-
Notifications
You must be signed in to change notification settings - Fork 70
Rewrite MOC southern boundary extractor #181
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Rewrite MOC southern boundary extractor #181
Conversation
|
@milenaveneziani, I'm still testing this but figured I'd do a PR so I can post the results of my testing here. |
|
@milenaveneziani, I was able to verify that the transect on the southern boundary had the expected sign for all of the following meshes: I'm not sure what the procedure will be for testing the MOC with this new transect. Presumably waiting until @vanroekel is back? |
|
This is intended to address #164. |
|
@xylar: this is great. I can test with the MOC postprocessing first, similarly to what I did before. I will try with the v0.43 tag on titan's login node. |
|
sorry it took so long: titan is able to compute the high-res climos without ncclimo, but takes very long.. |
|
@milenaveneziani, okay. I'm disappointed that things still don't look "right" in your tests. I don't know how I can be more help unless you can put a set of files in my hands with which I can reproduce a case that looks "right" and another with my transect that doesn't. Is that something you can put together? |
|
Yes, I will do that. I will transfer 1 year of high-res data to NERSC, and then bring over the old transect file and the new one, for both the high-res and low-res cases. And yes, I was disappointed too: I really thought things were solved. Something tells me that we are close though. At least we don't get the weird structure with flipped signs.. |
|
@xylar: you can find year 11 and 30 of Luke's high-res run here at nersc: A low-res run could be this one: The newer MOC regions with transects mask files are these: The old ones are these: At OLCF, I have completed the MOC for year 11 using the new mask file, and I am now computing the same but using the old mask file. I shall see the results of this tomorrow morning since it takes about 2 hours to compute the 1-year climatologies on titan's login node. As a reminder you should check out v0.43 to do any test with the high-res data. Also, I have used Hopefully all permissions are OK. |
|
@milenaveneziani, thanks very much for making these available. I'm busy packing for the US and probably won't have a chance to look at this until Monday or Tuesday. But it's on my radar... |
|
@milenaveneziani Just to update, I was able to test those four mask files, each with a single monthly average and using the simple old python script for a single time. The old and new masks produce completely different results. I have another meeting now, but will update more soon. |
|
