-nat: NeuroAnatomy Toolbox
@@ -248,19 +247,7 @@
-
+
nat: NeuroAnatomy Toolbox 
 +
-
-
+
+
-
-
+
An R package for the (3D) visualisation and analysis of biological image data, especially tracings of single neurons. nat is the core package of a wider suite of neuroanatomy tools introduced at http://natverse.github.io. nat (and its ancestors) have been used in a number of papers from our group including:
- + [](https://natverse.org/)
[](http://dx.doi.org/10.5281/zenodo.10171)
@@ -17,7 +19,7 @@ have been used in a number of papers from our group including:
[](http://dx.doi.org/10.1016/j.cell.2007.01.040)
[](http://dx.doi.org/10.1016/j.cub.2010.07.045)
-[](http://dx.doi.org/10.1038/nature10428)
+[
[](https://natverse.org/)
[](http://dx.doi.org/10.5281/zenodo.10171)
@@ -17,7 +19,7 @@ have been used in a number of papers from our group including:
[](http://dx.doi.org/10.1016/j.cell.2007.01.040)
[](http://dx.doi.org/10.1016/j.cub.2010.07.045)
-[](http://dx.doi.org/10.1038/nature10428)
+[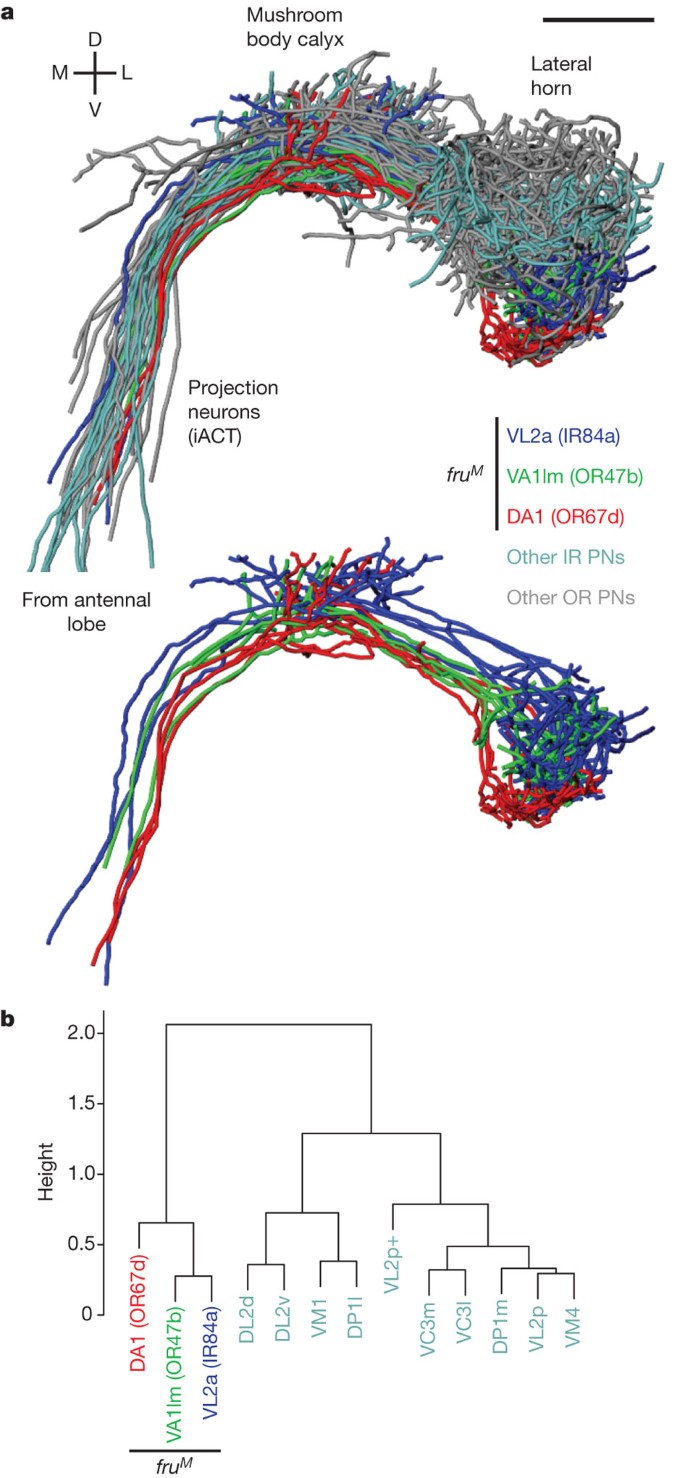 ](http://dx.doi.org/10.1038/nature10428)
[
](http://dx.doi.org/10.1038/nature10428)
[ ](http://dx.doi.org/10.1016/j.cell.2013.11.025)
[](http://dx.doi.org/10.1016/j.neuron.2016.06.012)
diff --git a/docs/index.html b/docs/index.html
index dc3c93e4..095dafac 100644
--- a/docs/index.html
+++ b/docs/index.html
@@ -122,12 +122,11 @@
](http://dx.doi.org/10.1016/j.cell.2013.11.025)
[](http://dx.doi.org/10.1016/j.neuron.2016.06.012)
diff --git a/docs/index.html b/docs/index.html
index dc3c93e4..095dafac 100644
--- a/docs/index.html
+++ b/docs/index.html
@@ -122,12 +122,11 @@