-
Notifications
You must be signed in to change notification settings - Fork 9
/
index.Rmd
76 lines (52 loc) · 2.76 KB
/
index.Rmd
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
---
title: "Home"
site: workflowr::wflow_site
output:
workflowr::wflow_html:
toc: false
---
# kallisto | bustools R notebooks
This repository has example notebooks that demonstrate usage of [`kallisto bus`](https://www.biorxiv.org/content/10.1101/673285v1) to generate gene count or transcript compatibility count (TCC) matrices from FASTQ files and some basic downstream analyses. To run the notebooks, please install [kallisto](https://pachterlab.github.io/kallisto/starting), [bustools](https://github.com/BUStools/bustools), and the [BUSpaRse](https://github.com/BUStools/BUSpaRse) R package.
## Notebooks
<div class = "row">
<div class = "col-md-6">
<center>
[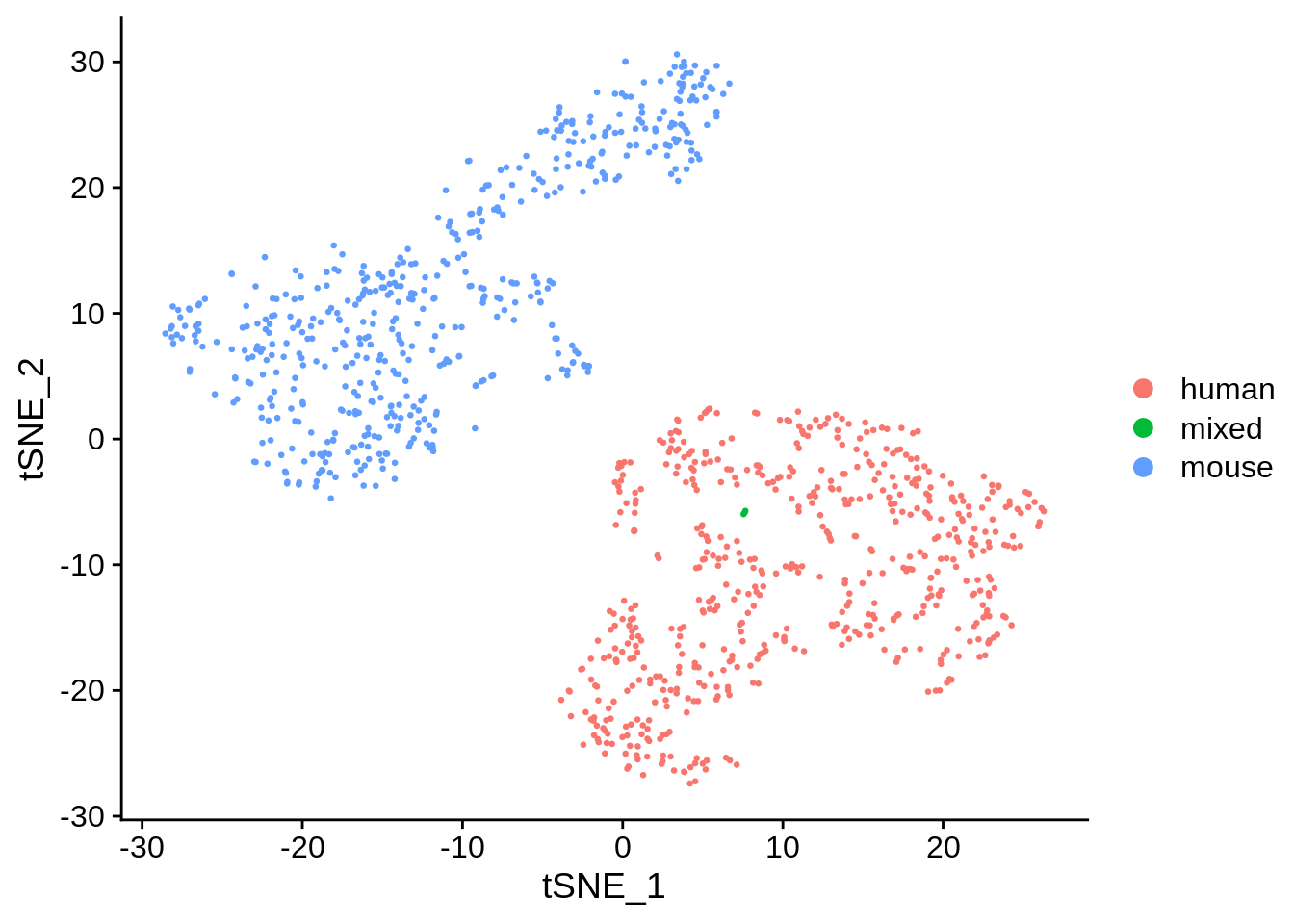{width=600px}](https://bustools.github.io/BUS_notebooks_R/10xv2.html)
