diff --git a/README.md b/README.md
index 9c784b5..023e737 100644
--- a/README.md
+++ b/README.md
@@ -43,7 +43,7 @@ Furthermore, this library presents other useful visualizations, such as **compar
- Classification Report
- Confusion Matrix
- ROC curve (AUROC)
- - y_prob histogram
+ - y_score histogram
- *multi-class classifier*
@@ -61,7 +61,7 @@ Furthermore, this library presents other useful visualizations, such as **compar
|  |
|  |
|  |
|:--------------------------------------------------:|:----------------------------------------------------------:|:-------------------------------------------------:|
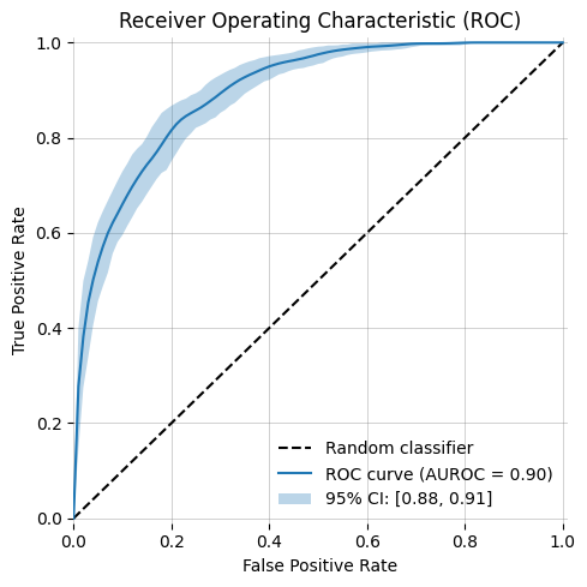
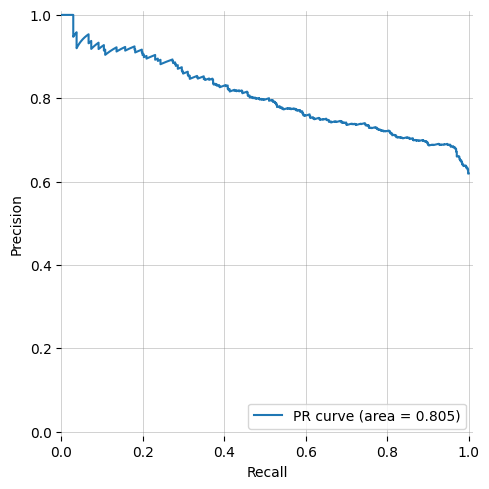
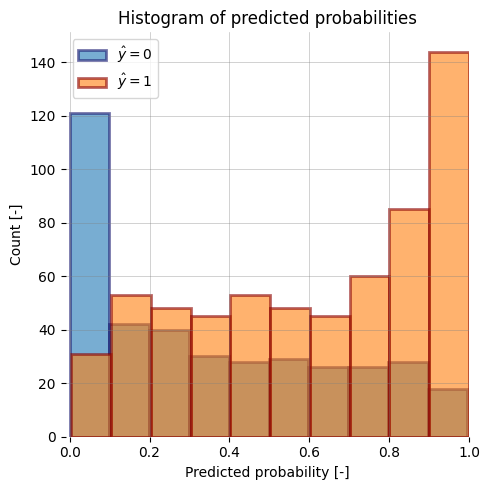
-| ROC Curve (AUROC) with bootstrapping | Precision-Recall Curve | y_prob histogram |
+| ROC Curve (AUROC) with bootstrapping | Precision-Recall Curve | y_score histogram |
|
|
|:--------------------------------------------------:|:----------------------------------------------------------:|:-------------------------------------------------:|
-| ROC Curve (AUROC) with bootstrapping | Precision-Recall Curve | y_prob histogram |
+| ROC Curve (AUROC) with bootstrapping | Precision-Recall Curve | y_score histogram |
| 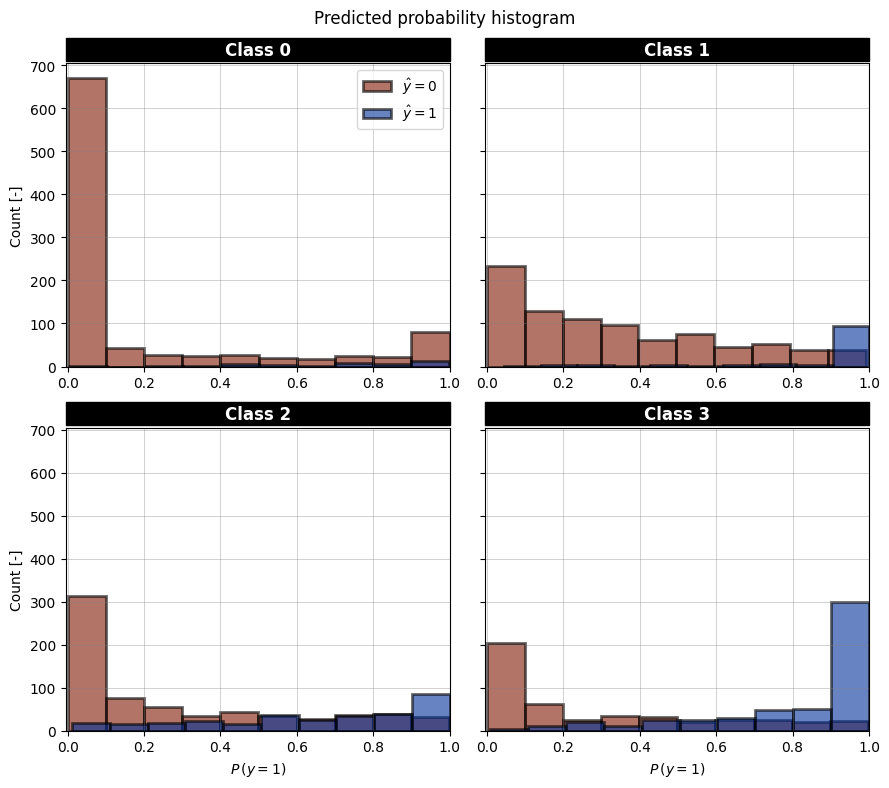 |
|  |
|  |
@@ -95,34 +95,28 @@ pip install -e .
# Usage
-Example usage of results from a binary classifier for a calibration curve.
+Get all classification metrics with **ONE** line of code. Here, for a binary classifier:
```python
-import matplotlib.pyplot as plt
-import numpy as np
import plotsandgraphs as pandg
+# ...
+pandg.pipeline.binary_classifier(y_true, y_score)
+```
-# create some predictions of a hypothetical binary classifier
-n_samples = 1000
-y_true = np.random.choice([0,1], n_samples, p=[0.4, 0.6]) # the true class labels 0 or 1, with class imbalance 40:60
-
-y_prob = np.zeros(y_true.shape) # a model's probability of class 1 predictions
-y_prob[y_true==1] = np.random.beta(1, 0.6, y_prob[y_true==1].shape)
-y_prob[y_true==0] = np.random.beta(0.5, 1, y_prob[y_true==0].shape)
-
-# show prob distribution
-fig_hist = pandg.binary_classifier.plot_y_prob_histogram(y_prob, y_true, save_fig_path=None)
-
-# create calibration curve
-fig_auroc = pandg.binary_classifier.plot_calibration_curve(y_prob, y_true, save_fig_path=None)
+Or with some more configs:
+```Python
+configs = {
+ 'roc': {'n_bootstraps': 10000},
+ 'pr': {'figsize': (8,10)}
+}
+pandg.pipeline.binary_classifier(y_true, y_score, save_fig_path='results/metrics', file_type='png', plot_kwargs=configs)
+```
+For multiclass classification:
-# --- OPTIONAL: Customize figure ---
-# get axis of figure and change title
-axes = fig_auroc.get_axes()
-ax0 = axes[0]
-ax0.set_title('New Title for Calibration Plot')
-fig_auroc.show()
+```Python
+# with multiclass data y_true (one-hot encoded) and y_score
+pandg.pipeline.multiclass_classifier(y_true, y_score)
```
# Requirements
diff --git a/plotsandgraphs/binary_classifier.py b/plotsandgraphs/binary_classifier.py
index e346749..4beda2e 100644
--- a/plotsandgraphs/binary_classifier.py
+++ b/plotsandgraphs/binary_classifier.py
@@ -45,7 +45,9 @@ def plot_accuracy(y_true, y_pred, name="", save_fig_path=None) -> Figure:
return fig
-def plot_confusion_matrix(y_true: np.ndarray, y_pred: np.ndarray, save_fig_path=None) -> Figure:
+def plot_confusion_matrix(
+ y_true: np.ndarray, y_pred: np.ndarray, save_fig_path=None
+) -> Figure:
import matplotlib.colors as colors
# Compute the confusion matrix
@@ -54,7 +56,9 @@ def plot_confusion_matrix(y_true: np.ndarray, y_pred: np.ndarray, save_fig_path=
cm = cm.astype("float") / cm.sum(axis=1)[:, np.newaxis]
# Create the ConfusionMatrixDisplay instance and plot it
- cmd = ConfusionMatrixDisplay(cm, display_labels=["class 0\nnegative", "class 1\npositive"])
+ cmd = ConfusionMatrixDisplay(
+ cm, display_labels=["class 0\nnegative", "class 1\npositive"]
+ )
fig, ax = plt.subplots(figsize=(4, 4))
cmd.plot(
cmap="YlOrRd",
@@ -144,8 +148,10 @@ def plot_classification_report(
ax : Matplotlib.pyplot.Axe
Axe object from matplotlib
"""
- print("Warning: plot_classification_report is not experiencing a bug and is, hence, currently skipped.")
- return
+ print(
+ "Warning: plot_classification_report is not experiencing a bug and is, hence, currently skipped."
+ )
+ return
import matplotlib as mpl
import matplotlib.colors as colors
@@ -156,7 +162,11 @@ def plot_classification_report(
cmap = "YlOrRd"
clf_report = classification_report(y_true, y_pred, output_dict=True, **kwargs)
- keys_to_plot = [key for key in clf_report.keys() if key not in ("accuracy", "macro avg", "weighted avg")]
+ keys_to_plot = [
+ key
+ for key in clf_report.keys()
+ if key not in ("accuracy", "macro avg", "weighted avg")
+ ]
df = pd.DataFrame(clf_report, columns=keys_to_plot).T
# the following line ensures that dataframe are sorted from the majority classes to the minority classes
df.sort_values(by=["support"], inplace=True)
@@ -325,7 +335,9 @@ def plot_roc_curve(
auc_upper = np.quantile(bootstrap_aucs, CI_upper)
auc_lower = np.quantile(bootstrap_aucs, CI_lower)
label = f"{confidence_interval:.0%} CI: [{auc_lower:.2f}, {auc_upper:.2f}]"
- plt.fill_between(base_fpr, tprs_lower, tprs_upper, alpha=0.3, label=label, zorder=2)
+ plt.fill_between(
+ base_fpr, tprs_lower, tprs_upper, alpha=0.3, label=label, zorder=2
+ )
if highlight_roc_area is True:
print(
@@ -357,7 +369,9 @@ def plot_roc_curve(
return fig
-def plot_calibration_curve(y_true: np.ndarray, y_score: np.ndarray, n_bins=10, save_fig_path=None) -> Figure:
+def plot_calibration_curve(
+ y_true: np.ndarray, y_score: np.ndarray, n_bins=10, save_fig_path=None
+) -> Figure:
"""
Creates calibration plot for a binary classifier and calculates the ECE.
