
cd CellProfiler-plugins
pip install -e .[FLAG]
```
+e.g. To install Cellpose the pip install command would be `pip install -e .[cellpose]`
+
+5. **Open and use CellProfiler.**
Please note that plugins that have separate install flags may have conflicting dependencies so we recommend making a separate python environment in which to run separate installations.
-(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
\ No newline at end of file
+(e.g. while having CellPose and StarDist in the same python environment is technically possible, it has been reported to be quite troublesome to install, so we recommend choosing either CellPose or StarDist.)
+
+### Installing plugins with dependencies, using pre-built CellProfiler
+
+1. **Install CellProfiler.**
+Download a binary (pre-built) version of CellProfiler from the [CellProfiler website](https://cellprofiler.org/releases).
+
+2. **Clone the CellProfiler-plugins repository.**
+In your terminal, type
+```bash
+git clone https://github.com/CellProfiler/CellProfiler-plugins.git
+```
+
+3. **Set the plugins path in CellProfiler.**
+- Open CellProfiler.
+- Go to `CellProfiler` => `Preferences` and set the path in the `CellProfiler plugins directory` to the GitHub repository that you just cloned.
+- Select `Save` at the bottom of the Preferences window
+- Close CellProfiler
+
+4. **Identify the Python dependencies of the plugins you want to use.**
+You can find this information for most plugins by looking in `setup.py`.
+```{admonition} e.g. using RunCellpose plugin
+If you would like to install the RunCellpose plugin, in `setup.py` you can see that `cellpose_deps = ["cellpose>=1.0.2"]` so the only dependency is `cellpose`.
+```
+
+Alternatively, you can find this information directly in the plugin code itself by looking at what is imported at the beginning of the plugin.
+Note that you will want to compare any imports to what is already required by CellProfiler with that caveat that dependencies often have dependencies.
+```{admonition} e.g. using RunImageJScript plugin
+If you would like to install the RunImageJScript plugin, you'll notice a number of imports in the beginning of the plugin code.
+You can ignore all those from `cellprofiler_core` as they are required for CellProfiler installation.
+You can ignore all those that are native to Python such as `sys`, `time`, and `threading`.
+A comparison with CellProfiler dependencies will show you that `skimage` is already installed (you can see this in CellProfiler's [environment.yml](https://github.com/CellProfiler/CellProfiler/blob/master/environment.yml) which lists `scikit-image` as a `cellprofiler-core` dependency or in CellProfiler's [setup.py](https://github.com/CellProfiler/CellProfiler/blob/master/setup.py) which lists `scikit-image` as an installation requirement).
+What remains is `pyimagej`
+```
+
+5. **Create a conda environment and install requirements in it**
+Create a conda (or other virtual) environment with a Python version matching that of CellProfiler.
+CellProfiler 4 uses Python 3.8 (which you can see in CellProfiler's [setup.py](https://github.com/CellProfiler/CellProfiler/blob/master/setup.py)).
+Activate the environment and install the plugin requirement/s into it.
+
+In your terminal, type
+```
+# Create a conda environment
+conda create --name ENV_NAME python=3.8
+# Activate the conda environment
+conda activate ENV_NAME
+# Install the plugin requirement/s
+pip install REQUIREMENT
+```
+```{admonition} e.g. using RunImageJScript plugin
+conda create --name cp-ij python=3.8
+conda activate cp-ij
+pip install pyimagej
+```
+
+6. **Copy plugin requirements into CellProfiler installation**
+Find the folder that you installed into in the previous step (where to find these libraries in your local environment) by entering `pip show REQUIREMENT` into your terminal e.g. `pip show pyimagej`.
+In the information it returns, under `Location` you will find a path that will look something like `/Users/eweisbar/mambaforge/envs/cp-ij/lib/python3.8/site-packages`.
+
+Find the folder in which you have installed CellProfiler.
+On Windows this is likely to be `C:\Program Files\CellProfiler` and on Mac it is likely to be `/Applications/CellProfiler.app/Contents/MacOS/`.
+In your terminal, if you `ls` (for Mac) or `dir` (for Windows) that folder (e.g. `ls /Applications/CellProfiler.app/Contents/MacOS/`), you should see a list of all the dependencies that were packaged with CellProfiler.
+
+Copy the folders and their corresponding `.dist-info` folders (e.g. `pytz` and `pytz-2023.3.dist-info`) for any dependencies that were installed with the installation of `pyimagej` that are not already in the CellProfiler folder.