That's both disappointing and also maybe progress... Can we plot the edge sign in Paraview for both masks and see if they are consistent with one another, at least where they overlap? Other than the edge sign, I just don't know what could be wrong. |
Only if the evtk package is not found do we use the old pyevtk package. The evtk package is being built from evtk source as part of newer e3sm-unified packages and is on the e3sm channel. pyevtk does not seem to reliably support python 3 and some versions seem to have bugs in the setup script.
The latest netCDF4 package automatically masks all numpy arrays, which doesn't play nicely with pyevtk. To fix this, set_auto_mask(False) has been used when opening NetCDF files.
672ea9f to
81befe3
Compare
|
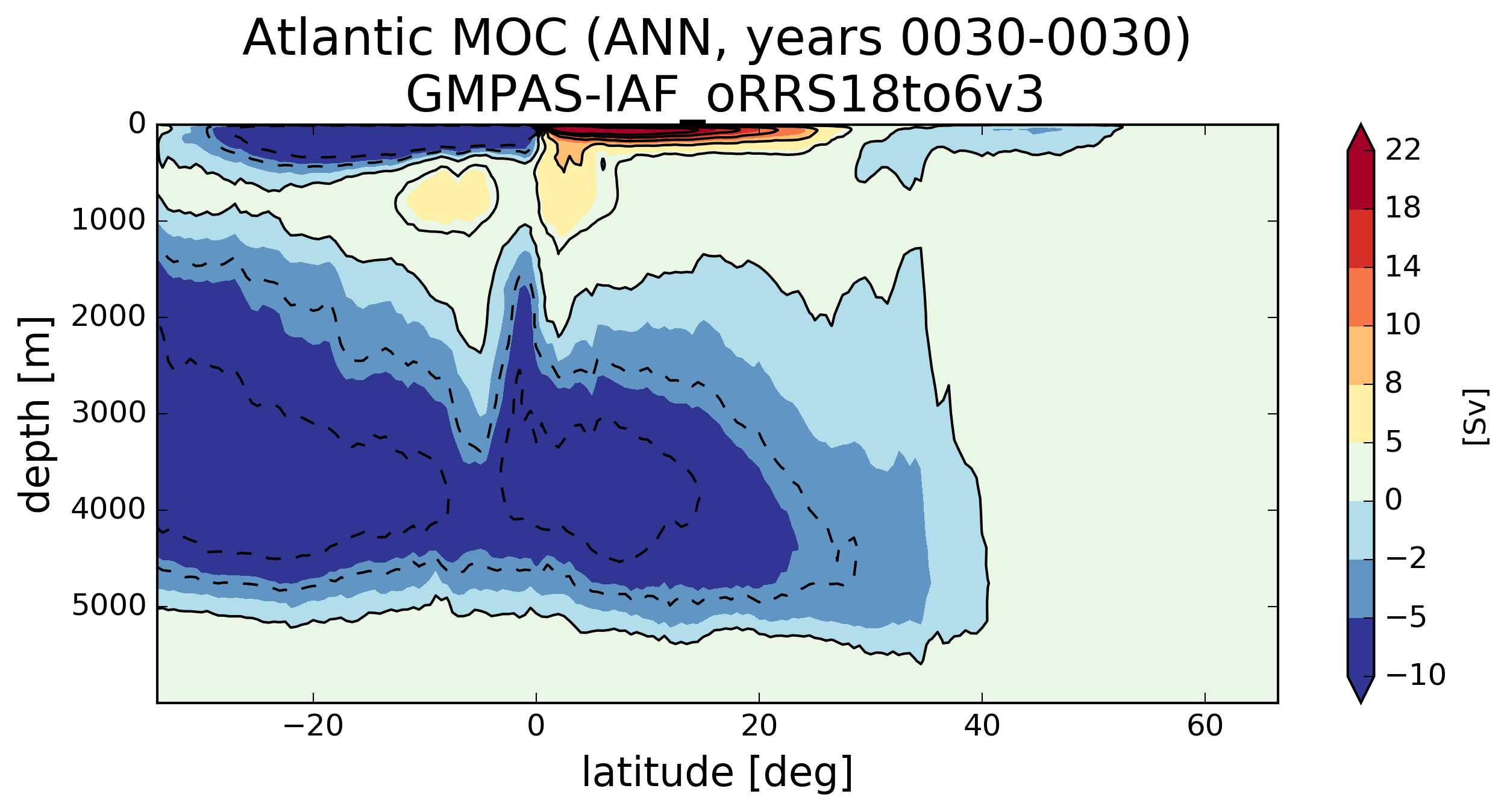
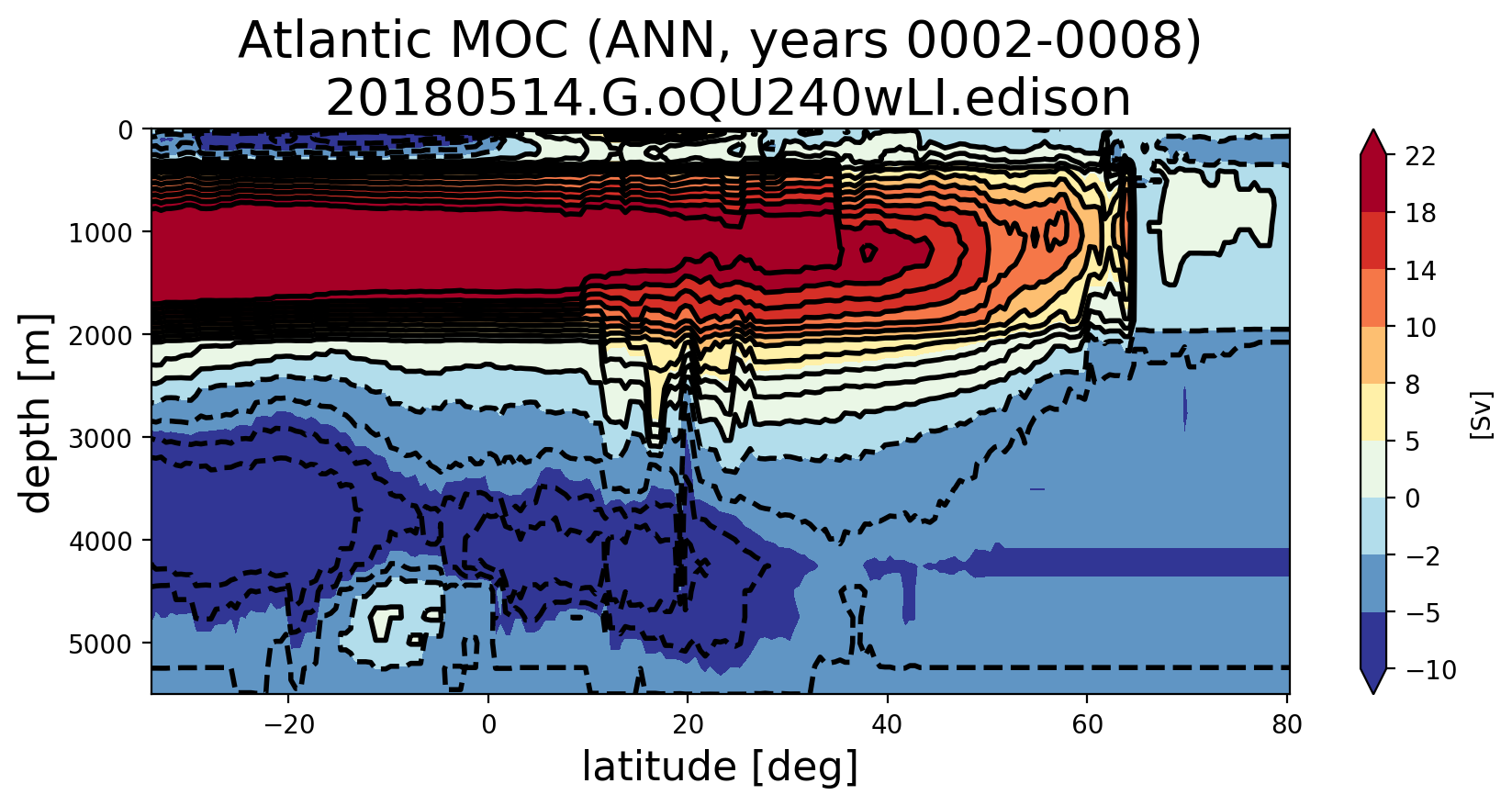
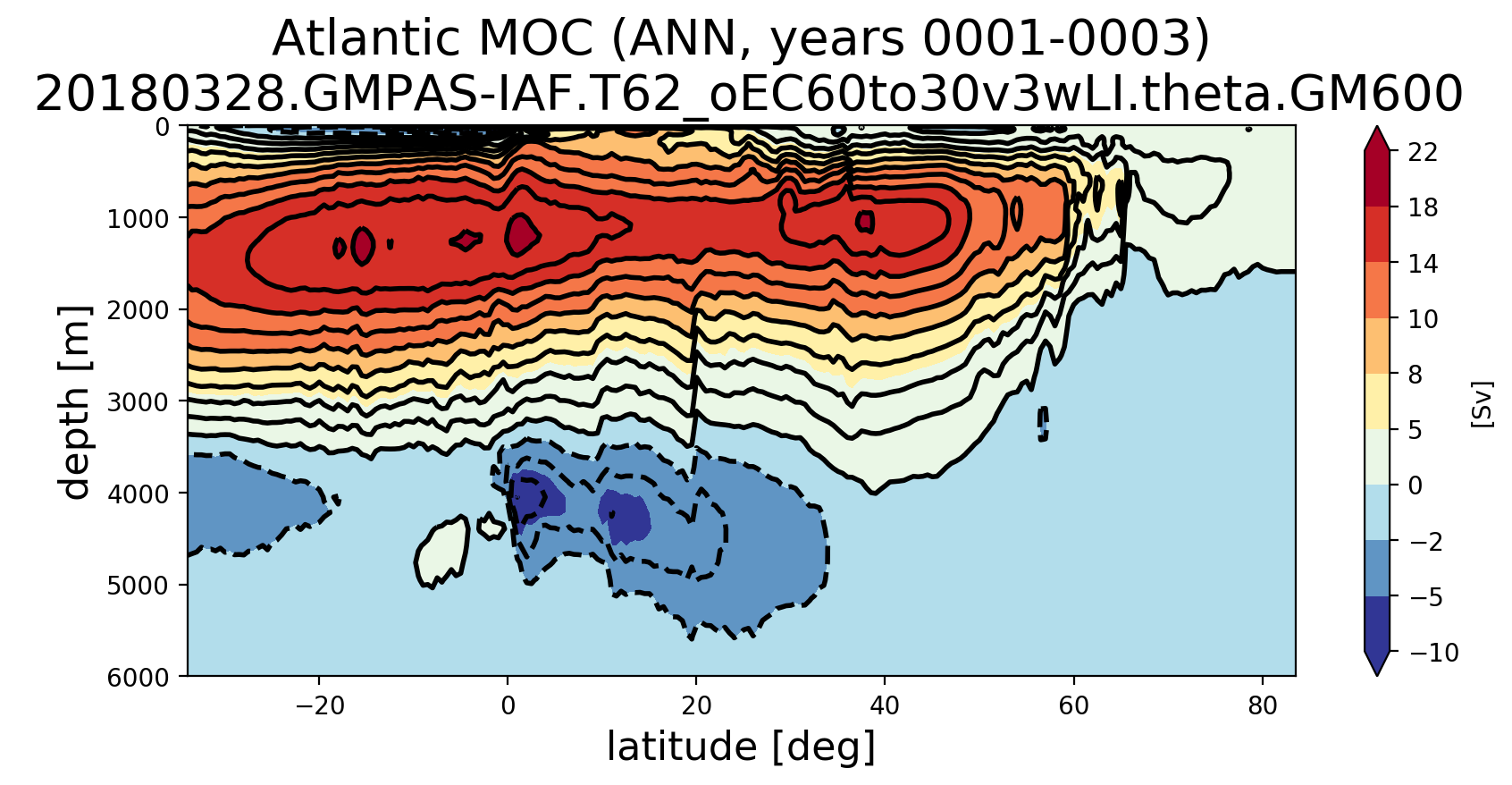
@milenaveneziani, I finally had time to get back to this. I believe a rewrite that I did earlier but didn't have a chance to fully test seems to be working. Here are Atlantic MOCs a very short time into runs at various resolutions: For comparison, here's are plots with the previous southern ocean transect. |
|
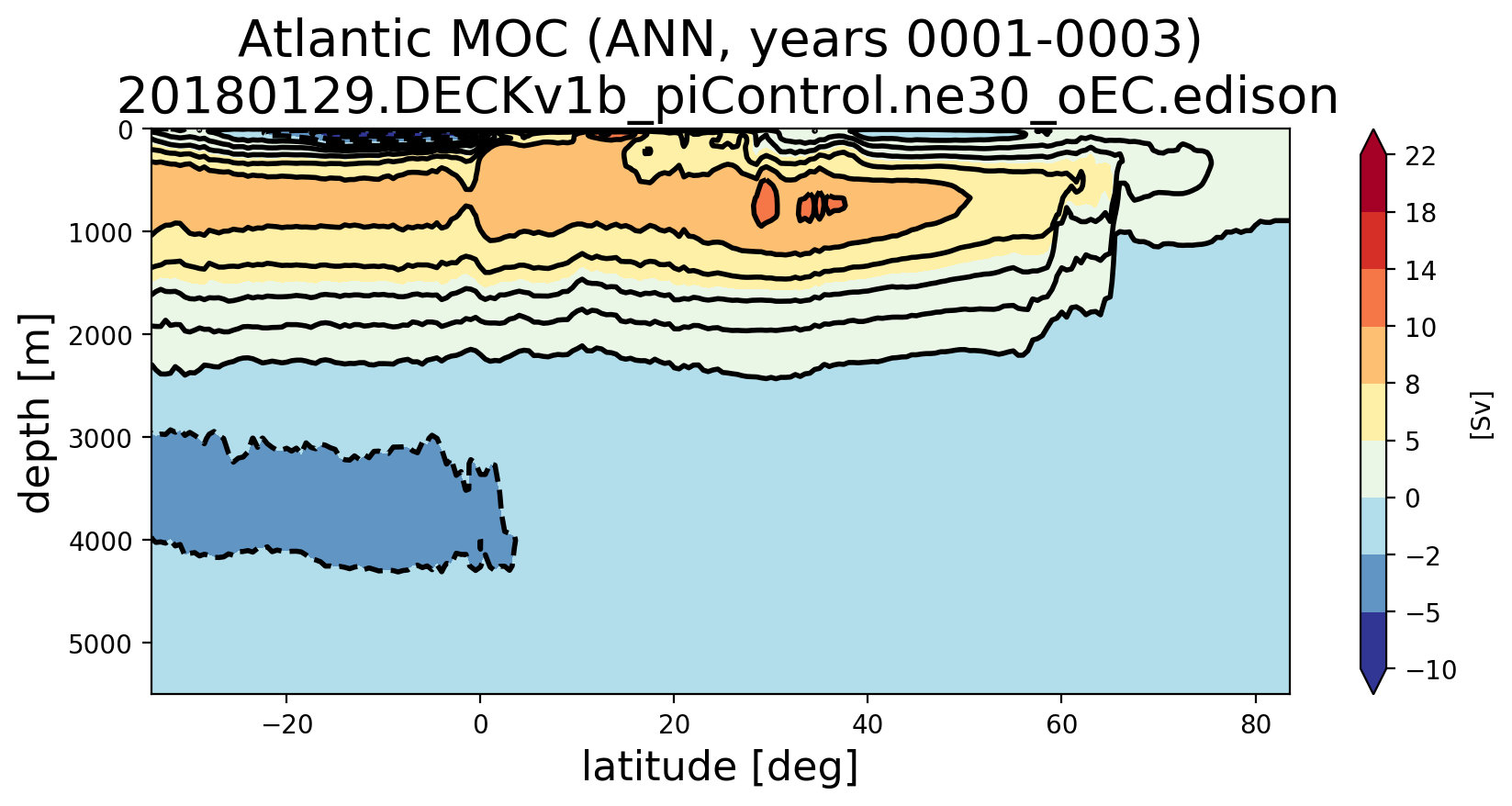
Here's the Atlantic MOC with the old transect: and new: from a different set of years from 20180129.DECKv1b_piControl.ne30_oEC.edison The other regional MOCs using the new analysis can be seen here: |
|