@@ -379,7 +393,9 @@ def plot_calibration_curve(y_true: np.ndarray, y_score: np.ndarray, n_bins=10, s
ece : float
The expected calibration error.
"""
- prob_true, prob_pred = calibration_curve(y_true, y_score, n_bins=n_bins, strategy="uniform")
+ prob_true, prob_pred = calibration_curve(
+ y_true, y_score, n_bins=n_bins, strategy="uniform"
+ )
# Find the number of samples in each bin
bin_counts = np.histogram(y_score, bins=n_bins, range=(0, 1))[0]
@@ -452,7 +468,9 @@ def plot_calibration_curve(y_true: np.ndarray, y_score: np.ndarray, n_bins=10, s
return fig
-def plot_y_score_histogram(y_true: Optional[np.ndarray], y_score: np.ndarray = None, save_fig_path=None) -> Figure:
+def plot_y_score_histogram(
+ y_true: Optional[np.ndarray], y_score: np.ndarray = None, save_fig_path=None
+) -> Figure:
"""
Provides a histogram for the predicted probabilities of a binary classifier. If ```y_true``` is provided, it divides the ```y_score``` values into the two classes and plots them jointly into the same plot with different colors.
@@ -474,7 +492,9 @@ def plot_y_score_histogram(y_true: Optional[np.ndarray], y_score: np.ndarray = N
ax = fig.add_subplot(111)
if y_true is None:
- ax.hist(y_score, bins=10, alpha=0.9, edgecolor="midnightblue", linewidth=2, rwidth=1)
+ ax.hist(
+ y_score, bins=10, alpha=0.9, edgecolor="midnightblue", linewidth=2, rwidth=1
+ )
# same histogram as above, but with border lines
# ax.hist(y_prob, bins=10, alpha=0.5, edgecolor='black', linewidth=1.2)
else:
diff --git a/plotsandgraphs/multiclass_classifier.py b/plotsandgraphs/multiclass_classifier.py
index af1c019..c105373 100644
--- a/plotsandgraphs/multiclass_classifier.py
+++ b/plotsandgraphs/multiclass_classifier.py
@@ -20,7 +20,12 @@
from sklearn.utils import resample
from tqdm import tqdm
-from plotsandgraphs.utils import bootstrap, set_black_title_boxes, scale_ax_bbox, get_cmap
+from plotsandgraphs.utils import (
+ bootstrap,
+ set_black_title_boxes,
+ scale_ax_bbox,
+ get_cmap,
+)
def plot_roc_curve(
@@ -32,7 +37,7 @@ def plot_roc_curve(
figsize: Optional[Tuple[float, float]] = None,
class_labels: Optional[List[str]] = None,
split_plots: bool = False,
- save_fig_path:Optional[Union[str, Tuple[str, str]]] = None,
+ save_fig_path: Optional[Union[str, Tuple[str, str]]] = None,
) -> Tuple[Figure, Union[Figure, None]]:
"""
Creates two plots.
@@ -190,9 +195,6 @@ def roc_metric_function(y_true, y_score):
# create the subplot tiles (and black boxes)
set_black_title_boxes(axes.flat[:num_classes], class_labels)
-
-
-
# ---------- AUROC overview plot comparing classes ----------
# Make an AUROC overview plot comparing the aurocs per class and combined
@@ -268,10 +270,12 @@ def auroc_metric_function(y_true, y_score, average, multi_class):
def plot_y_score_histogram(
- y_true: np.ndarray, y_score: Optional[np.ndarray] = None, save_fig_path: Optional[str]=None
+ y_true: np.ndarray,
+ y_score: Optional[np.ndarray] = None,
+ save_fig_path: Optional[str] = None,
) -> Figure:
"""
- Histogram plot that is intended to show the distribution of the predicted probabilities for different classes, where the the different classes (y_true==0 and y_true==1) are plotted in different colors.
+ Histogram plot that is intended to show the distribution of the predicted probabilities for different classes, where the the different classes (y_true==0 and y_true==1) are plotted in different colors.
Limitations: Does not work for samples, that can be part of multiple classes (e.g. multilabel classification).
Parameters
@@ -288,15 +292,19 @@ def plot_y_score_histogram(
Figure
The figure of the histogram plot.
"""
-
+
num_classes = y_true.shape[-1]
class_labels = [f"Class {i}" for i in range(num_classes)]
-
+
cmap, colors = get_cmap("roma", n_colors=2) # 2 colors for y==0 and y==1 per class
-
+
# Aiming for a square plot
- plot_cols = np.ceil(np.sqrt(num_classes)).astype(int) # Number of plots in a row # noqa
- plot_rows = np.ceil(num_classes / plot_cols).astype(int) # Number of plots in a column # noqa
+ plot_cols = np.ceil(np.sqrt(num_classes)).astype(
+ int
+ ) # Number of plots in a row # noqa
+ plot_rows = np.ceil(num_classes / plot_cols).astype(
+ int
+ ) # Number of plots in a column # noqa
fig, axes = plt.subplots(
nrows=plot_rows,
ncols=plot_cols,
diff --git a/plotsandgraphs/pipeline.py b/plotsandgraphs/pipeline.py
index 00eb099..24c8844 100644
--- a/plotsandgraphs/pipeline.py
+++ b/plotsandgraphs/pipeline.py
@@ -6,109 +6,105 @@
from . import multiclass_classifier as mc
+FILE_ENDINGS = Literal["pdf", "png", "jpg", "jpeg", "svg"]
-FILE_ENDINGS = Literal['pdf', 'png', 'jpg', 'jpeg', 'svg']
-
-
-def binary_classifier(y_true, y_score, save_fig_path=None, plot_kwargs={}, file_type:FILE_ENDINGS='png'):
-
-
+def binary_classifier(
+ y_true, y_score, save_fig_path=None, plot_kwargs={}, file_type: FILE_ENDINGS = "png"
+):
# Create new tqdm instance
- tqdm_instance = tqdm(total=6, desc='Binary classifier metrics', leave=True)
-
+ tqdm_instance = tqdm(total=6, desc="Binary classifier metrics", leave=True)
+
# Update tqdm instance
tqdm_instance.update()
-
+
# 1) Plot ROC curve
- roc_kwargs = plot_kwargs.get('roc', {})
- save_path = get_file_path(save_fig_path, 'roc_curve', file_type)
+ roc_kwargs = plot_kwargs.get("roc", {})
+ save_path = get_file_path(save_fig_path, "roc_curve", file_type)
bc.plot_roc_curve(y_true, y_score, save_fig_path=save_path, **roc_kwargs)
tqdm_instance.update()
-
+
# 2) Plot precision-recall curve
- pr_kwargs = plot_kwargs.get('pr', {})
- save_path = get_file_path(save_fig_path, 'pr_curve', file_type)
+ pr_kwargs = plot_kwargs.get("pr", {})
+ save_path = get_file_path(save_fig_path, "pr_curve", file_type)
bc.plot_pr_curve(y_true, y_score, save_fig_path=save_path, **pr_kwargs)
tqdm_instance.update()
-
+
# 3) Plot calibration curve
- cal_kwargs = plot_kwargs.get('cal', {})
- save_path = get_file_path(save_fig_path, 'calibration_curve', file_type)
+ cal_kwargs = plot_kwargs.get("cal", {})
+ save_path = get_file_path(save_fig_path, "calibration_curve", file_type)
bc.plot_calibration_curve(y_true, y_score, save_fig_path=save_path, **cal_kwargs)
tqdm_instance.update()
-
+
# 3) Plot confusion matrix
- cm_kwargs = plot_kwargs.get('cm', {})
- save_path = get_file_path(save_fig_path, 'confusion_matrix', file_type)
+ cm_kwargs = plot_kwargs.get("cm", {})
+ save_path = get_file_path(save_fig_path, "confusion_matrix", file_type)
bc.plot_confusion_matrix(y_true, y_score, save_fig_path=save_path, **cm_kwargs)
tqdm_instance.update()
-
+
# 5) Plot classification report
- cr_kwargs = plot_kwargs.get('cr', {})
- save_path = get_file_path(save_fig_path, 'classification_report', file_type)
+ cr_kwargs = plot_kwargs.get("cr", {})
+ save_path = get_file_path(save_fig_path, "classification_report", file_type)
bc.plot_classification_report(y_true, y_score, save_fig_path=save_path, **cr_kwargs)
tqdm_instance.update()
-
+
# 6) Plot y_score histogram
- hist_kwargs = plot_kwargs.get('hist', {})
- save_path = get_file_path(save_fig_path, 'y_score_histogram', file_type)
+ hist_kwargs = plot_kwargs.get("hist", {})
+ save_path = get_file_path(save_fig_path, "y_score_histogram", file_type)
bc.plot_y_score_histogram(y_true, y_score, save_fig_path=save_path, **hist_kwargs)
tqdm_instance.update()
-
- return
-
+ return
-def multiclass_classifier(y_true, y_score, save_fig_path=None, plot_kwargs={}, file_type:FILE_ENDINGS = 'png'):
-
+def multiclass_classifier(
+ y_true, y_score, save_fig_path=None, plot_kwargs={}, file_type: FILE_ENDINGS = "png"
+):
# Create new tqdm instance
- tqdm_instance = tqdm(total=6, desc='Binary classifier metrics', leave=True)
-
+ tqdm_instance = tqdm(total=6, desc="Binary classifier metrics", leave=True)
+
# Update tqdm instance
tqdm_instance.update()
-
+
# 1) Plot ROC curve
- roc_kwargs = plot_kwargs.get('roc', {})
- save_path = get_file_path(save_fig_path, 'roc_curve', '')
+ roc_kwargs = plot_kwargs.get("roc", {})
+ save_path = get_file_path(save_fig_path, "roc_curve", "")
mc.plot_roc_curve(y_true, y_score, save_fig_path=save_path, **roc_kwargs)
tqdm_instance.update()
-
+
# 2) Plot precision-recall curve
# pr_kwargs = plot_kwargs.get('pr', {})
# mc.plot_pr_curve(y_true, y_score, save_fig_path=save_fig_path, **pr_kwargs)
# tqdm_instance.update()
-
+
# 3) Plot calibration curve
# cal_kwargs = plot_kwargs.get('cal', {})
# mc.plot_calibration_curve(y_true, y_score, save_fig_path=save_fig_path, **cal_kwargs)
# tqdm_instance.update()
-
+
# 3) Plot confusion matrix
# cm_kwargs = plot_kwargs.get('cm', {})
# mc.plot_confusion_matrix(y_true, y_score, save_fig_path=save_fig_path, **cm_kwargs)
# tqdm_instance.update()
-
+
# 5) Plot classification report
# cr_kwargs = plot_kwargs.get('cr', {})
# mc.plot_classification_report(y_true, y_score, save_fig_path=save_fig_path, **cr_kwargs)
# tqdm_instance.update()
-
+
# 6) Plot y_score histogram
- hist_kwargs = plot_kwargs.get('hist', {})
- save_path = get_file_path(save_fig_path, 'y_score_histogram', file_type)
+ hist_kwargs = plot_kwargs.get("hist", {})
+ save_path = get_file_path(save_fig_path, "y_score_histogram", file_type)
mc.plot_y_score_histogram(y_true, y_score, save_fig_path=save_path, **hist_kwargs)
tqdm_instance.update()
-
- return
+ return
-def get_file_path(save_fig_path: Union[Path,None, str], name:str, ending:str):
+def get_file_path(save_fig_path: Union[Path, None, str], name: str, ending: str):
if save_fig_path is None:
return None
else:
result = Path(save_fig_path) / f"{name}.{ending}"
print(result)
- return str(result)
\ No newline at end of file
+ return str(result)
diff --git a/plotsandgraphs/utils.py b/plotsandgraphs/utils.py
index 65b801d..b8c2a32 100644
--- a/plotsandgraphs/utils.py
+++ b/plotsandgraphs/utils.py
@@ -1,4 +1,4 @@
-from typing import Optional, List, Callable, Dict, Tuple, Union, TYPE_CHECKING, Literal
+from typing import Optional, List, Callable, Dict, Tuple, TYPE_CHECKING
from tqdm import tqdm
from sklearn.utils import resample
import numpy as np
@@ -11,7 +11,12 @@
from matplotlib.axes import Axes
-def bootstrap(metric_function: Callable, input_resample: List[np.ndarray], n_bootstraps: int, metric_kwargs: Dict={}) -> List:
+def bootstrap(
+ metric_function: Callable,
+ input_resample: List[np.ndarray],
+ n_bootstraps: int,
+ metric_kwargs: Dict = {},
+) -> List:
"""
A bootstrapping function for a metric function. The metric function should take the same number of arguments as the length of input_resample.