[1k 1:1 Mixture of Fresh Frozen Human (HEK293T) and Mouse (NIH3T3) Cells (10x v2 chemistry); demonstration of `bustools`](https://bustools.github.io/BUS_notebooks_R/10xv2.html)
</center>
</div>
<div class = "col-md-6">
<center>
[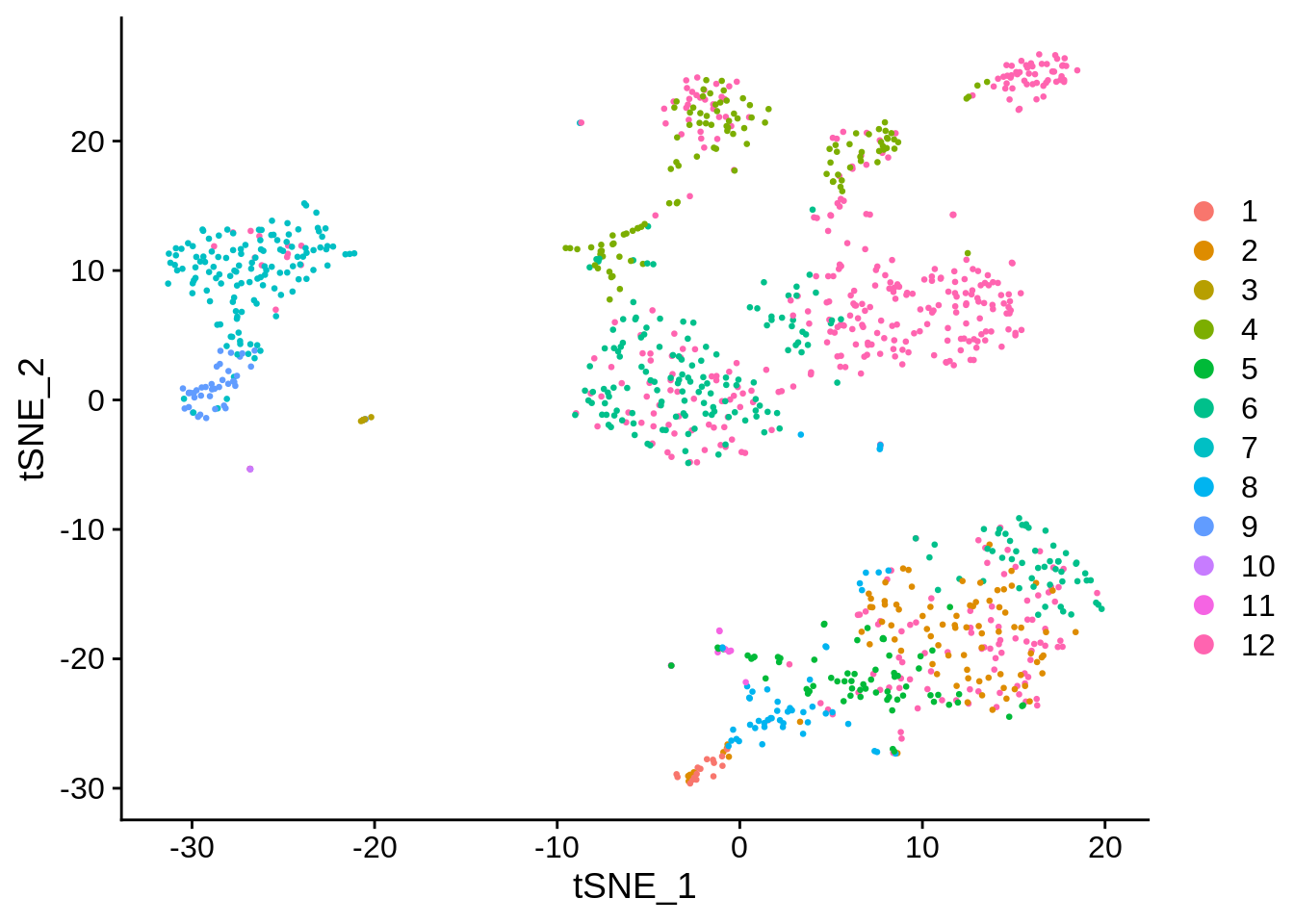{width=600px}](https://bustools.github.io/BUS_notebooks_R/10xv3.html)
[1k PBMCs from a Healthy Donor (10x v3 chemistry); demonstration of `TENxBUSData` and `BUSpaRse`](https://bustools.github.io/BUS_notebooks_R/10xv3.html)
</center>
</div>
</div>
<div class = "row">
<div class = "col-md-6">
<center>
[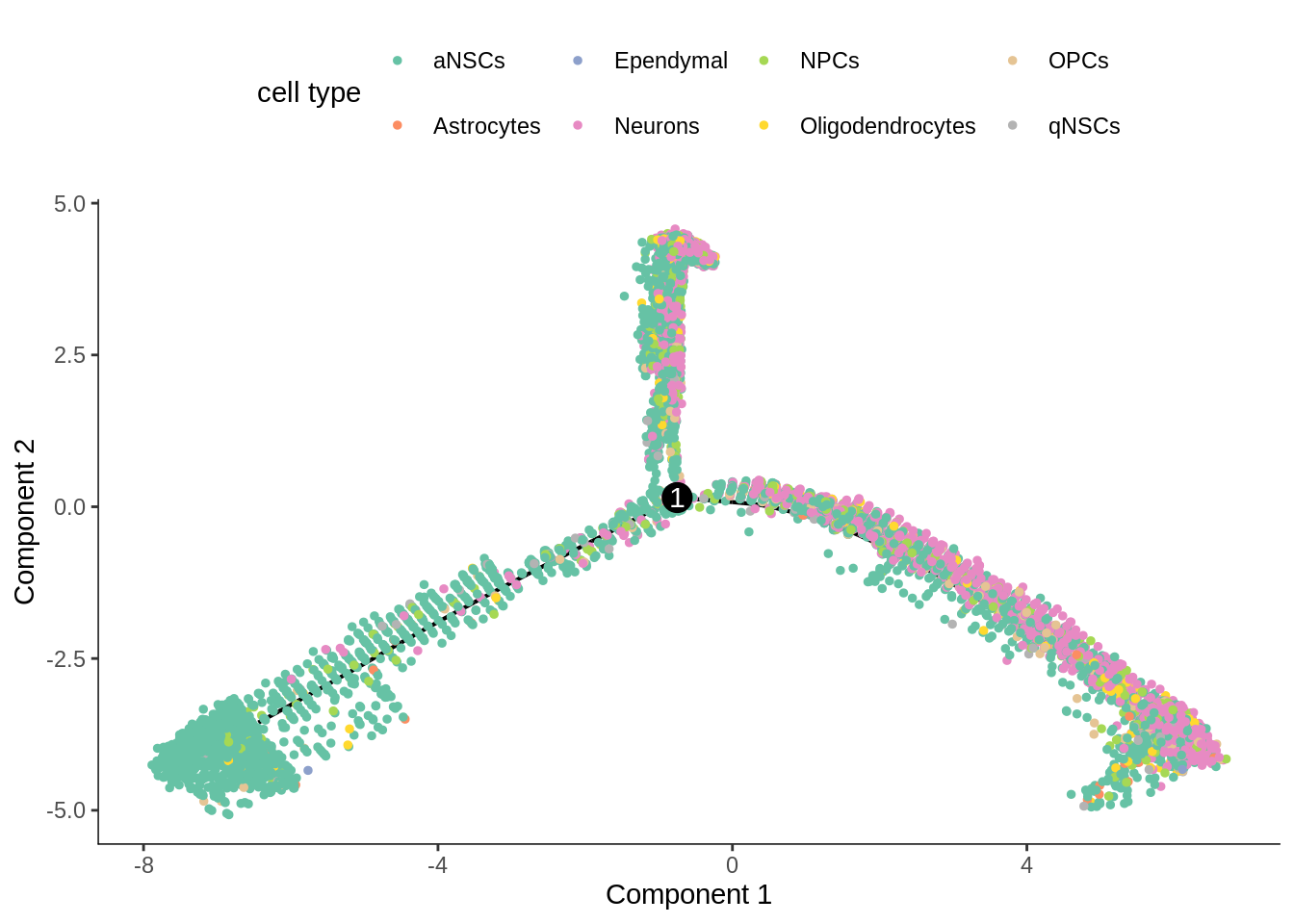{width=600px}](https://bustools.github.io/BUS_notebooks_R/monocle2.html)
[Monocle 2 pseudotime analysis with 10k E18 mouse neurons from 10x](https://bustools.github.io/BUS_notebooks_R/monocle2.html)
</center>
</div>
<div class = "col-md-6">
<center>