Here's the results from analyzing an RRS30to10 run: Here's the rest of the MOC plots for that same set of analysis: Here's the MOC over years 5-22 computed with the old transect : |
|
@milenaveneziani, @mark-petersen, @maltrud, @vanroekel Can you think of ways I can further test these new southern boundary transects to gain confidence? I did a test where I computed the horizontal transport into each cell and computed that the resulting vertical velocity was super tiny (0.0004 m/s or something). I then summed the transport from individual cells and proved to myself that was equal to the transport computed over a whole boundary of the MOC region (not just the southern boundary). So I'm quite confident that I have the edge signs right (finally). At a glance, all of the MOC plots look "not too crazy" to me in the sense that the MOC at the bottom of the ocean is not too far from zero and we're seeing similar values at EC60to30 resolution and higher between the old and new transects. However, we're seeing enough change that I want to make sure we're okay with this. It certainly won't help us explain why the AMOC is too weak, but it does seem to change a peak here and there in there in both the time series and the lat/depth plots. |
|
@xylar: @vanroekel noticed that the new transect file has the dimension |
|
@milenaveneziani, the bigger number is likely necessary to handle the metadata that is included in the files. I would think the better approach would be to modify the MOC AM to handle the longer string length. |
|
I agree that it would be nice to leave 1024, but, as you mentioned in the MPAS-Model PR, we could temporary change it to 64 and raise an issue in MPAS-Model to change this in the framework. |
Add bare-bones docs with Read The Docs
In such cases, we need to cd to the location of setup.py before finding package modules and scripts.
Fix conda package setup.py when called with a path
This was being done externally in scripts but it is easier to include it in setup.py. This also seems to be the only way to do this within Read The Docs, which doesn't support custom install scripts.
In conda package, copy directories within setup.py
Simple upgrade to python 3. Just print parenthesis and spacing.
Add some missing tests in conda recipe. Remove redundant listing of a script in setup.py
This is needed only for Read The Docs, which won't allow a build script.
Updates to the conda package for release of v0.0.4 Add SCRIP script to package fix some tests fix copying of scripts/code from outside of the package update to v0.0.4
This commit allows the script to use input arguments for the radius beyond which to cull and the fractional distance beyond which to cull. Previously those were both hard-coded values, which complicated using those methods within COMPASS.