@@ -33,29 +38,28 @@ def bootstrap(metric_function: Callable, input_resample: List[np.ndarray], n_boo
"""
results = []
# for each bootstrap iteration
- for _ in tqdm(range(n_bootstraps), desc='Bootsrapping', leave=True):
+ for _ in tqdm(range(n_bootstraps), desc="Bootsrapping", leave=True):
# resample indices with replacement
indices = resample(np.arange(len(input_resample[0])), replace=True)
input_resampled = [x[indices] for x in input_resample]
# calculate metric
result = metric_function(*input_resampled, **metric_kwargs)
-
+
results.append(result)
-
- return results
+ return results
class ExtendedTextBox_v2:
"""
Black background boxes for titles in maptlolib subplots
-
+
From:
https://stackoverflow.com/questions/40796117/how-do-i-make-the-width-of-the-title-box-span-the-entire-plot
https://matplotlib.org/stable/gallery/userdemo/custom_boxstyle01.html?highlight=boxstyle+_style_list
"""
- def __init__(self, pad=0.3, width=500.):
+ def __init__(self, pad=0.3, width=500.0):
"""
The arguments must be floats and have default values.
@@ -85,22 +89,25 @@ def __call__(self, x0, y0, width, height, mutation_size):
# padding
pad = mutation_size * self.pad
# width and height with padding added
- #width = width + 2.*pad
- height = height + 3 * pad
+ # width = width + 2.*pad
+ height = height + 3 * pad
# boundary of the padded box
y0 = y0 - pad # change this to move the text
- y1 = y0 + height
+ y1 = y0 + height
_x0 = x0
- x0 = _x0 +width /2. - self.width/2.
- x1 = _x0 +width /2. + self.width/2.
+ x0 = _x0 + width / 2.0 - self.width / 2.0
+ x1 = _x0 + width / 2.0 + self.width / 2.0
# return the new path
- return Path([(x0, y0),
- (x1, y0), (x1, y1), (x0, y1),
- (x0, y0)],
- closed=True)
+ return Path([(x0, y0), (x1, y0), (x1, y1), (x0, y1), (x0, y0)], closed=True)
-def _set_black_title_box(ax: "Axes", title:str, backgroundcolor='black', color='white', title_kwargs: Optional[Dict]=None):
+def _set_black_title_box(
+ ax: "Axes",
+ title: str,
+ backgroundcolor="black",
+ color="white",
+ title_kwargs: Optional[Dict] = None,
+):
"""
Note: Do not use this function by itself, instead use `set_black_title_boxes()`.
Sets the title of the given axes with a black bounding box.
@@ -114,16 +121,25 @@ def _set_black_title_box(ax: "Axes", title:str, backgroundcolor='black', color='
- set_title_kwargs: Keyword arguments to pass to `ax.set_title()`.
"""
if title_kwargs is None:
- title_kwargs = {'fontdict': {"fontname": "Arial Black", "fontweight": "bold"}}
- BoxStyle._style_list["ext"] = ExtendedTextBox_v2
+ title_kwargs = {"fontdict": {"fontname": "Arial Black", "fontweight": "bold"}}
+ BoxStyle._style_list["ext"] = ExtendedTextBox_v2
ax_width = ax.get_window_extent().width
# make title with black bounding box
- title_instance = ax.set_title(title, backgroundcolor=backgroundcolor, color=color, **title_kwargs)
- bb = title_instance.get_bbox_patch() # get bbox from title
- bb.set_boxstyle("ext", pad=0.1, width=ax_width) # use custom style
-
-
-def set_black_title_boxes(axes: "np.ndarray[Axes]", titles: List[str], backgroundcolor='black', color='white', title_kwargs: Optional[Dict]=None, tight_layout_kwargs: Dict={}):
+ title_instance = ax.set_title(
+ title, backgroundcolor=backgroundcolor, color=color, **title_kwargs
+ )
+ bb = title_instance.get_bbox_patch() # get bbox from title
+ bb.set_boxstyle("ext", pad=0.1, width=ax_width) # use custom style
+
+
+def set_black_title_boxes(
+ axes: "np.ndarray[Axes]",
+ titles: List[str],
+ backgroundcolor="black",
+ color="white",
+ title_kwargs: Optional[Dict] = None,
+ tight_layout_kwargs: Dict = {},
+):
"""
Creates black boxes for the subtitles above the given axes with the given titles. The subtitles are centered above the axes.
@@ -145,18 +161,15 @@ def set_black_title_boxes(axes: "np.ndarray[Axes]", titles: List[str], backgroun
for i, ax in enumerate(axes.flat):
_set_black_title_box(ax, titles[i], backgroundcolor, color, title_kwargs)
-
+
plt.tight_layout(**tight_layout_kwargs)
-
+
for i, ax in enumerate(axes.flat):
_set_black_title_box(ax, titles[i], backgroundcolor, color, title_kwargs)
-
-
+
return
-
-
-
-
+
+
def scale_ax_bbox(ax: "Axes", factor: float):
# Get the current position of the subplot
box = ax.get_position()
@@ -167,12 +180,13 @@ def scale_ax_bbox(ax: "Axes", factor: float):
# Set the new position
ax.set_position([box.x0 + adjustment, box.y0, new_width, box.height])
-
- return
+ return
-def get_cmap(cmap_name: str, n_colors: Optional[int]=None) -> Tuple[LinearSegmentedColormap, Tuple]:
+def get_cmap(
+ cmap_name: str, n_colors: Optional[int] = None
+) -> Tuple[LinearSegmentedColormap, Tuple]:
"""
Loads one of the custom cmaps from the cmaps folder.
@@ -185,23 +199,19 @@ def get_cmap(cmap_name: str, n_colors: Optional[int]=None) -> Tuple[LinearSegmen
-------
Tuple[LinearSegmentedColormap, Union[None, Tuple]]
A tuple of the cmap and a list of colors if n_colors is not None.
-
+
Example
-------
>>> cmap_name = 'hawaii'
>>> cmap, color_list = get_cmap(cmap_name, n_colors=10)
"""
from pathlib import Path as PathClass
-
- cm_path = PathClass(__file__).parent / ('cmaps/' + cmap_name + '.txt')
+
+ cm_path = PathClass(__file__).parent / ("cmaps/" + cmap_name + ".txt")
cm_data = np.loadtxt(cm_path)
- cmap_name = cmap_name.split('.')[0]
+ cmap_name = cmap_name.split(".")[0]
cmap = LinearSegmentedColormap.from_list(cmap_name, cm_data)
if n_colors is None:
n_colors = 10
color_list = cmap(np.linspace(0, 1, n_colors))
return cmap, color_list
-
-
-
-
\ No newline at end of file
diff --git a/tests/test_binary_classifier.py b/tests/test_binary_classifier.py
index a8e72b4..9426287 100644
--- a/tests/test_binary_classifier.py
+++ b/tests/test_binary_classifier.py
@@ -41,8 +41,12 @@ def test_hist_plot(random_data_binary_classifier):
"""
y_true, y_score = random_data_binary_classifier
print(TEST_RESULTS_PATH)
- binary.plot_y_score_histogram(y_true=None, y_score=y_score, save_fig_path=TEST_RESULTS_PATH / "histogram.png")
- binary.plot_y_score_histogram(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "histogram_2_classes.png")
+ binary.plot_y_score_histogram(
+ y_true=None, y_score=y_score, save_fig_path=TEST_RESULTS_PATH / "histogram.png"
+ )
+ binary.plot_y_score_histogram(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "histogram_2_classes.png"
+ )
# test roc curve without bootstrapping
@@ -56,7 +60,9 @@ def test_roc_curve(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_roc_curve(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "roc_curve.png")
+ binary.plot_roc_curve(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "roc_curve.png"
+ )
# test roc curve with bootstrapping
@@ -71,7 +77,10 @@ def test_roc_curve_bootstrap(random_data_binary_classifier):
"""
y_true, y_score = random_data_binary_classifier
binary.plot_roc_curve(
- y_true, y_score, n_bootstraps=10000, save_fig_path=TEST_RESULTS_PATH / "roc_curve_bootstrap.png"
+ y_true,
+ y_score,
+ n_bootstraps=10000,
+ save_fig_path=TEST_RESULTS_PATH / "roc_curve_bootstrap.png",
)
@@ -86,7 +95,9 @@ def test_pr_curve(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_pr_curve(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "pr_curve.png")
+ binary.plot_pr_curve(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "pr_curve.png"
+ )
# test confusion matrix
@@ -100,7 +111,9 @@ def test_confusion_matrix(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_confusion_matrix(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "confusion_matrix.png")
+ binary.plot_confusion_matrix(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "confusion_matrix.png"
+ )
# test classification report
@@ -114,7 +127,10 @@ def test_classification_report(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_classification_report(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "classification_report.png")
+ binary.plot_classification_report(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "classification_report.png"
+ )
+
# test calibration curve
def test_calibration_curve(random_data_binary_classifier):
@@ -127,7 +143,10 @@ def test_calibration_curve(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_calibration_curve(y_score, y_true, save_fig_path=TEST_RESULTS_PATH / "calibration_curve.png")
+ binary.plot_calibration_curve(
+ y_score, y_true, save_fig_path=TEST_RESULTS_PATH / "calibration_curve.png"
+ )
+
# test accuracy
def test_accuracy(random_data_binary_classifier):
@@ -140,4 +159,6 @@ def test_accuracy(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_accuracy(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "accuracy.png")
+ binary.plot_accuracy(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "accuracy.png"
+ )
diff --git a/tests/test_multiclass_classifier.py b/tests/test_multiclass_classifier.py
index d47e7e0..3cbdb70 100644
--- a/tests/test_multiclass_classifier.py
+++ b/tests/test_multiclass_classifier.py
@@ -3,6 +3,7 @@
from itertools import product
import numpy as np
import pytest
+
# import plotsandgraphs.binary_classifier as binary
import plotsandgraphs.multiclass_classifier as multiclass
@@ -10,7 +11,9 @@
# @pytest.fixture(scope="module")
-def random_data_multiclass_classifier(num_classes:int = 3) -> Tuple[np.ndarray, np.ndarray]:
+def random_data_multiclass_classifier(
+ num_classes: int = 3,
+) -> Tuple[np.ndarray, np.ndarray]:
"""
Create random data for binary classifier tests.
