-
Notifications
You must be signed in to change notification settings - Fork 370
sparse_matmul implementation, softmax stability, layer configs #155
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Conversation
|
@julienvalentin @amakadia this supercedes PR 101. |
|
Thanks for making this change. I'm testing this now, and should be able to respond soon. |
|
Hi Jack, |
|
Hi Jack, Could you also test this change on the mesh-segmentation model shared in the mesh-segmentation-demo.ipynb colab? We want to verify consistency with the existing implementation as a baseline. |
|
@avneesh-sud sorry about the delay on this - ECCV deadline is taking priority right now. |
@jackd - hope you are safe and recovered from ECCV deadline. Checking in if you have bandwidth to benchmark the changes and test with the segmentation model now? |
|
@avneesh-sud - sorry about the delay, rolled straight into a new role after ECCV. I've had another look and... things aren't as straight forward as I remember. I'm running experiments on my laptop now with a 1050-Ti (previous results were on a desktop / 1070), but having done some more digging on experiments with full forward/backwards pass and sorted inputs, I'm observing:
Standard implementation OOMs with default I've never really trusted the memory usage values that come out of jitted code, and this experiment hasn't given me any more faith. I don't know of any other profiling tools for this - happy to take suggestions though. Will hopefully have a chance to play with the model tomorrow :) |
|
For the models from the notebook, the sparse implementation (proposed in this PR) takes ~170ms per step, vs 215ms for the original implementation on my laptop with 1050Ti. Not much observed difference in memory usage based on random polling of In terms of model accuracy, there are tests in place that show the results are the same but I wanted to do up a keras model anyway so I did up this gist. I got a bit carried away changing stuff and it's entirely possible I stuffed something up, but overall loss numbers look roughly equivalent to the notebook file. Unsurprisingly, the results using different layer implementations are very similar. Somewhat surprising was that my sparse keras implementation in tf 2.1 was ~10% slower than the notebook version and tf 1.15, where as my keras implementation using the default layer implementations ~30% slower than the notebook equivalent. Again, not really an apples-to-apples comparison because I changed a fair bit (though I'd hoped my changes would have made things faster if anything). Maybe I screwed up something... |
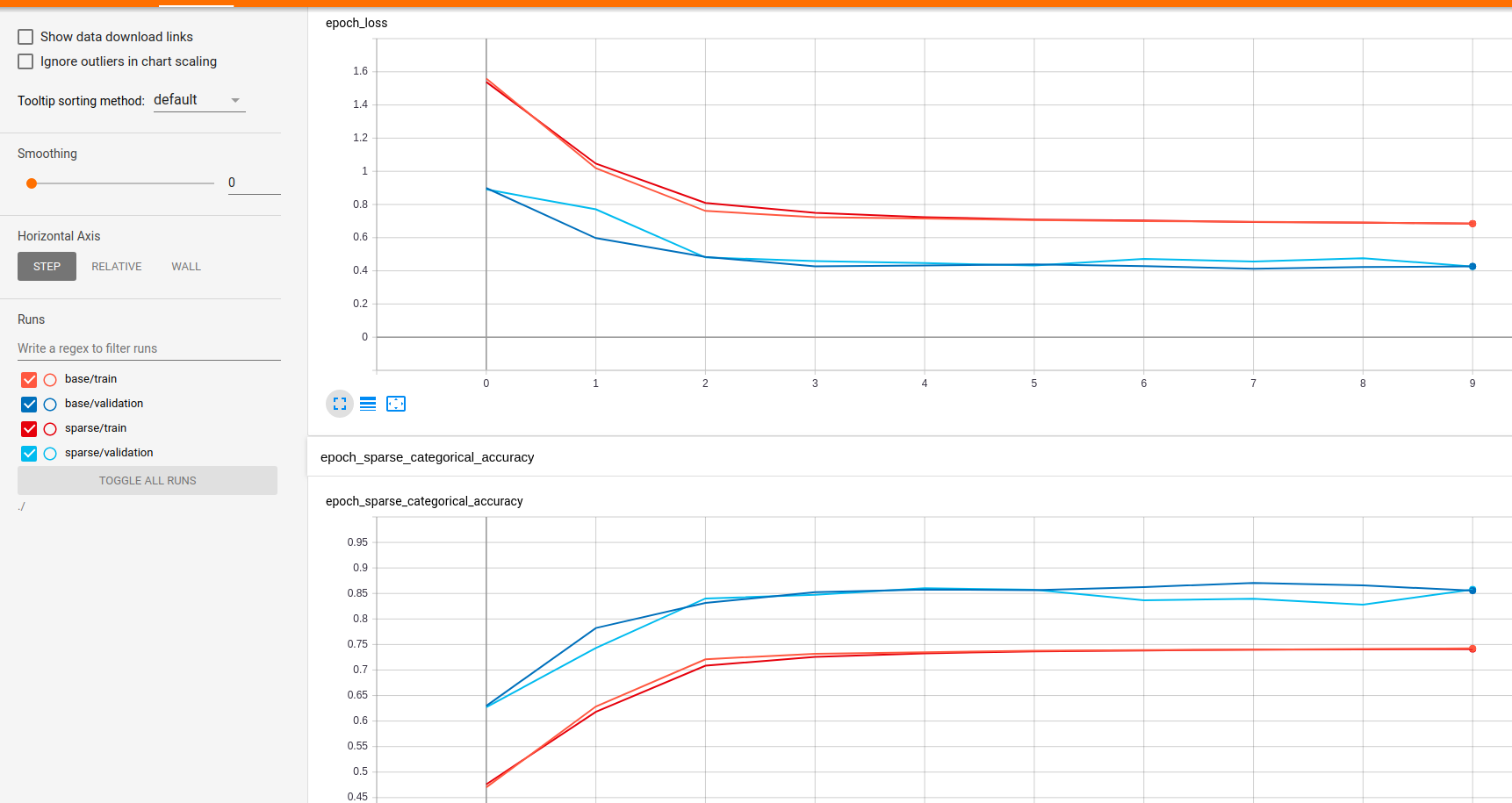
This PR is mostly focused on adding a
'sparse_matmul'implementation offeature_steered_convolutionwhich is faster than the base version ('gather_sum') in certain circumstances.Differences can be seen in the benchmarking script.
Benefits are similar though reduced when using
--jitoption.Note these benefits are not across-the-board, hence the choice of implementation is left to the user. For example, smaller networks are faster using
gather_sum. Verify with--num_layers=2or similar.Other minor changes:
tf.nn.softmaxwhen using JIT)feature_steered_convolutioncan take non-square neighborhood tensors. Updated documentation to reflect this.get_configimplementations andtf.keras.*.getcalls in constructors, allowingLayer.from_configto work as intended.