[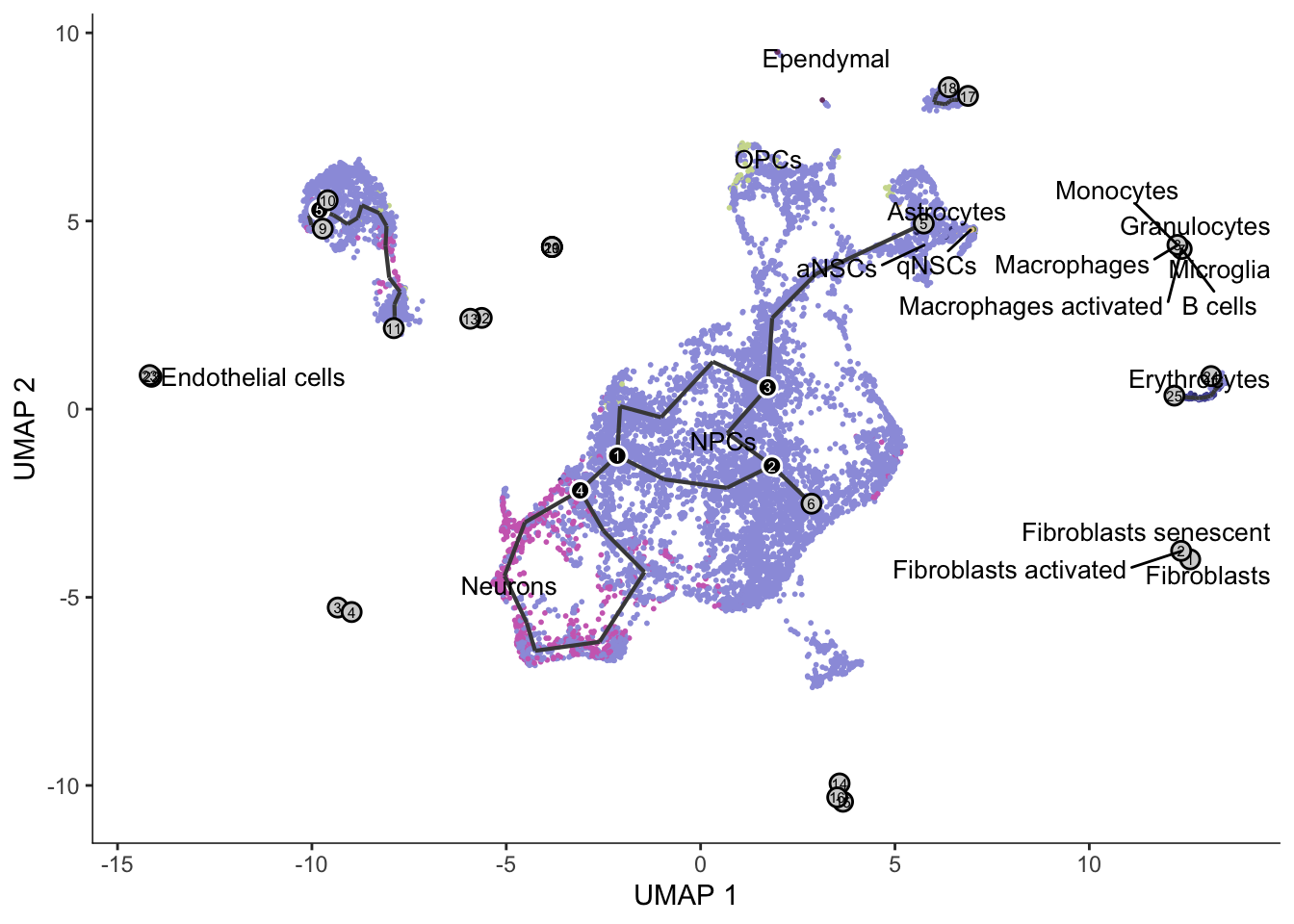{width=600px}](https://bustools.github.io/BUS_notebooks_R/monocle3.html)
[Monocle 3 pseudotime analysis with 10k E18 mouse neurons from 10x](https://bustools.github.io/BUS_notebooks_R/monocle3.html)
</center>
</div>
</div>
<div class = "row">
<div class = "col-md-6">
<center>
[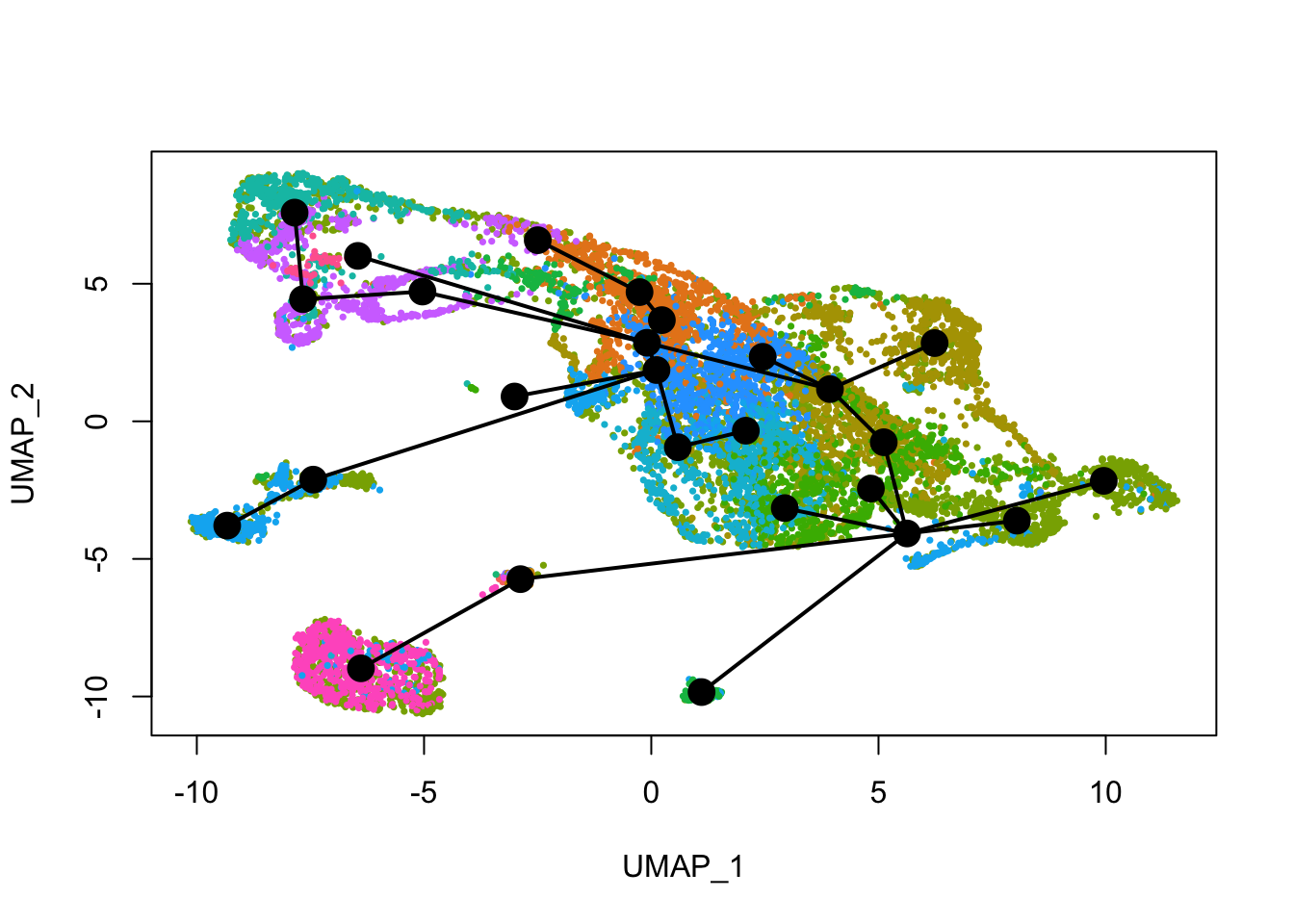{width=600px}](https://bustools.github.io/BUS_notebooks_R/slingshot.html)