@@ -21,26 +24,26 @@ def random_data_multiclass_classifier(num_classes:int = 3) -> Tuple[np.ndarray,
"""
class_labels = np.arange(num_classes)
class_probs = np.random.random(num_classes)
- class_probs = class_probs / class_probs.sum() # normalize
+ class_probs = class_probs / class_probs.sum() # normalize
# True labels
y_true = np.random.choice(class_labels, p=class_probs, size=1000)
# one hot encoding
- y_true_one_hot = np.eye(num_classes)[y_true]
+ y_true_one_hot = np.eye(num_classes)[y_true]
# Predicted labels
y_pred = np.ones(y_true_one_hot.shape)
# parameters for Beta distribution for each label (a0,b0 for class 0, a1,b1 for class 1)
- a0, b0 = [0.1, 0.6, 0.3, 0.4, 2]*10, [0.4, 1.2, 0.8, 1, 5]*10
- a1, b1 = [0.9, 0.8, 0.9, 1.2, 5]*10, [0.4, 0.1, 0.5, 0.3, 2]*10
+ a0, b0 = [0.1, 0.6, 0.3, 0.4, 2] * 10, [0.4, 1.2, 0.8, 1, 5] * 10
+ a1, b1 = [0.9, 0.8, 0.9, 1.2, 5] * 10, [0.4, 0.1, 0.5, 0.3, 2] * 10
# iterate through all the columns/labels and create a beta distribution for each label
for i in range(y_pred.shape[1]):
y = y_pred[:, i]
y_t = y_true_one_hot[:, i]
- y[y_t==0] = np.random.beta(a0[i], b0[i], size=y[y_t==0].shape)
- y[y_t==1] = np.random.beta(a1[i], b1[i], size=y[y_t==1].shape)
-
+ y[y_t == 0] = np.random.beta(a0[i], b0[i], size=y[y_t == 0].shape)
+ y[y_t == 1] = np.random.beta(a1[i], b1[i], size=y[y_t == 1].shape)
+
return y_true_one_hot, y_pred
@@ -57,42 +60,71 @@ def test_hist_plot():
for num_classes in [1, 2, 3, 4, 5, 10, 16, 25]:
y_true, y_prob = random_data_multiclass_classifier(num_classes=num_classes)
print(TEST_RESULTS_PATH)
- multiclass.plot_y_prob_histogram(y_true=y_true, y_score=y_prob, save_fig_path=TEST_RESULTS_PATH / f"histogram_{num_classes}_classes.png")
+ multiclass.plot_y_prob_histogram(
+ y_true=y_true,
+ y_score=y_prob,
+ save_fig_path=TEST_RESULTS_PATH / f"histogram_{num_classes}_classes.png",
+ )
# multiclass.plot_y_prob_histogram(y_prob=y_prob, save_fig_path=TEST_RESULTS_PATH / "histogram_classes.png")
-
-
+
+
def test_roc_curve():
"""
Test roc curve.
-
+
Parameters
----------
random_data_binary_classifier : Tuple[np.ndarray, np.ndarray]
The simulated data.
"""
+
# helper function for file name, to avoid repeating code
- def get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots):
+ def get_path_name(
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots
+ ):
if split_plots is False:
- fig_path = TEST_RESULTS_PATH / f"roc_curves_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ fig_path = (
+ TEST_RESULTS_PATH
+ / f"roc_curves_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ )
else:
- fig_path_1 = TEST_RESULTS_PATH / f"roc_curves_split_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
- fig_path_2 = TEST_RESULTS_PATH / f"auroc_comparison_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ fig_path_1 = (
+ TEST_RESULTS_PATH
+ / f"roc_curves_split_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ )
+ fig_path_2 = (
+ TEST_RESULTS_PATH
+ / f"auroc_comparison_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ )
fig_path = [fig_path_1, fig_path_2]
return fig_path
-
-
+
y_true, y_prob = random_data_multiclass_classifier(num_classes=3)
-
+
confidence_intervals = [None, 0.99]
highlight_roc_area = [True, False]
n_bootstraps = [1, 1000]
figsizes = [None]
split_plots = [True, False]
-
- # From the previous lists I want all possible combinations
- combinations = list(product(confidence_intervals, highlight_roc_area, n_bootstraps, figsizes, split_plots))
-
- for confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots in combinations:
+
+ # From the previous lists I want all possible combinations
+ combinations = list(
+ product(
+ confidence_intervals,
+ highlight_roc_area,
+ n_bootstraps,
+ figsizes,
+ split_plots,
+ )
+ )
+
+ for (
+ confidence_interval,
+ highlight_roc_area,
+ n_bootstraps,
+ figsize,
+ split_plots,
+ ) in combinations:
# check if one or two figures should be saved (splot_plots=True or False)
# if split_plots is False:
# fig_path = TEST_RESULTS_PATH / f"roc_curves_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
@@ -100,51 +132,77 @@ def get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize
# fig_path_1 = TEST_RESULTS_PATH / f"roc_curves_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
# fig_path_2 = TEST_RESULTS_PATH / f"auroc_comparison_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
# fig_path = [fig_path_1, fig_path_2]
-
- fig_path = get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots)
-
+
+ fig_path = get_path_name(
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots
+ )
+
# It should raise an error when confidence_interval is None but highlight_roc_area is True
if confidence_interval is None and highlight_roc_area is True:
with pytest.raises(ValueError):
- multiclass.plot_roc_curve(y_true=y_true, y_score=y_prob,
- confidence_interval=confidence_interval,
- highlight_roc_area=highlight_roc_area,
- n_bootstraps=n_bootstraps,
- figsize=figsize,
- split_plots=split_plots,
- save_fig_path=fig_path)
+ multiclass.plot_roc_curve(
+ y_true=y_true,
+ y_score=y_prob,
+ confidence_interval=confidence_interval,
+ highlight_roc_area=highlight_roc_area,
+ n_bootstraps=n_bootstraps,
+ figsize=figsize,
+ split_plots=split_plots,
+ save_fig_path=fig_path,
+ )
# Otherwise no error
else:
- multiclass.plot_roc_curve(y_true=y_true, y_score=y_prob,
- confidence_interval=confidence_interval,
- highlight_roc_area=highlight_roc_area,
- n_bootstraps=n_bootstraps,
- figsize=figsize,
- split_plots=split_plots,
- save_fig_path=fig_path)
-
- # check for SMALL figure size
- confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots = 0.95, True, 100, (3,3), False
- fig_path = get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots)
- multiclass.plot_roc_curve(y_true=y_true, y_score=y_prob,
- confidence_interval=confidence_interval,
- highlight_roc_area=highlight_roc_area,
- n_bootstraps=n_bootstraps,
- figsize=figsize,
- split_plots=split_plots,
- save_fig_path=fig_path)
-
- # check for BIG figure size
- confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots = 0.95, True, 100, (15, 15), False
- fig_path = get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots)
- multiclass.plot_roc_curve(y_true=y_true, y_score=y_prob,
- confidence_interval=confidence_interval,
- highlight_roc_area=highlight_roc_area,
- n_bootstraps=n_bootstraps,
- figsize=figsize,
- split_plots=split_plots,
- save_fig_path=fig_path)
-
-
+ multiclass.plot_roc_curve(
+ y_true=y_true,
+ y_score=y_prob,
+ confidence_interval=confidence_interval,
+ highlight_roc_area=highlight_roc_area,
+ n_bootstraps=n_bootstraps,
+ figsize=figsize,
+ split_plots=split_plots,
+ save_fig_path=fig_path,
+ )
+ # check for SMALL figure size
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots = (
+ 0.95,
+ True,
+ 100,
+ (3, 3),
+ False,
+ )
+ fig_path = get_path_name(
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots
+ )
+ multiclass.plot_roc_curve(
+ y_true=y_true,
+ y_score=y_prob,
+ confidence_interval=confidence_interval,
+ highlight_roc_area=highlight_roc_area,
+ n_bootstraps=n_bootstraps,
+ figsize=figsize,
+ split_plots=split_plots,
+ save_fig_path=fig_path,
+ )
+ # check for BIG figure size
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots = (
+ 0.95,
+ True,
+ 100,
+ (15, 15),
+ False,
+ )
+ fig_path = get_path_name(
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots
+ )
+ multiclass.plot_roc_curve(
+ y_true=y_true,
+ y_score=y_prob,
+ confidence_interval=confidence_interval,
+ highlight_roc_area=highlight_roc_area,
+ n_bootstraps=n_bootstraps,
+ figsize=figsize,
+ split_plots=split_plots,
+ save_fig_path=fig_path,
+ )
diff --git a/tests/test_pipeline.py b/tests/test_pipeline.py
index 1ad1fe3..dd9cdaf 100644
--- a/tests/test_pipeline.py
+++ b/tests/test_pipeline.py
@@ -1,7 +1,4 @@
from pathlib import Path
-from typing import Tuple
-import numpy as np
-import pytest
import shutil
from plotsandgraphs import pipeline
@@ -11,6 +8,7 @@
TEST_RESULTS_PATH = Path("tests/test_results/pipeline")
+
def test_binary_classification_pipeline(random_data_binary_classifier):
"""
Test binary classification pipeline.
@@ -21,36 +19,40 @@ def test_binary_classification_pipeline(random_data_binary_classifier):
The simulated data.
"""
save_fig_path = TEST_RESULTS_PATH / "binary_classifier"
-
+
# Delete the folder and its previous contents
if save_fig_path.exists() and save_fig_path.is_dir():
shutil.rmtree(save_fig_path)
y_true, y_score = random_data_binary_classifier
- pipeline.binary_classifier(y_true, y_score, save_fig_path=save_fig_path, file_type='png')
-
+ pipeline.binary_classifier(
+ y_true, y_score, save_fig_path=save_fig_path, file_type="png"
+ )
+
# assert that there are files with the names xy in the save_fig_path
assert (save_fig_path / "roc_curve.png").exists()
assert (save_fig_path / "y_score_histogram.png").exists()
assert (save_fig_path / "calibration_curve.png").exists()
assert (save_fig_path / "confusion_matrix.png").exists()
assert (save_fig_path / "pr_curve.png").exists()
-
-
+
+
def test_multiclassification_pipeline():
"""
Test multiclassification pipeline.
"""
for num_classes in [3]:
save_fig_path = TEST_RESULTS_PATH / f"multiclass_{num_classes}_classes"
-
+
# Delete the folder and its previous contents
if save_fig_path.exists() and save_fig_path.is_dir():
shutil.rmtree(save_fig_path)
-
+
y_true, y_score = random_data_multiclass_classifier(num_classes=num_classes)
- pipeline.multiclass_classifier(y_true, y_score, save_fig_path=save_fig_path, file_type='png')
-
+ pipeline.multiclass_classifier(
+ y_true, y_score, save_fig_path=save_fig_path, file_type="png"
+ )
+
# assert that there are files with the names xy in the save_fig_path
assert (save_fig_path / "roc_curve.png").exists()
- assert (save_fig_path / "y_score_histogram.png").exists()
\ No newline at end of file
+ assert (save_fig_path / "y_score_histogram.png").exists()
diff --git a/tests/utils.py b/tests/utils.py
index ee27bdb..780b451 100644
--- a/tests/utils.py
+++ b/tests/utils.py
@@ -25,7 +25,9 @@ def random_data_binary_classifier() -> Tuple[np.ndarray, np.ndarray]:
return y_true, y_score
-def random_data_multiclass_classifier(num_classes:int = 3) -> Tuple[np.ndarray, np.ndarray]:
+def random_data_multiclass_classifier(
+ num_classes: int = 3,
+) -> Tuple[np.ndarray, np.ndarray]:
"""
Create random data for binary classifier tests.