…script Update define_cullMask.py to allow more input This merge allows the script to use input arguments for the radius beyond which to cull and the fractional distance beyond which to cull. Previously those were both hard-coded values, which complicated using those methods within COMPASS.
|
I believe the remaining issue with 1024-character strings (instead of the expected 64) should be addressed by #275, followed by a release of MPAS-Tools and an update of the various COMPASS conda environments. |
It now computes edge signs based on the order of cells on a given edge rather than of vertices, which may fix the issue seen previously.
This flag should not be used in nearly all standar cases.
|
@milenaveneziani, can we go ahead and merge this very old PR? Presumably any remaining issues can be addressed later. |
 milenaveneziani
left a comment
milenaveneziani
left a comment
There was a problem hiding this comment.
Choose a reason for hiding this comment
The reason will be displayed to describe this comment to others. Learn more.
Sure.
This merge fixes issues with the southern boundary transect extracted for MOC regions through a substantial rewrite. One significant change is that the edge sign is determined by the order of cells on an edge, not vertices (the latter of which may have caused issues with the previous approach). The logic has also been simplified.







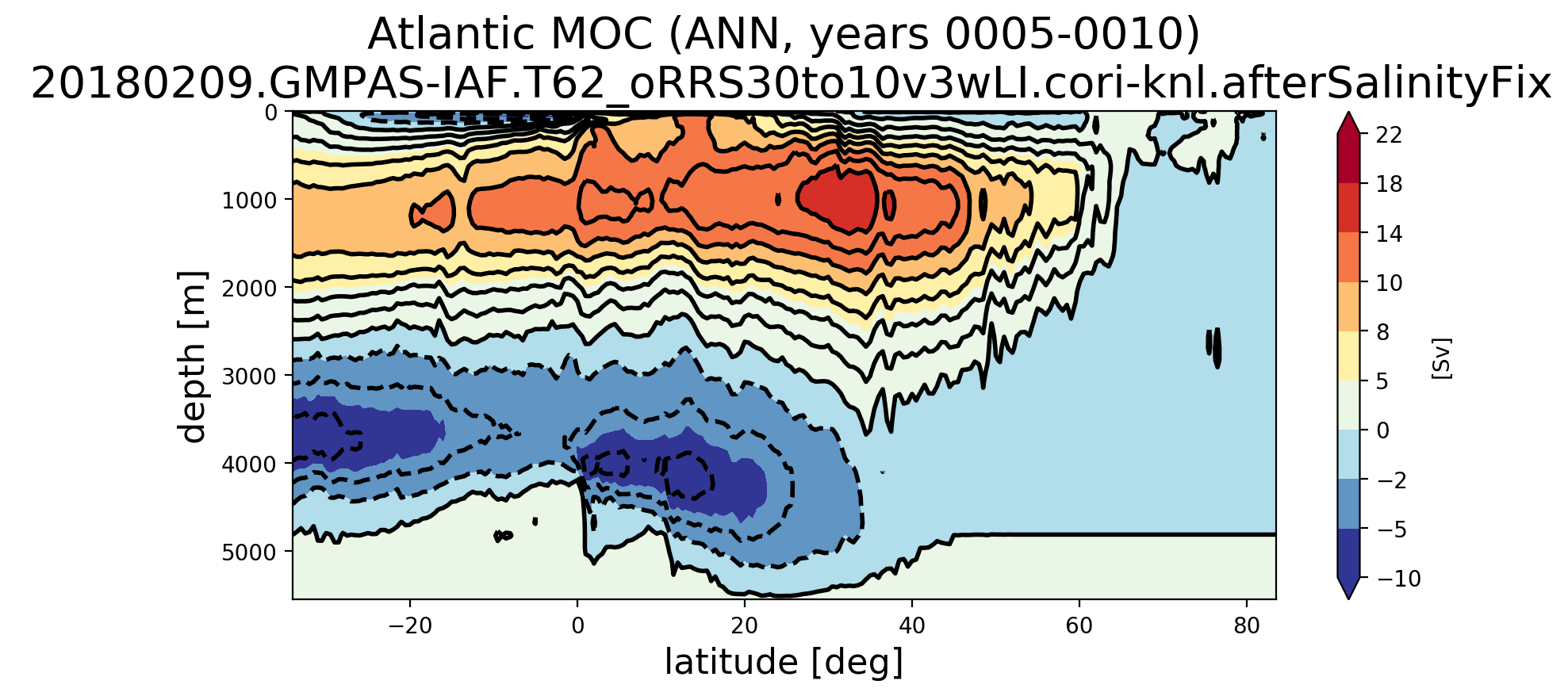
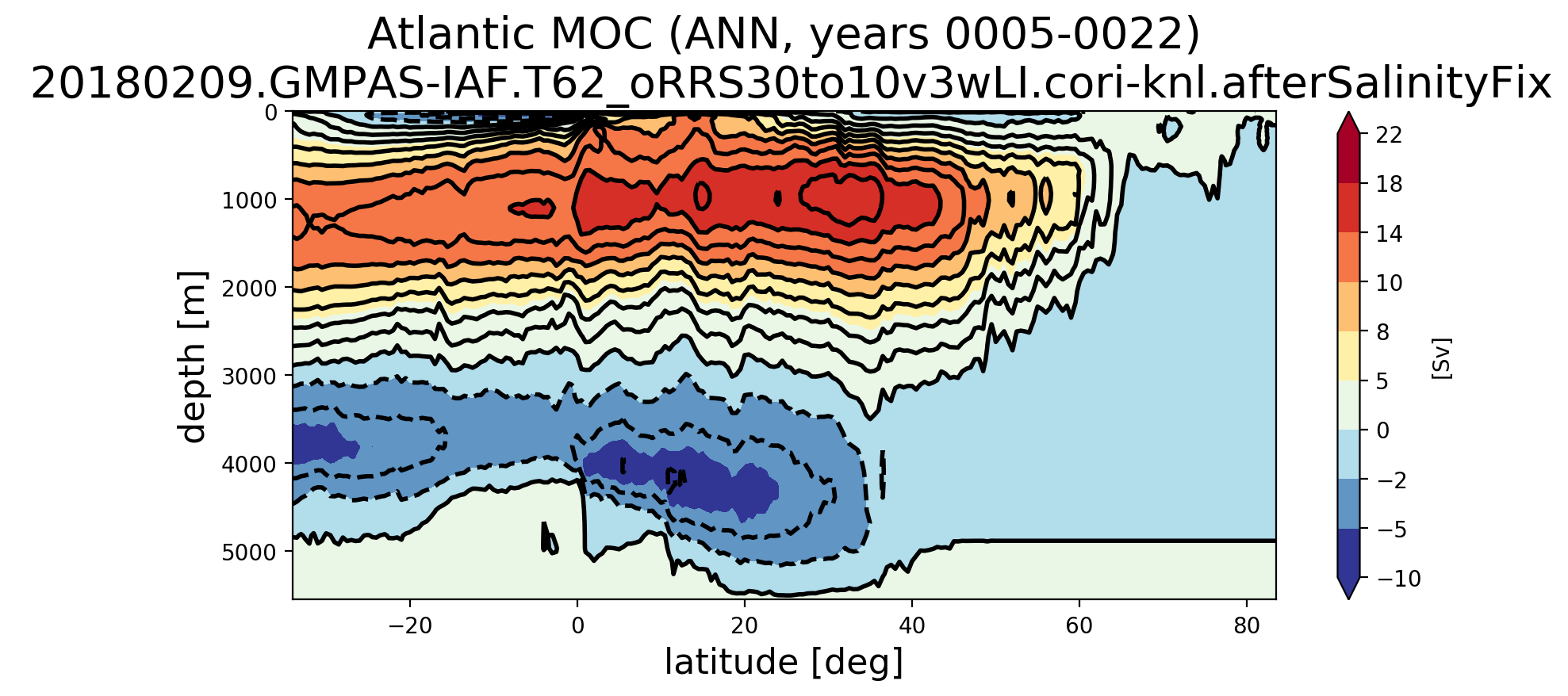

This merge fixes issues with the southern boundary transect extracted for MOC regions through a substantial rewrite. One significant change is that the edge sign is determined by the order of cells on an edge, not vertices (the latter of which may have caused issues with the previous approach). The logic has also been simplified.