+
+```{admonition} e.g. using RunImageJScript plugin
+These are all the folders you need to copy over:
+`pytz`
+`pytz-2023.3.dist-info`
+`xarray`
+`xarray-2023.1.0.dist-info`
+`imglyb`
+`imglyb-2.1.0.dist-info`
+`jgo`
+`jgo-1.0.5.dist-info`
+`jpype`
+`_jpype.cpython-38-darwin.so`
+`imagej`
+`pyimagej-1.4.1.dist-info`
+```
+
+7. **Open and use CellProfiler.**
+When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/versions.md b/documentation/CP-plugins-documentation/versions.md
index 3755e75b..0f8db604 100644
--- a/documentation/CP-plugins-documentation/versions.md
+++ b/documentation/CP-plugins-documentation/versions.md
@@ -1,55 +1,8 @@
# Versions
-The most current release can always be found on DockerHub at `cellprofiler/distributed-cellprofiler`.
-Current version is 2.0.0.
-Our current tag system is dcpversion_CellProfilerversion, e.g. 1.2.0_3.1.8
-
-Previous release versions can be accessed at `bethcimini/distributed-cellprofiler:versionnumber`
+Current version is
---
# Version History
-## 2.1.0 (forthcoming)
-* Increase support for using plugins
-* Improve role handling and allow file download/upload from/to S3 buckets in different accounts
-* Improve file download to S3FS can be completely bypassed
-* Name EBS volumes
-
-## 2.0.0rc2 - Released 20201110
-* Add optional ability to download files to EBS rather than reading from S3 (helpful for pipelines that access many files/image sets)
-
-## 2.0.0rc1 - Released 20201105
-
-* Remove requirement for boto and fabric, using only boto3
-* Add support for Python 3 and CellProfiler 4
-* Move cluster size, machine type, and machine price to the config file from the fleet file, eliminating mismatches between the two
-* Add the ability to filter only files with certain names when running CHECK_IF_DONE
-* Don't cancel a fleet over capacity errors
-* Add "cheapest" mode to the monitor, allowing you to run more cheaply (at possible expense of running more slowly)
-
-## 1.2.2 - Released 20201103
-
-* Allow pipelines using batch files to also designate an input output_top_directory
-* Add support for multiple LaunchData specifications
-* Add CellProfiler-plugins
-* Additional way to create job submissions with run_batch_general.py
-
-## 1.2.1 - Released 20200109, Updated through 20191002
-
-* Allow monitor to downscale machines when number of jobs < number of machines
-* Add a parameter to discount files when running CHECK_IF_DONE checks if less than a certain size
-
-## 1.2.0 - Released 20181108, Updated through 20200109
-
-* Improved compatibility with CellProfiler 2 and 3
-* Better handling of logging when using output_structure
-
-## 1.1.0 - Released 20170217, Updated 20170221 (bugfixes) - 20181018
-
-* Changes in this release:
-
- * Added the `output_structure` variable to the job file, which allows you to structure the output folders created by DCP (ie `Plate/Well-Site` rather than `Plate-Well-Site`). Job files lacking this variable will still default to the previous settings (hyphenating all Metadata items in order they are presented in the Metadata grouping).
- * Added support for creating the list of groups via `cellprofiler --print-groups`- see [this issue](https://github.com/CellProfiler/Distributed-CellProfiler/issues/52) for example and discussion. Groups listed in this way MUST use the `output_structure` variable to state their desired folder structure or an error will be returned.
-
-## 1.0.0 - Version as of 20170213
From fffb5566c3b4d2892c75c30b14469e66e94bca6a Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 15:04:09 -0700
Subject: [PATCH 5/6] more doc changes
---
.../CP-plugins-documentation/RunCellPose.md | 34 +++++++++++++++++++
.../contributing_plugins.md | 9 ++++-
.../supported_plugins.md | 5 ++-
.../unsupported_plugins.md | 1 +
.../CP-plugins-documentation/using_GPU.md | 34 -------------------
.../CP-plugins-documentation/using_plugins.md | 3 +-
6 files changed, 49 insertions(+), 37 deletions(-)
create mode 100644 documentation/CP-plugins-documentation/RunCellPose.md
delete mode 100644 documentation/CP-plugins-documentation/using_GPU.md
diff --git a/documentation/CP-plugins-documentation/RunCellPose.md b/documentation/CP-plugins-documentation/RunCellPose.md
new file mode 100644
index 00000000..168fa7cc
--- /dev/null
+++ b/documentation/CP-plugins-documentation/RunCellPose.md
@@ -0,0 +1,34 @@
+# RunCellPose
+
+## Using RunCellPose with a GPU
+
+If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU.
+General instructions are available at this [link](https://pytorch.org/get-started/locally/).
+
+1. Your GPU should be visible in Device Manager under Display Adaptors.
+If your GPU isn't there, you likely need to install drivers.
+[Here](https://www.nvidia.com/Download/Find.aspx) is where you can find NVIDIA GPU drivers if you need to install them.