@@ -36,24 +38,24 @@ def random_data_multiclass_classifier(num_classes:int = 3) -> Tuple[np.ndarray,
"""
class_labels = np.arange(num_classes)
class_probs = np.random.random(num_classes)
- class_probs = class_probs / class_probs.sum() # normalize
+ class_probs = class_probs / class_probs.sum() # normalize
# True labels
y_true = np.random.choice(class_labels, p=class_probs, size=1000)
# one hot encoding
- y_true_one_hot = np.eye(num_classes)[y_true]
+ y_true_one_hot = np.eye(num_classes)[y_true]
# Predicted labels
y_pred = np.ones(y_true_one_hot.shape)
# parameters for Beta distribution for each label (a0,b0 for class 0, a1,b1 for class 1)
- a0, b0 = [0.1, 0.6, 0.3, 0.4, 2]*10, [0.4, 1.2, 0.8, 1, 5]*10
- a1, b1 = [0.9, 0.8, 0.9, 1.2, 5]*10, [0.4, 0.1, 0.5, 0.3, 2]*10
+ a0, b0 = [0.1, 0.6, 0.3, 0.4, 2] * 10, [0.4, 1.2, 0.8, 1, 5] * 10
+ a1, b1 = [0.9, 0.8, 0.9, 1.2, 5] * 10, [0.4, 0.1, 0.5, 0.3, 2] * 10
# iterate through all the columns/labels and create a beta distribution for each label
for i in range(y_pred.shape[1]):
y = y_pred[:, i]
y_t = y_true_one_hot[:, i]
- y[y_t==0] = np.random.beta(a0[i], b0[i], size=y[y_t==0].shape)
- y[y_t==1] = np.random.beta(a1[i], b1[i], size=y[y_t==1].shape)
-
- return y_true_one_hot, y_pred
\ No newline at end of file
+ y[y_t == 0] = np.random.beta(a0[i], b0[i], size=y[y_t == 0].shape)
+ y[y_t == 1] = np.random.beta(a1[i], b1[i], size=y[y_t == 1].shape)
+
+ return y_true_one_hot, y_pred
|
@@ -95,34 +95,28 @@ pip install -e .
# Usage
-Example usage of results from a binary classifier for a calibration curve.
+Get all classification metrics with **ONE** line of code. Here, for a binary classifier:
```python
-import matplotlib.pyplot as plt
-import numpy as np
import plotsandgraphs as pandg
+# ...
+pandg.pipeline.binary_classifier(y_true, y_score)
+```
-# create some predictions of a hypothetical binary classifier
-n_samples = 1000
-y_true = np.random.choice([0,1], n_samples, p=[0.4, 0.6]) # the true class labels 0 or 1, with class imbalance 40:60
-
-y_prob = np.zeros(y_true.shape) # a model's probability of class 1 predictions
-y_prob[y_true==1] = np.random.beta(1, 0.6, y_prob[y_true==1].shape)
-y_prob[y_true==0] = np.random.beta(0.5, 1, y_prob[y_true==0].shape)
-
-# show prob distribution
-fig_hist = pandg.binary_classifier.plot_y_prob_histogram(y_prob, y_true, save_fig_path=None)
-
-# create calibration curve
-fig_auroc = pandg.binary_classifier.plot_calibration_curve(y_prob, y_true, save_fig_path=None)
+Or with some more configs:
+```Python
+configs = {
+ 'roc': {'n_bootstraps': 10000},
+ 'pr': {'figsize': (8,10)}
+}
+pandg.pipeline.binary_classifier(y_true, y_score, save_fig_path='results/metrics', file_type='png', plot_kwargs=configs)
+```
+For multiclass classification:
-# --- OPTIONAL: Customize figure ---
-# get axis of figure and change title
-axes = fig_auroc.get_axes()
-ax0 = axes[0]
-ax0.set_title('New Title for Calibration Plot')
-fig_auroc.show()
+```Python
+# with multiclass data y_true (one-hot encoded) and y_score
+pandg.pipeline.multiclass_classifier(y_true, y_score)
```
# Requirements
diff --git a/plotsandgraphs/binary_classifier.py b/plotsandgraphs/binary_classifier.py
index e346749..4beda2e 100644
--- a/plotsandgraphs/binary_classifier.py
+++ b/plotsandgraphs/binary_classifier.py
@@ -45,7 +45,9 @@ def plot_accuracy(y_true, y_pred, name="", save_fig_path=None) -> Figure:
return fig
-def plot_confusion_matrix(y_true: np.ndarray, y_pred: np.ndarray, save_fig_path=None) -> Figure:
+def plot_confusion_matrix(
+ y_true: np.ndarray, y_pred: np.ndarray, save_fig_path=None
+) -> Figure:
import matplotlib.colors as colors
# Compute the confusion matrix
@@ -54,7 +56,9 @@ def plot_confusion_matrix(y_true: np.ndarray, y_pred: np.ndarray, save_fig_path=
cm = cm.astype("float") / cm.sum(axis=1)[:, np.newaxis]
# Create the ConfusionMatrixDisplay instance and plot it
- cmd = ConfusionMatrixDisplay(cm, display_labels=["class 0\nnegative", "class 1\npositive"])
+ cmd = ConfusionMatrixDisplay(
+ cm, display_labels=["class 0\nnegative", "class 1\npositive"]
+ )
fig, ax = plt.subplots(figsize=(4, 4))
cmd.plot(
cmap="YlOrRd",
@@ -144,8 +148,10 @@ def plot_classification_report(
ax : Matplotlib.pyplot.Axe
Axe object from matplotlib
"""
- print("Warning: plot_classification_report is not experiencing a bug and is, hence, currently skipped.")
- return
+ print(
+ "Warning: plot_classification_report is not experiencing a bug and is, hence, currently skipped."
+ )
+ return
import matplotlib as mpl
import matplotlib.colors as colors
@@ -156,7 +162,11 @@ def plot_classification_report(
cmap = "YlOrRd"
clf_report = classification_report(y_true, y_pred, output_dict=True, **kwargs)
- keys_to_plot = [key for key in clf_report.keys() if key not in ("accuracy", "macro avg", "weighted avg")]
+ keys_to_plot = [
+ key
+ for key in clf_report.keys()
+ if key not in ("accuracy", "macro avg", "weighted avg")
+ ]
df = pd.DataFrame(clf_report, columns=keys_to_plot).T
# the following line ensures that dataframe are sorted from the majority classes to the minority classes
df.sort_values(by=["support"], inplace=True)
@@ -325,7 +335,9 @@ def plot_roc_curve(
auc_upper = np.quantile(bootstrap_aucs, CI_upper)
auc_lower = np.quantile(bootstrap_aucs, CI_lower)
label = f"{confidence_interval:.0%} CI: [{auc_lower:.2f}, {auc_upper:.2f}]"
- plt.fill_between(base_fpr, tprs_lower, tprs_upper, alpha=0.3, label=label, zorder=2)
+ plt.fill_between(
+ base_fpr, tprs_lower, tprs_upper, alpha=0.3, label=label, zorder=2
+ )
if highlight_roc_area is True:
print(
@@ -357,7 +369,9 @@ def plot_roc_curve(
return fig
-def plot_calibration_curve(y_true: np.ndarray, y_score: np.ndarray, n_bins=10, save_fig_path=None) -> Figure:
+def plot_calibration_curve(
+ y_true: np.ndarray, y_score: np.ndarray, n_bins=10, save_fig_path=None
+) -> Figure:
"""
Creates calibration plot for a binary classifier and calculates the ECE.
@@ -379,7 +393,9 @@ def plot_calibration_curve(y_true: np.ndarray, y_score: np.ndarray, n_bins=10, s
ece : float
The expected calibration error.
"""
- prob_true, prob_pred = calibration_curve(y_true, y_score, n_bins=n_bins, strategy="uniform")
+ prob_true, prob_pred = calibration_curve(
+ y_true, y_score, n_bins=n_bins, strategy="uniform"
+ )
# Find the number of samples in each bin
bin_counts = np.histogram(y_score, bins=n_bins, range=(0, 1))[0]
@@ -452,7 +468,9 @@ def plot_calibration_curve(y_true: np.ndarray, y_score: np.ndarray, n_bins=10, s
return fig
-def plot_y_score_histogram(y_true: Optional[np.ndarray], y_score: np.ndarray = None, save_fig_path=None) -> Figure:
+def plot_y_score_histogram(
+ y_true: Optional[np.ndarray], y_score: np.ndarray = None, save_fig_path=None
+) -> Figure:
"""
Provides a histogram for the predicted probabilities of a binary classifier. If ```y_true``` is provided, it divides the ```y_score``` values into the two classes and plots them jointly into the same plot with different colors.
@@ -474,7 +492,9 @@ def plot_y_score_histogram(y_true: Optional[np.ndarray], y_score: np.ndarray = N
ax = fig.add_subplot(111)
if y_true is None:
- ax.hist(y_score, bins=10, alpha=0.9, edgecolor="midnightblue", linewidth=2, rwidth=1)
+ ax.hist(
+ y_score, bins=10, alpha=0.9, edgecolor="midnightblue", linewidth=2, rwidth=1
+ )
# same histogram as above, but with border lines
# ax.hist(y_prob, bins=10, alpha=0.5, edgecolor='black', linewidth=1.2)
else:
diff --git a/plotsandgraphs/multiclass_classifier.py b/plotsandgraphs/multiclass_classifier.py
index af1c019..c105373 100644
--- a/plotsandgraphs/multiclass_classifier.py
+++ b/plotsandgraphs/multiclass_classifier.py
@@ -20,7 +20,12 @@
from sklearn.utils import resample
from tqdm import tqdm
-from plotsandgraphs.utils import bootstrap, set_black_title_boxes, scale_ax_bbox, get_cmap
+from plotsandgraphs.utils import (
+ bootstrap,
+ set_black_title_boxes,
+ scale_ax_bbox,
+ get_cmap,
+)
def plot_roc_curve(
@@ -32,7 +37,7 @@ def plot_roc_curve(
figsize: Optional[Tuple[float, float]] = None,
class_labels: Optional[List[str]] = None,
split_plots: bool = False,
- save_fig_path:Optional[Union[str, Tuple[str, str]]] = None,
+ save_fig_path: Optional[Union[str, Tuple[str, str]]] = None,
) -> Tuple[Figure, Union[Figure, None]]:
"""
Creates two plots.
@@ -190,9 +195,6 @@ def roc_metric_function(y_true, y_score):
# create the subplot tiles (and black boxes)
set_black_title_boxes(axes.flat[:num_classes], class_labels)
-
-
-
# ---------- AUROC overview plot comparing classes ----------
# Make an AUROC overview plot comparing the aurocs per class and combined
@@ -268,10 +270,12 @@ def auroc_metric_function(y_true, y_score, average, multi_class):
def plot_y_score_histogram(
- y_true: np.ndarray, y_score: Optional[np.ndarray] = None, save_fig_path: Optional[str]=None
+ y_true: np.ndarray,
+ y_score: Optional[np.ndarray] = None,
+ save_fig_path: Optional[str] = None,
) -> Figure:
"""
- Histogram plot that is intended to show the distribution of the predicted probabilities for different classes, where the the different classes (y_true==0 and y_true==1) are plotted in different colors.