[Slingshot pseudotime analysis with 10k E18 mouse neurons from 10x](https://bustools.github.io/BUS_notebooks_R/slingshot.html)
</center>
</div>
<div class = "col-md-6">
[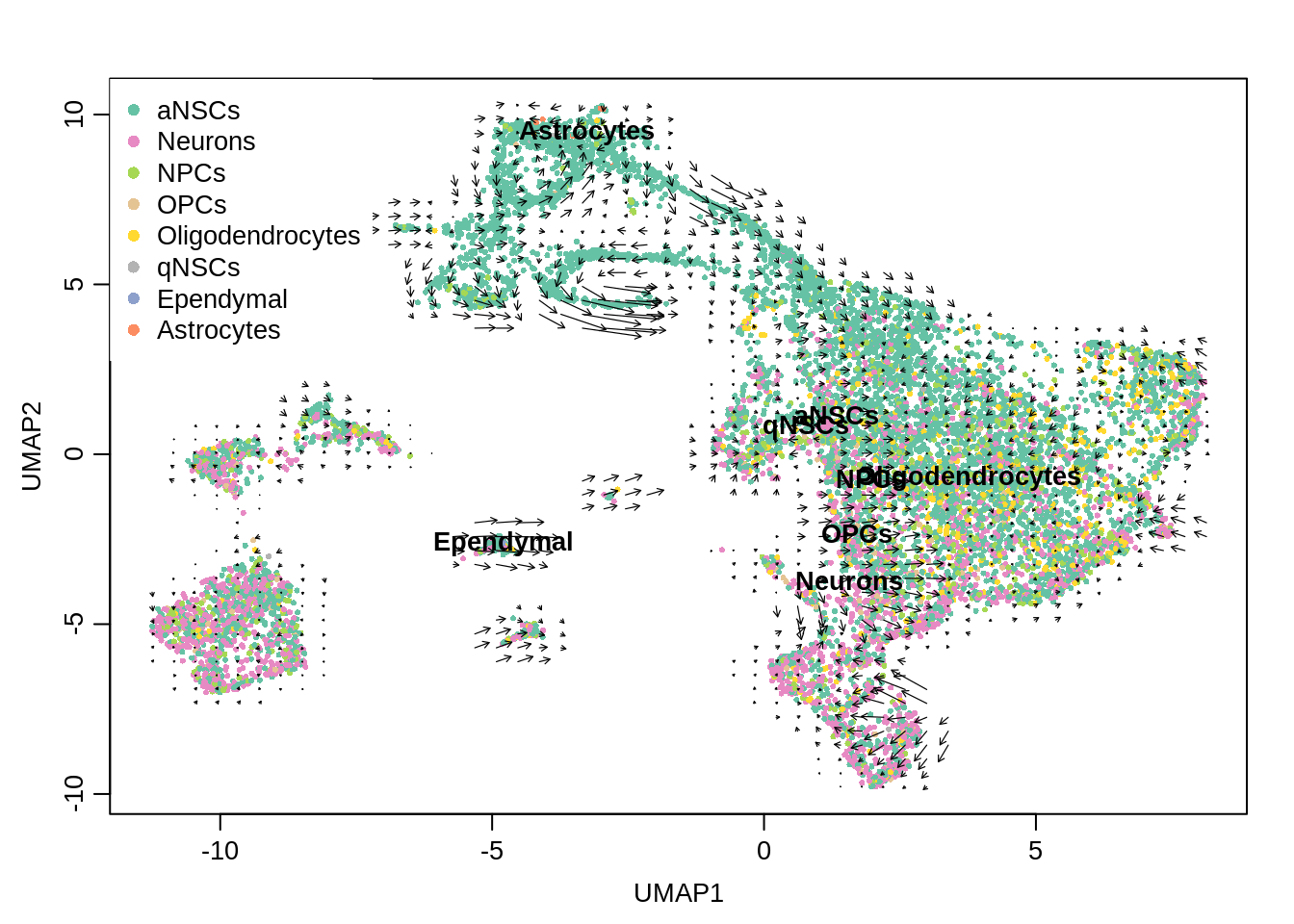{width=600px}](https://bustools.github.io/BUS_notebooks_R/velocity.html)
[RNA velocity analysis with 10k E18 mouse neurons from 10x](https://bustools.github.io/BUS_notebooks_R/velocity.html)
</div>
</div>
-----------
Differential expression with equivalence classes in TCC matrix (coming soon)