+
+
+2. To test whether the GPU is configured correctly:
+ * Run `python` on the command line (i.e., in Command Prompt or Terminal) to start an interactive session
+ * Then run the following
+ ```
+ import torch
+ torch.cuda.is_available()
+ ```
+ * If this returns `True`, you're all set
+ * If this returns `False`, you likely need to install/reinstall torch. See [here](https://pytorch.org/get-started/locally/) for your exact command.
+ * Exit the session with `exit()` then install torch if necessary
+ ```
+ pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
+ ```
+ If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
+
+
+**NOTE**: You might get a warning like this:
+```
+W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
+2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
+```
+If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/contributing_plugins.md b/documentation/CP-plugins-documentation/contributing_plugins.md
index f0084668..8907b250 100644
--- a/documentation/CP-plugins-documentation/contributing_plugins.md
+++ b/documentation/CP-plugins-documentation/contributing_plugins.md
@@ -20,6 +20,13 @@ In your PR, we appreciate if you also include:
- unit tests for your plugin
- customized installation in setup.py, if your plugin has dependencies
+## Contributing bug fixes or updating deprecated plugins
+
+Please create a Pull Request to the CellProfiler-plugins repository to submit a bug fix or plugin un-deprecation.
+If you would like to update a currently deprecated plugin, we encourage you to read closed Issues in GitHub relevant to the plugin as they may contain helpful information about bugs present at the time of deprecation.
+
## Having your plugin cited
-While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
\ No newline at end of file
+While we cannot guarantee that users will cite your plugin, we have introduced a Citation generator into CellProfiler v? that scans all modules in a user's pipeline and generators a citation file for them that includes citation information for any modules (including plugins) that have specific citation information in them.
+
+See [Citing Plugins](citing.md) for information on citing CellProfiler Plugins.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/supported_plugins.md b/documentation/CP-plugins-documentation/supported_plugins.md
index a17866c0..5ccdda93 100644
--- a/documentation/CP-plugins-documentation/supported_plugins.md
+++ b/documentation/CP-plugins-documentation/supported_plugins.md
@@ -6,6 +6,9 @@ For details about using any particular plugin, please read the module documentat
Most plugins will run without any special installation of either CellProfiler or the plugins.
See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin use as well as for installation information for those plugins that do require installation of dependencies.
+Most plugin documentation can be found within the plugin itself and can be accessed through CellProfiler help.
+Those plugins that do have extra documentation contain links below.
+
| Plugin | Description | Requires installation of dependencies? | Install flag |
|--------|-------------|----------------------------------------|--------------|
| CalculateMoments | CalculateMoments extracts moments statistics from a given distribution of pixel values. | No | |
@@ -17,7 +20,7 @@ See [using plugins](using_plugins.md) for how to set up CellProfiler for plugin
| HistogramMatching | HistogramMatching manipulates the pixel intensity values an input image and matches them to the histogram of a reference image. It can be used as a way to normalize intensities across different 2D or 3D images or different frames of the same 3D image. It allows you to choose which frame to use as the reference. | No | |
| PixelShuffle | PixelShuffle takes the intensity of each pixel in an image and randomly shuffles its position. | No | |
| Predict | Predict allows you to use an ilastik pixel classifier to generate a probability image. CellProfiler supports two types of ilastik projects: Pixel Classification and Autocontext (2-stage). | No | |
-| RunCellpose | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
+| [RunCellpose](RunCellPose.md) | RunCellpose allows you to run Cellpose within CellProfiler. Cellpose is a generalist machine-learning algorithm for cellular segmentation and is a great starting point for segmenting non-round cells. You can use pre-trained Cellpose models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. | Yes | `cellpose` |
| RunImageJScript | RunImageJScript allows you to run any supported ImageJ script directly within CellProfiler. It is significantly more performant than RunImageJMacro, and is also less likely to leave behind temporary files. | Yes | XXXXX |
| RunOmnipose | RunOmnipose allows you to run Omnipose within CellProfiler. Omnipose is a general image segmentation tool that builds on Cellpose. | Yes | `omnipose` |
| RunStarDist | RunStarDist allows you to run StarDist within CellProfiler. StarDist is a machine-learning algorithm for object detection with star-convex shapes making it best suited for nuclei or round-ish cells. You can use pre-trained StarDist models or your custom model with this plugin. You can use a GPU with this module to dramatically increase your speed/efficiency. RunStarDist is generally faster than RunCellpose. | Yes | `stardist` |
diff --git a/documentation/CP-plugins-documentation/unsupported_plugins.md b/documentation/CP-plugins-documentation/unsupported_plugins.md
index b7cf3fe8..5e277b61 100644
--- a/documentation/CP-plugins-documentation/unsupported_plugins.md
+++ b/documentation/CP-plugins-documentation/unsupported_plugins.md
@@ -19,4 +19,5 @@ Those plugins in the `CellProfiler4_autoconverted` folder were automatically con
We cannot provide comprehensive information about why we are not supporting a given plugin.