+ Histogram plot that is intended to show the distribution of the predicted probabilities for different classes, where the the different classes (y_true==0 and y_true==1) are plotted in different colors.
Limitations: Does not work for samples, that can be part of multiple classes (e.g. multilabel classification).
Parameters
@@ -288,15 +292,19 @@ def plot_y_score_histogram(
Figure
The figure of the histogram plot.
"""
-
+
num_classes = y_true.shape[-1]
class_labels = [f"Class {i}" for i in range(num_classes)]
-
+
cmap, colors = get_cmap("roma", n_colors=2) # 2 colors for y==0 and y==1 per class
-
+
# Aiming for a square plot
- plot_cols = np.ceil(np.sqrt(num_classes)).astype(int) # Number of plots in a row # noqa
- plot_rows = np.ceil(num_classes / plot_cols).astype(int) # Number of plots in a column # noqa
+ plot_cols = np.ceil(np.sqrt(num_classes)).astype(
+ int
+ ) # Number of plots in a row # noqa
+ plot_rows = np.ceil(num_classes / plot_cols).astype(
+ int
+ ) # Number of plots in a column # noqa
fig, axes = plt.subplots(
nrows=plot_rows,
ncols=plot_cols,
diff --git a/plotsandgraphs/pipeline.py b/plotsandgraphs/pipeline.py
index 00eb099..24c8844 100644
--- a/plotsandgraphs/pipeline.py
+++ b/plotsandgraphs/pipeline.py
@@ -6,109 +6,105 @@
from . import multiclass_classifier as mc
+FILE_ENDINGS = Literal["pdf", "png", "jpg", "jpeg", "svg"]
-FILE_ENDINGS = Literal['pdf', 'png', 'jpg', 'jpeg', 'svg']
-
-
-def binary_classifier(y_true, y_score, save_fig_path=None, plot_kwargs={}, file_type:FILE_ENDINGS='png'):
-
-
+def binary_classifier(
+ y_true, y_score, save_fig_path=None, plot_kwargs={}, file_type: FILE_ENDINGS = "png"
+):
# Create new tqdm instance
- tqdm_instance = tqdm(total=6, desc='Binary classifier metrics', leave=True)
-
+ tqdm_instance = tqdm(total=6, desc="Binary classifier metrics", leave=True)
+
# Update tqdm instance
tqdm_instance.update()
-
+
# 1) Plot ROC curve
- roc_kwargs = plot_kwargs.get('roc', {})
- save_path = get_file_path(save_fig_path, 'roc_curve', file_type)
+ roc_kwargs = plot_kwargs.get("roc", {})
+ save_path = get_file_path(save_fig_path, "roc_curve", file_type)
bc.plot_roc_curve(y_true, y_score, save_fig_path=save_path, **roc_kwargs)
tqdm_instance.update()
-
+
# 2) Plot precision-recall curve
- pr_kwargs = plot_kwargs.get('pr', {})
- save_path = get_file_path(save_fig_path, 'pr_curve', file_type)
+ pr_kwargs = plot_kwargs.get("pr", {})
+ save_path = get_file_path(save_fig_path, "pr_curve", file_type)
bc.plot_pr_curve(y_true, y_score, save_fig_path=save_path, **pr_kwargs)
tqdm_instance.update()
-
+
# 3) Plot calibration curve
- cal_kwargs = plot_kwargs.get('cal', {})
- save_path = get_file_path(save_fig_path, 'calibration_curve', file_type)
+ cal_kwargs = plot_kwargs.get("cal", {})
+ save_path = get_file_path(save_fig_path, "calibration_curve", file_type)
bc.plot_calibration_curve(y_true, y_score, save_fig_path=save_path, **cal_kwargs)
tqdm_instance.update()
-
+
# 3) Plot confusion matrix
- cm_kwargs = plot_kwargs.get('cm', {})
- save_path = get_file_path(save_fig_path, 'confusion_matrix', file_type)
+ cm_kwargs = plot_kwargs.get("cm", {})
+ save_path = get_file_path(save_fig_path, "confusion_matrix", file_type)
bc.plot_confusion_matrix(y_true, y_score, save_fig_path=save_path, **cm_kwargs)
tqdm_instance.update()
-
+
# 5) Plot classification report
- cr_kwargs = plot_kwargs.get('cr', {})
- save_path = get_file_path(save_fig_path, 'classification_report', file_type)
+ cr_kwargs = plot_kwargs.get("cr", {})
+ save_path = get_file_path(save_fig_path, "classification_report", file_type)
bc.plot_classification_report(y_true, y_score, save_fig_path=save_path, **cr_kwargs)
tqdm_instance.update()
-
+
# 6) Plot y_score histogram
- hist_kwargs = plot_kwargs.get('hist', {})
- save_path = get_file_path(save_fig_path, 'y_score_histogram', file_type)
+ hist_kwargs = plot_kwargs.get("hist", {})
+ save_path = get_file_path(save_fig_path, "y_score_histogram", file_type)
bc.plot_y_score_histogram(y_true, y_score, save_fig_path=save_path, **hist_kwargs)
tqdm_instance.update()
-
- return
-
+ return
-def multiclass_classifier(y_true, y_score, save_fig_path=None, plot_kwargs={}, file_type:FILE_ENDINGS = 'png'):
-
+def multiclass_classifier(
+ y_true, y_score, save_fig_path=None, plot_kwargs={}, file_type: FILE_ENDINGS = "png"
+):
# Create new tqdm instance
- tqdm_instance = tqdm(total=6, desc='Binary classifier metrics', leave=True)
-
+ tqdm_instance = tqdm(total=6, desc="Binary classifier metrics", leave=True)
+
# Update tqdm instance
tqdm_instance.update()
-
+
# 1) Plot ROC curve
- roc_kwargs = plot_kwargs.get('roc', {})
- save_path = get_file_path(save_fig_path, 'roc_curve', '')
+ roc_kwargs = plot_kwargs.get("roc", {})
+ save_path = get_file_path(save_fig_path, "roc_curve", "")
mc.plot_roc_curve(y_true, y_score, save_fig_path=save_path, **roc_kwargs)
tqdm_instance.update()
-
+
# 2) Plot precision-recall curve
# pr_kwargs = plot_kwargs.get('pr', {})
# mc.plot_pr_curve(y_true, y_score, save_fig_path=save_fig_path, **pr_kwargs)
# tqdm_instance.update()
-
+
# 3) Plot calibration curve
# cal_kwargs = plot_kwargs.get('cal', {})
# mc.plot_calibration_curve(y_true, y_score, save_fig_path=save_fig_path, **cal_kwargs)
# tqdm_instance.update()
-
+
# 3) Plot confusion matrix
# cm_kwargs = plot_kwargs.get('cm', {})
# mc.plot_confusion_matrix(y_true, y_score, save_fig_path=save_fig_path, **cm_kwargs)
# tqdm_instance.update()
-
+
# 5) Plot classification report
# cr_kwargs = plot_kwargs.get('cr', {})
# mc.plot_classification_report(y_true, y_score, save_fig_path=save_fig_path, **cr_kwargs)
# tqdm_instance.update()
-
+
# 6) Plot y_score histogram
- hist_kwargs = plot_kwargs.get('hist', {})
- save_path = get_file_path(save_fig_path, 'y_score_histogram', file_type)
+ hist_kwargs = plot_kwargs.get("hist", {})
+ save_path = get_file_path(save_fig_path, "y_score_histogram", file_type)
mc.plot_y_score_histogram(y_true, y_score, save_fig_path=save_path, **hist_kwargs)
tqdm_instance.update()
-
- return
+ return
-def get_file_path(save_fig_path: Union[Path,None, str], name:str, ending:str):
+def get_file_path(save_fig_path: Union[Path, None, str], name: str, ending: str):
if save_fig_path is None:
return None
else:
result = Path(save_fig_path) / f"{name}.{ending}"
print(result)
- return str(result)
\ No newline at end of file
+ return str(result)
diff --git a/plotsandgraphs/utils.py b/plotsandgraphs/utils.py
index 65b801d..b8c2a32 100644
--- a/plotsandgraphs/utils.py
+++ b/plotsandgraphs/utils.py
@@ -1,4 +1,4 @@
-from typing import Optional, List, Callable, Dict, Tuple, Union, TYPE_CHECKING, Literal
+from typing import Optional, List, Callable, Dict, Tuple, TYPE_CHECKING
from tqdm import tqdm
from sklearn.utils import resample
import numpy as np
@@ -11,7 +11,12 @@
from matplotlib.axes import Axes
-def bootstrap(metric_function: Callable, input_resample: List[np.ndarray], n_bootstraps: int, metric_kwargs: Dict={}) -> List:
+def bootstrap(
+ metric_function: Callable,
+ input_resample: List[np.ndarray],
+ n_bootstraps: int,
+ metric_kwargs: Dict = {},
+) -> List:
"""
A bootstrapping function for a metric function. The metric function should take the same number of arguments as the length of input_resample.
@@ -33,29 +38,28 @@ def bootstrap(metric_function: Callable, input_resample: List[np.ndarray], n_boo
"""
results = []
# for each bootstrap iteration
- for _ in tqdm(range(n_bootstraps), desc='Bootsrapping', leave=True):
+ for _ in tqdm(range(n_bootstraps), desc="Bootsrapping", leave=True):
# resample indices with replacement
indices = resample(np.arange(len(input_resample[0])), replace=True)
input_resampled = [x[indices] for x in input_resample]
# calculate metric
result = metric_function(*input_resampled, **metric_kwargs)
-
+
results.append(result)
-
- return results
+ return results
class ExtendedTextBox_v2:
"""
Black background boxes for titles in maptlolib subplots
-
+
From:
https://stackoverflow.com/questions/40796117/how-do-i-make-the-width-of-the-title-box-span-the-entire-plot
https://matplotlib.org/stable/gallery/userdemo/custom_boxstyle01.html?highlight=boxstyle+_style_list
"""
- def __init__(self, pad=0.3, width=500.):
+ def __init__(self, pad=0.3, width=500.0):
"""
The arguments must be floats and have default values.
@@ -85,22 +89,25 @@ def __call__(self, x0, y0, width, height, mutation_size):
# padding
pad = mutation_size * self.pad
# width and height with padding added
- #width = width + 2.*pad
- height = height + 3 * pad
+ # width = width + 2.*pad
+ height = height + 3 * pad
# boundary of the padded box
y0 = y0 - pad # change this to move the text
- y1 = y0 + height
+ y1 = y0 + height
_x0 = x0
- x0 = _x0 +width /2. - self.width/2.
- x1 = _x0 +width /2. + self.width/2.
+ x0 = _x0 + width / 2.0 - self.width / 2.0
+ x1 = _x0 + width / 2.0 + self.width / 2.0
# return the new path
- return Path([(x0, y0),
- (x1, y0), (x1, y1), (x0, y1),
- (x0, y0)],
- closed=True)
+ return Path([(x0, y0), (x1, y0), (x1, y1), (x0, y1), (x0, y0)], closed=True)
-def _set_black_title_box(ax: "Axes", title:str, backgroundcolor='black', color='white', title_kwargs: Optional[Dict]=None):
+def _set_black_title_box(
+ ax: "Axes",
+ title: str,
+ backgroundcolor="black",
+ color="white",
+ title_kwargs: Optional[Dict] = None,
+):
"""
Note: Do not use this function by itself, instead use `set_black_title_boxes()`.
