-
Notifications
You must be signed in to change notification settings - Fork 217
Description
Hi,
I am not sure that bamCompare is doing exactly what I think it is doing so I wanted to check to see what I might be missing.
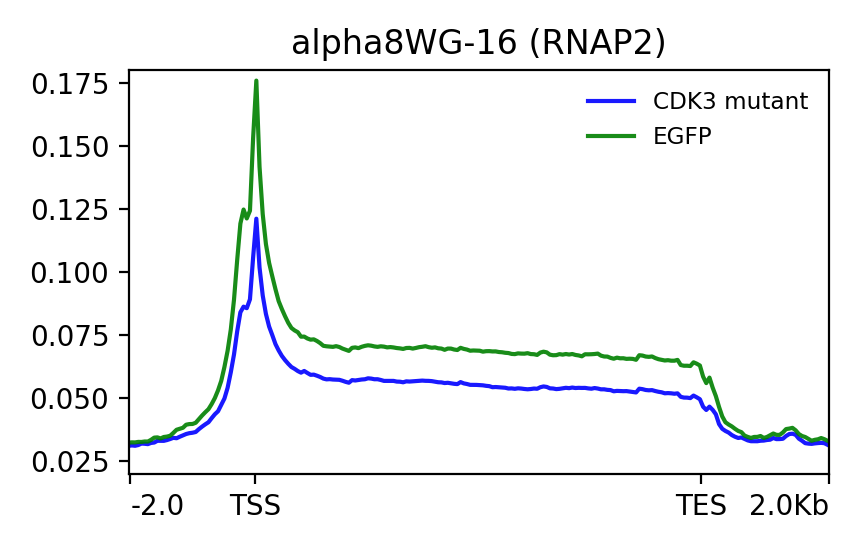
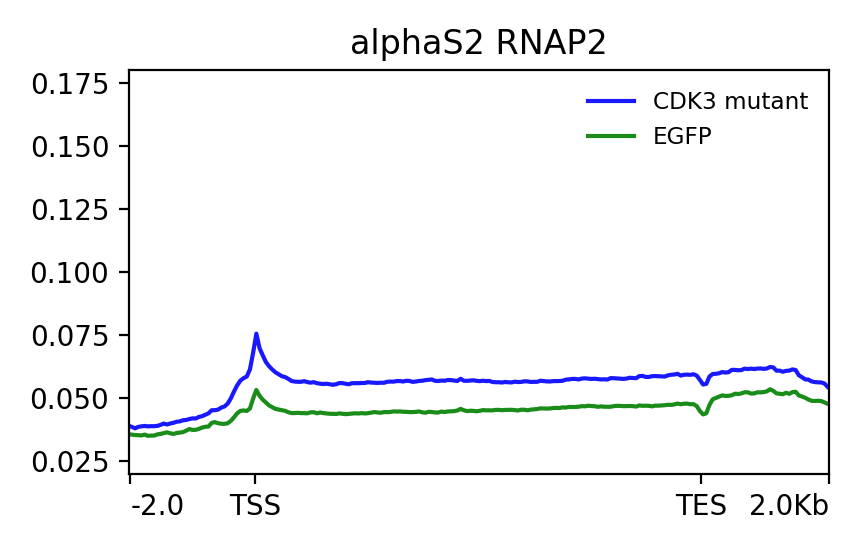
I have attached two profile plots that I have generated with deepTools and looks as expected to me. What I want to do is get the log2 ratio of blue line (2nd image divided by the first image) and the same for the green line. And then plot the two as profile plots.
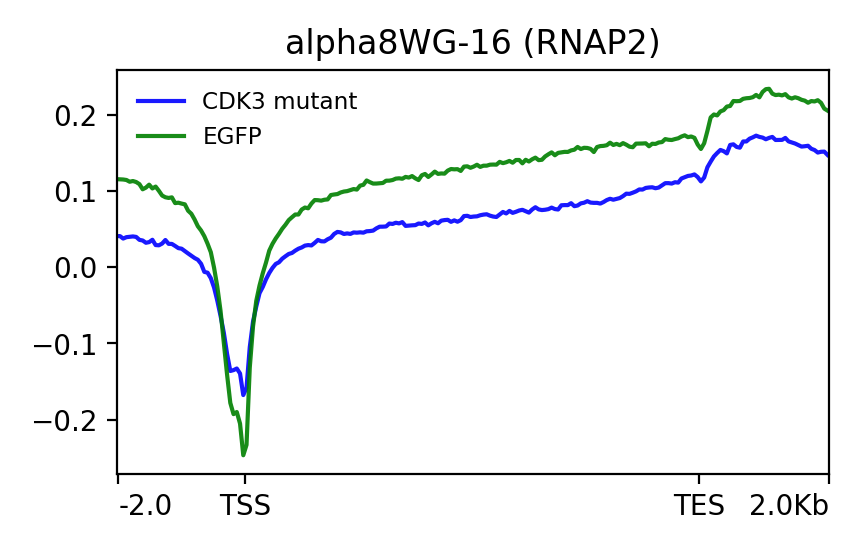
When I do this I get the following plot which doesn't really make sense to me. So the values at the TSS make sense - bigger difference for the green line, therefore smaller ratio and more negative value (dips lower than the green).
But what about across the gene body? The difference is still larger with the green line but yet the blue line dips lower on the plot. Am I misunderstanding how the command works?
Code and version info included below. Any help would be great, thanks!
Meeta
Version Info:
deeptools 3.1.3
Python 2.7.15 :: Anaconda, Inc.
# Correct order but removing scaling
bamCompare -b1 bams/MI-4_hg19/MI-4_hg19.sorted.bam -b2 bams/MI-3_hg19/MI-3_hg19.sorted.bam -p 6 --scaleFactorsMethod None -bs 20 -o MI-4_log2ratio_MI-3_noscaling.bw
bamCompare -b1 bams/MI-8_hg19/MI-8_hg19.sorted.bam -b2 bams/MI-7_hg19/MI-7_hg19.sorted.bam -p 6 --scaleFactorsMethod None -bs 20 -o MI-8_log2ratio_MI-7_noscaling.bw
# Compute matrix
computeMatrix scale-regions -S MI-4_log2ratio_MI-3_noscaling.bw MI-8_log2ratio_MI-7_noscaling.bw -R hg19.bed -b 2000 -a 2000 --skipZeros -p 6 -o deeptools/log2ratio_hg19_noscaling_mat.gz --smartLabels --regionBodyLength 7000 -bs 50
plotProfile -m deeptools/log2ratio_hg19_noscaling_mat.gz -out deeptools/log2ratio_hg19_noscaling_profile.png --perGroup --plotTitle "" --samplesLabel "CDK3 mutant" "EGFP" --regionsLabel "" --colors blue green