Information about select plugins is as follows:
+**ClassifyPixelsUNET**: ClassifyPixelsUNET is a pixel classifier for background/object edge/object body. As far as we are aware, other deep learning based plugins that we do currently support (such as RunCellpose) work better.
**DeclumpObjects**: DeclumpObjects will split objects based on a seeded watershed method. Functionality from this module was [added into CellProfiler](https://github.com/CellProfiler/CellProfiler/pull/4397) in the Watershed module as of CellProfiler 4.2.0.
diff --git a/documentation/CP-plugins-documentation/using_GPU.md b/documentation/CP-plugins-documentation/using_GPU.md
deleted file mode 100644
index b5dfc4d1..00000000
--- a/documentation/CP-plugins-documentation/using_GPU.md
+++ /dev/null
@@ -1,34 +0,0 @@
-# Using a GPU with CellProfiler Plugins
-
-These CellProfiler modules currently support using a GPU: LIST MODULES
-
-## Tips for Setting Up Your GPU (if you have one)
-
-If you want to use a GPU to run the model (this is recommended for speed), you'll need a compatible version of PyTorch and a supported GPU. General instructions are available at this [link](https://pytorch.org/get-started/locally/).
-
-1. Your GPU should be visible in Device Manager under Display Adaptors. If your GPU isn't there, you likely need to install drivers.
- * [Here](https://www.nvidia.com/Download/Find.aspx) is where you can find NVIDIA GPU drivers if you need to install them.
-
-
-2. To test whether the GPU is configured correctly:
- * Run `python` on the command line (i.e., in Command Prompt or Terminal) to start an interactive session
- * Then run the following
- ```
- import torch
- torch.cuda.is_available()
- ```
- * If this returns `True`, you're all set
- * If this returns `False`, you likely need to install/reinstall torch. See [here](https://pytorch.org/get-started/locally/) for your exact command.
- * Exit the session with `exit()` then install torch if necessary
- ```
- pip3 install torch torchvision torchaudio --extra-index-url https://download.pytorch.org/whl/cu113
- ```
- If you have a previous version of torch installed, make sure to run `pip uninstall torch` first.
-
-
- **NOTE**: You might get a warning like this:
- ```
- W tensorflow/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'cudart64_110.dll'; dlerror: cudart64_110.dll not found
- 2022-05-26 20:24:21.906286: I tensorflow/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
- ```
- If you don't have a GPU, this is not a problem. If you do, your configuration is incorrect and you need to try reinstalling drivers and the correct version of CUDA for your system.
\ No newline at end of file
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index bce4bc70..946b9ab5 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -153,4 +153,5 @@ These are all the folders you need to copy over:
```
7. **Open and use CellProfiler.**
-When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing.
\ No newline at end of file
+When you try to run your plugin in your pipeline, if you have missed copying over any specific requirements, it will give you an error message that will tell you what dependency is missing in the terminal window that opens with CellProfiler on Windows machines.
+This information is not available in Mac machines.
\ No newline at end of file
From f99b97e8c94b878f6e7a6059bd5f926f9901b41c Mon Sep 17 00:00:00 2001
From: Erin Weisbart <54687786+ErinWeisbart@users.noreply.github.com>
Date: Wed, 31 May 2023 15:09:02 -0700
Subject: [PATCH 6/6] docs
---
documentation/CP-plugins-documentation/using_plugins.md | 5 ++++-
1 file changed, 4 insertions(+), 1 deletion(-)
diff --git a/documentation/CP-plugins-documentation/using_plugins.md b/documentation/CP-plugins-documentation/using_plugins.md
index 946b9ab5..bab82fe8 100644
--- a/documentation/CP-plugins-documentation/using_plugins.md
+++ b/documentation/CP-plugins-documentation/using_plugins.md
@@ -1,7 +1,10 @@
# Using plugins
-Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all plugins in the plugin folder that you have set.
+Once you have installed your plugins and their dependencies (see below), CellProfiler will automatically detect all useable plugins in the plugin folder that you have set.
The plugins will appear in the "Add Modules" panel like all standard modules and you can use the plugins as you would any other CellProfiler module.
+Typically, if a plugin's dependencies are not installed, it will not be visible in the "Add Modules" panel.
+If you cannot find any plugins in the "Add Modules" panel then you have not properly set the plugins path (see below).
+If you can find some but not all plugins in the "Add Modules" panel then the not-visible plugins have unmet dependencies.
Please note that, as CellProfiler-plugins are considered experimental, they may not be as well documented as standard modules and they may not have a window that shows on run.
Please report any installation issues or bugs related to plugins in the [CellProfiler-plugins repository](https://github.com/CellProfiler/CellProfiler-plugins) and not in the main CellProfiler repository.