Sets the title of the given axes with a black bounding box.
@@ -114,16 +121,25 @@ def _set_black_title_box(ax: "Axes", title:str, backgroundcolor='black', color='
- set_title_kwargs: Keyword arguments to pass to `ax.set_title()`.
"""
if title_kwargs is None:
- title_kwargs = {'fontdict': {"fontname": "Arial Black", "fontweight": "bold"}}
- BoxStyle._style_list["ext"] = ExtendedTextBox_v2
+ title_kwargs = {"fontdict": {"fontname": "Arial Black", "fontweight": "bold"}}
+ BoxStyle._style_list["ext"] = ExtendedTextBox_v2
ax_width = ax.get_window_extent().width
# make title with black bounding box
- title_instance = ax.set_title(title, backgroundcolor=backgroundcolor, color=color, **title_kwargs)
- bb = title_instance.get_bbox_patch() # get bbox from title
- bb.set_boxstyle("ext", pad=0.1, width=ax_width) # use custom style
-
-
-def set_black_title_boxes(axes: "np.ndarray[Axes]", titles: List[str], backgroundcolor='black', color='white', title_kwargs: Optional[Dict]=None, tight_layout_kwargs: Dict={}):
+ title_instance = ax.set_title(
+ title, backgroundcolor=backgroundcolor, color=color, **title_kwargs
+ )
+ bb = title_instance.get_bbox_patch() # get bbox from title
+ bb.set_boxstyle("ext", pad=0.1, width=ax_width) # use custom style
+
+
+def set_black_title_boxes(
+ axes: "np.ndarray[Axes]",
+ titles: List[str],
+ backgroundcolor="black",
+ color="white",
+ title_kwargs: Optional[Dict] = None,
+ tight_layout_kwargs: Dict = {},
+):
"""
Creates black boxes for the subtitles above the given axes with the given titles. The subtitles are centered above the axes.
@@ -145,18 +161,15 @@ def set_black_title_boxes(axes: "np.ndarray[Axes]", titles: List[str], backgroun
for i, ax in enumerate(axes.flat):
_set_black_title_box(ax, titles[i], backgroundcolor, color, title_kwargs)
-
+
plt.tight_layout(**tight_layout_kwargs)
-
+
for i, ax in enumerate(axes.flat):
_set_black_title_box(ax, titles[i], backgroundcolor, color, title_kwargs)
-
-
+
return
-
-
-
-
+
+
def scale_ax_bbox(ax: "Axes", factor: float):
# Get the current position of the subplot
box = ax.get_position()
@@ -167,12 +180,13 @@ def scale_ax_bbox(ax: "Axes", factor: float):
# Set the new position
ax.set_position([box.x0 + adjustment, box.y0, new_width, box.height])
-
- return
+ return
-def get_cmap(cmap_name: str, n_colors: Optional[int]=None) -> Tuple[LinearSegmentedColormap, Tuple]:
+def get_cmap(
+ cmap_name: str, n_colors: Optional[int] = None
+) -> Tuple[LinearSegmentedColormap, Tuple]:
"""
Loads one of the custom cmaps from the cmaps folder.
@@ -185,23 +199,19 @@ def get_cmap(cmap_name: str, n_colors: Optional[int]=None) -> Tuple[LinearSegmen
-------
Tuple[LinearSegmentedColormap, Union[None, Tuple]]
A tuple of the cmap and a list of colors if n_colors is not None.
-
+
Example
-------
>>> cmap_name = 'hawaii'
>>> cmap, color_list = get_cmap(cmap_name, n_colors=10)
"""
from pathlib import Path as PathClass
-
- cm_path = PathClass(__file__).parent / ('cmaps/' + cmap_name + '.txt')
+
+ cm_path = PathClass(__file__).parent / ("cmaps/" + cmap_name + ".txt")
cm_data = np.loadtxt(cm_path)
- cmap_name = cmap_name.split('.')[0]
+ cmap_name = cmap_name.split(".")[0]
cmap = LinearSegmentedColormap.from_list(cmap_name, cm_data)
if n_colors is None:
n_colors = 10
color_list = cmap(np.linspace(0, 1, n_colors))
return cmap, color_list
-
-
-
-
\ No newline at end of file
diff --git a/tests/test_binary_classifier.py b/tests/test_binary_classifier.py
index a8e72b4..9426287 100644
--- a/tests/test_binary_classifier.py
+++ b/tests/test_binary_classifier.py
@@ -41,8 +41,12 @@ def test_hist_plot(random_data_binary_classifier):
"""
y_true, y_score = random_data_binary_classifier
print(TEST_RESULTS_PATH)
- binary.plot_y_score_histogram(y_true=None, y_score=y_score, save_fig_path=TEST_RESULTS_PATH / "histogram.png")
- binary.plot_y_score_histogram(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "histogram_2_classes.png")
+ binary.plot_y_score_histogram(
+ y_true=None, y_score=y_score, save_fig_path=TEST_RESULTS_PATH / "histogram.png"
+ )
+ binary.plot_y_score_histogram(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "histogram_2_classes.png"
+ )
# test roc curve without bootstrapping
@@ -56,7 +60,9 @@ def test_roc_curve(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_roc_curve(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "roc_curve.png")
+ binary.plot_roc_curve(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "roc_curve.png"
+ )
# test roc curve with bootstrapping
@@ -71,7 +77,10 @@ def test_roc_curve_bootstrap(random_data_binary_classifier):
"""
y_true, y_score = random_data_binary_classifier
binary.plot_roc_curve(
- y_true, y_score, n_bootstraps=10000, save_fig_path=TEST_RESULTS_PATH / "roc_curve_bootstrap.png"
+ y_true,
+ y_score,
+ n_bootstraps=10000,
+ save_fig_path=TEST_RESULTS_PATH / "roc_curve_bootstrap.png",
)
@@ -86,7 +95,9 @@ def test_pr_curve(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_pr_curve(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "pr_curve.png")
+ binary.plot_pr_curve(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "pr_curve.png"
+ )
# test confusion matrix
@@ -100,7 +111,9 @@ def test_confusion_matrix(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_confusion_matrix(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "confusion_matrix.png")
+ binary.plot_confusion_matrix(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "confusion_matrix.png"
+ )
# test classification report
@@ -114,7 +127,10 @@ def test_classification_report(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_classification_report(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "classification_report.png")
+ binary.plot_classification_report(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "classification_report.png"
+ )
+
# test calibration curve
def test_calibration_curve(random_data_binary_classifier):
@@ -127,7 +143,10 @@ def test_calibration_curve(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_calibration_curve(y_score, y_true, save_fig_path=TEST_RESULTS_PATH / "calibration_curve.png")
+ binary.plot_calibration_curve(
+ y_score, y_true, save_fig_path=TEST_RESULTS_PATH / "calibration_curve.png"
+ )
+
# test accuracy
def test_accuracy(random_data_binary_classifier):
@@ -140,4 +159,6 @@ def test_accuracy(random_data_binary_classifier):
The simulated data.
"""
y_true, y_score = random_data_binary_classifier
- binary.plot_accuracy(y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "accuracy.png")
+ binary.plot_accuracy(
+ y_true, y_score, save_fig_path=TEST_RESULTS_PATH / "accuracy.png"
+ )
diff --git a/tests/test_multiclass_classifier.py b/tests/test_multiclass_classifier.py
index d47e7e0..3cbdb70 100644
--- a/tests/test_multiclass_classifier.py
+++ b/tests/test_multiclass_classifier.py
@@ -3,6 +3,7 @@
from itertools import product
import numpy as np
import pytest
+
# import plotsandgraphs.binary_classifier as binary
import plotsandgraphs.multiclass_classifier as multiclass
@@ -10,7 +11,9 @@
# @pytest.fixture(scope="module")
-def random_data_multiclass_classifier(num_classes:int = 3) -> Tuple[np.ndarray, np.ndarray]:
+def random_data_multiclass_classifier(
+ num_classes: int = 3,
+) -> Tuple[np.ndarray, np.ndarray]:
"""
Create random data for binary classifier tests.
@@ -21,26 +24,26 @@ def random_data_multiclass_classifier(num_classes:int = 3) -> Tuple[np.ndarray,
"""
class_labels = np.arange(num_classes)
class_probs = np.random.random(num_classes)
- class_probs = class_probs / class_probs.sum() # normalize
+ class_probs = class_probs / class_probs.sum() # normalize
# True labels
y_true = np.random.choice(class_labels, p=class_probs, size=1000)
# one hot encoding
- y_true_one_hot = np.eye(num_classes)[y_true]
+ y_true_one_hot = np.eye(num_classes)[y_true]
# Predicted labels
y_pred = np.ones(y_true_one_hot.shape)
# parameters for Beta distribution for each label (a0,b0 for class 0, a1,b1 for class 1)
- a0, b0 = [0.1, 0.6, 0.3, 0.4, 2]*10, [0.4, 1.2, 0.8, 1, 5]*10
- a1, b1 = [0.9, 0.8, 0.9, 1.2, 5]*10, [0.4, 0.1, 0.5, 0.3, 2]*10
+ a0, b0 = [0.1, 0.6, 0.3, 0.4, 2] * 10, [0.4, 1.2, 0.8, 1, 5] * 10
+ a1, b1 = [0.9, 0.8, 0.9, 1.2, 5] * 10, [0.4, 0.1, 0.5, 0.3, 2] * 10
# iterate through all the columns/labels and create a beta distribution for each label
for i in range(y_pred.shape[1]):
y = y_pred[:, i]
y_t = y_true_one_hot[:, i]
- y[y_t==0] = np.random.beta(a0[i], b0[i], size=y[y_t==0].shape)
- y[y_t==1] = np.random.beta(a1[i], b1[i], size=y[y_t==1].shape)
-
+ y[y_t == 0] = np.random.beta(a0[i], b0[i], size=y[y_t == 0].shape)
+ y[y_t == 1] = np.random.beta(a1[i], b1[i], size=y[y_t == 1].shape)
+
return y_true_one_hot, y_pred
@@ -57,42 +60,71 @@ def test_hist_plot():
for num_classes in [1, 2, 3, 4, 5, 10, 16, 25]:
y_true, y_prob = random_data_multiclass_classifier(num_classes=num_classes)
print(TEST_RESULTS_PATH)
- multiclass.plot_y_prob_histogram(y_true=y_true, y_score=y_prob, save_fig_path=TEST_RESULTS_PATH / f"histogram_{num_classes}_classes.png")
+ multiclass.plot_y_prob_histogram(
+ y_true=y_true,
+ y_score=y_prob,
+ save_fig_path=TEST_RESULTS_PATH / f"histogram_{num_classes}_classes.png",
+ )
# multiclass.plot_y_prob_histogram(y_prob=y_prob, save_fig_path=TEST_RESULTS_PATH / "histogram_classes.png")
-
-
+
+
def test_roc_curve():
"""
Test roc curve.
-
+
Parameters
----------
random_data_binary_classifier : Tuple[np.ndarray, np.ndarray]
The simulated data.
"""
+
# helper function for file name, to avoid repeating code
- def get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots):
+ def get_path_name(
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots
+ ):
if split_plots is False:
- fig_path = TEST_RESULTS_PATH / f"roc_curves_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ fig_path = (
+ TEST_RESULTS_PATH
+ / f"roc_curves_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ )
else:
- fig_path_1 = TEST_RESULTS_PATH / f"roc_curves_split_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
- fig_path_2 = TEST_RESULTS_PATH / f"auroc_comparison_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ fig_path_1 = (
+ TEST_RESULTS_PATH
+ / f"roc_curves_split_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ )
+ fig_path_2 = (
+ TEST_RESULTS_PATH
+ / f"auroc_comparison_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
+ )
fig_path = [fig_path_1, fig_path_2]
return fig_path
-
-
+
y_true, y_prob = random_data_multiclass_classifier(num_classes=3)
-
+
confidence_intervals = [None, 0.99]
highlight_roc_area = [True, False]
n_bootstraps = [1, 1000]
figsizes = [None]
split_plots = [True, False]
-
- # From the previous lists I want all possible combinations
- combinations = list(product(confidence_intervals, highlight_roc_area, n_bootstraps, figsizes, split_plots))
-
- for confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots in combinations:
+
+ # From the previous lists I want all possible combinations
+ combinations = list(
+ product(
+ confidence_intervals,
+ highlight_roc_area,
+ n_bootstraps,
+ figsizes,
+ split_plots,
+ )
+ )
+
+ for (
+ confidence_interval,
+ highlight_roc_area,
+ n_bootstraps,
+ figsize,
+ split_plots,
+ ) in combinations:
# check if one or two figures should be saved (splot_plots=True or False)
# if split_plots is False:
# fig_path = TEST_RESULTS_PATH / f"roc_curves_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
@@ -100,51 +132,77 @@ def get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize
# fig_path_1 = TEST_RESULTS_PATH / f"roc_curves_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
# fig_path_2 = TEST_RESULTS_PATH / f"auroc_comparison_conf_{confidence_interval}_highlight_{highlight_roc_area}_nboot_{n_bootstraps}_figsize_{figsize}.png"
# fig_path = [fig_path_1, fig_path_2]
-
- fig_path = get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots)
-
+
+ fig_path = get_path_name(
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots
+ )
+
# It should raise an error when confidence_interval is None but highlight_roc_area is True
if confidence_interval is None and highlight_roc_area is True:
with pytest.raises(ValueError):
- multiclass.plot_roc_curve(y_true=y_true, y_score=y_prob,
- confidence_interval=confidence_interval,
- highlight_roc_area=highlight_roc_area,
- n_bootstraps=n_bootstraps,
- figsize=figsize,
- split_plots=split_plots,
- save_fig_path=fig_path)
+ multiclass.plot_roc_curve(
+ y_true=y_true,
+ y_score=y_prob,
+ confidence_interval=confidence_interval,
+ highlight_roc_area=highlight_roc_area,
+ n_bootstraps=n_bootstraps,
+ figsize=figsize,
+ split_plots=split_plots,
+ save_fig_path=fig_path,
+ )
# Otherwise no error
else:
- multiclass.plot_roc_curve(y_true=y_true, y_score=y_prob,
- confidence_interval=confidence_interval,
- highlight_roc_area=highlight_roc_area,
- n_bootstraps=n_bootstraps,
- figsize=figsize,
- split_plots=split_plots,
- save_fig_path=fig_path)
-
- # check for SMALL figure size
- confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots = 0.95, True, 100, (3,3), False
- fig_path = get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots)
- multiclass.plot_roc_curve(y_true=y_true, y_score=y_prob,
- confidence_interval=confidence_interval,
- highlight_roc_area=highlight_roc_area,
- n_bootstraps=n_bootstraps,
- figsize=figsize,
- split_plots=split_plots,
- save_fig_path=fig_path)
-
- # check for BIG figure size
- confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots = 0.95, True, 100, (15, 15), False
- fig_path = get_path_name(confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots)
- multiclass.plot_roc_curve(y_true=y_true, y_score=y_prob,
- confidence_interval=confidence_interval,
- highlight_roc_area=highlight_roc_area,
- n_bootstraps=n_bootstraps,
- figsize=figsize,
- split_plots=split_plots,
- save_fig_path=fig_path)
-
-
+ multiclass.plot_roc_curve(
+ y_true=y_true,
+ y_score=y_prob,
+ confidence_interval=confidence_interval,
+ highlight_roc_area=highlight_roc_area,
+ n_bootstraps=n_bootstraps,
+ figsize=figsize,
+ split_plots=split_plots,
+ save_fig_path=fig_path,
+ )
+ # check for SMALL figure size
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots = (
+ 0.95,
+ True,
+ 100,
+ (3, 3),
+ False,
+ )
+ fig_path = get_path_name(
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots
+ )
+ multiclass.plot_roc_curve(
+ y_true=y_true,
+ y_score=y_prob,
+ confidence_interval=confidence_interval,
+ highlight_roc_area=highlight_roc_area,
+ n_bootstraps=n_bootstraps,
+ figsize=figsize,
+ split_plots=split_plots,
+ save_fig_path=fig_path,
+ )
+ # check for BIG figure size
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots = (
+ 0.95,
+ True,
+ 100,
+ (15, 15),
+ False,
+ )
+ fig_path = get_path_name(
+ confidence_interval, highlight_roc_area, n_bootstraps, figsize, split_plots
+ )
+ multiclass.plot_roc_curve(
+ y_true=y_true,
+ y_score=y_prob,
+ confidence_interval=confidence_interval,
+ highlight_roc_area=highlight_roc_area,
+ n_bootstraps=n_bootstraps,
+ figsize=figsize,
+ split_plots=split_plots,
+ save_fig_path=fig_path,
+ )
diff --git a/tests/test_pipeline.py b/tests/test_pipeline.py
index 1ad1fe3..dd9cdaf 100644
--- a/tests/test_pipeline.py
+++ b/tests/test_pipeline.py
@@ -1,7 +1,4 @@
from pathlib import Path
-from typing import Tuple
-import numpy as np
-import pytest
import shutil
from plotsandgraphs import pipeline
@@ -11,6 +8,7 @@
TEST_RESULTS_PATH = Path("tests/test_results/pipeline")
+
def test_binary_classification_pipeline(random_data_binary_classifier):
"""
Test binary classification pipeline.
@@ -21,36 +19,40 @@ def test_binary_classification_pipeline(random_data_binary_classifier):
The simulated data.
"""
save_fig_path = TEST_RESULTS_PATH / "binary_classifier"
-
+
# Delete the folder and its previous contents
if save_fig_path.exists() and save_fig_path.is_dir():
shutil.rmtree(save_fig_path)
y_true, y_score = random_data_binary_classifier
- pipeline.binary_classifier(y_true, y_score, save_fig_path=save_fig_path, file_type='png')
-
+ pipeline.binary_classifier(
+ y_true, y_score, save_fig_path=save_fig_path, file_type="png"
+ )
+
# assert that there are files with the names xy in the save_fig_path
assert (save_fig_path / "roc_curve.png").exists()
assert (save_fig_path / "y_score_histogram.png").exists()
assert (save_fig_path / "calibration_curve.png").exists()
assert (save_fig_path / "confusion_matrix.png").exists()
assert (save_fig_path / "pr_curve.png").exists()
-
-
+
+
def test_multiclassification_pipeline():
"""
Test multiclassification pipeline.
"""
for num_classes in [3]:
save_fig_path = TEST_RESULTS_PATH / f"multiclass_{num_classes}_classes"
-
+
# Delete the folder and its previous contents
if save_fig_path.exists() and save_fig_path.is_dir():
shutil.rmtree(save_fig_path)
-
+
y_true, y_score = random_data_multiclass_classifier(num_classes=num_classes)
- pipeline.multiclass_classifier(y_true, y_score, save_fig_path=save_fig_path, file_type='png')
-
+ pipeline.multiclass_classifier(
+ y_true, y_score, save_fig_path=save_fig_path, file_type="png"
+ )
+
# assert that there are files with the names xy in the save_fig_path
assert (save_fig_path / "roc_curve.png").exists()
- assert (save_fig_path / "y_score_histogram.png").exists()
\ No newline at end of file
+ assert (save_fig_path / "y_score_histogram.png").exists()
diff --git a/tests/utils.py b/tests/utils.py
index ee27bdb..780b451 100644
--- a/tests/utils.py
+++ b/tests/utils.py
@@ -25,7 +25,9 @@ def random_data_binary_classifier() -> Tuple[np.ndarray, np.ndarray]:
return y_true, y_score
-def random_data_multiclass_classifier(num_classes:int = 3) -> Tuple[np.ndarray, np.ndarray]:
+def random_data_multiclass_classifier(
+ num_classes: int = 3,
+) -> Tuple[np.ndarray, np.ndarray]:
"""
Create random data for binary classifier tests.
@@ -36,24 +38,24 @@ def random_data_multiclass_classifier(num_classes:int = 3) -> Tuple[np.ndarray,
"""
class_labels = np.arange(num_classes)
class_probs = np.random.random(num_classes)
- class_probs = class_probs / class_probs.sum() # normalize
+ class_probs = class_probs / class_probs.sum() # normalize
# True labels
y_true = np.random.choice(class_labels, p=class_probs, size=1000)
# one hot encoding
- y_true_one_hot = np.eye(num_classes)[y_true]
+ y_true_one_hot = np.eye(num_classes)[y_true]
# Predicted labels
y_pred = np.ones(y_true_one_hot.shape)
# parameters for Beta distribution for each label (a0,b0 for class 0, a1,b1 for class 1)
- a0, b0 = [0.1, 0.6, 0.3, 0.4, 2]*10, [0.4, 1.2, 0.8, 1, 5]*10
- a1, b1 = [0.9, 0.8, 0.9, 1.2, 5]*10, [0.4, 0.1, 0.5, 0.3, 2]*10
+ a0, b0 = [0.1, 0.6, 0.3, 0.4, 2] * 10, [0.4, 1.2, 0.8, 1, 5] * 10
+ a1, b1 = [0.9, 0.8, 0.9, 1.2, 5] * 10, [0.4, 0.1, 0.5, 0.3, 2] * 10
# iterate through all the columns/labels and create a beta distribution for each label
for i in range(y_pred.shape[1]):
y = y_pred[:, i]
y_t = y_true_one_hot[:, i]
- y[y_t==0] = np.random.beta(a0[i], b0[i], size=y[y_t==0].shape)
- y[y_t==1] = np.random.beta(a1[i], b1[i], size=y[y_t==1].shape)
-
- return y_true_one_hot, y_pred
\ No newline at end of file
+ y[y_t == 0] = np.random.beta(a0[i], b0[i], size=y[y_t == 0].shape)
+ y[y_t == 1] = np.random.beta(a1[i], b1[i], size=y[y_t == 1].shape)
+
+ return y_true_one_hot, y_pred
 |
|  |
|  |
|:--------------------------------------------------:|:----------------------------------------------------------:|:-------------------------------------------------:|
-| ROC Curve (AUROC) with bootstrapping | Precision-Recall Curve | y_prob histogram |
+| ROC Curve (AUROC) with bootstrapping | Precision-Recall Curve | y_score histogram |
|
|
|:--------------------------------------------------:|:----------------------------------------------------------:|:-------------------------------------------------:|
-| ROC Curve (AUROC) with bootstrapping | Precision-Recall Curve | y_prob histogram |
+| ROC Curve (AUROC) with bootstrapping | Precision-Recall Curve | y_score histogram |
|  |
|  |
|