New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Unknown SSL protocol error in connections with RCurl under Windows #208
Comments
|
I suggest to replace the RCurl routines with
Attaching package: ‘httr’ The following object is masked from ‘package:curl’:
[1] "InChIKey=LFTLOKWAGJYHHR-UHFFFAOYSA-N" |
|
@tsufz thanks for the work, I have provisionally added the workaround (slightly adapted) in 0c82f3b. It is building successfully on Windows. I am waiting for a successful Bioc build in order to close the issue. RCurl should probably be removed completely at this point; up to now it's only a minimal workaround. |
|
Building on Bioconductor. |
|
It might be building on BioC but I am only getting NAs for all our test data and everything else? |
|
OK our bad ... fresh install of BioC for Loulou didn't pull the latest RMB for some reason ... httr cactus is working... |
|
Hi, remember that BioC/RMB package versions are tied to the R version. |
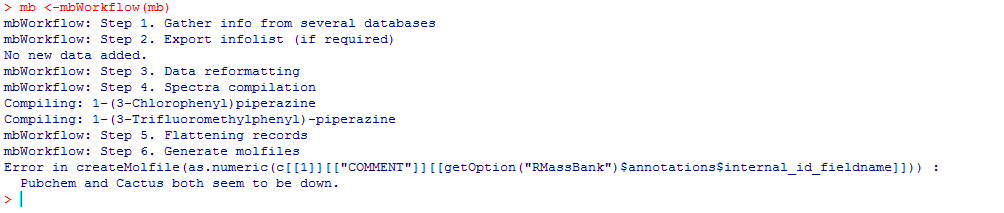
RCurl derives an unknown SSL protocol error during queries with Cactus
library(RCurl)Loading required package: bitops
getURLContent("https://cactus.nci.nih.gov/chemical/structure/7529-22-8/stdinchikey")error in function (type, msg, asError = TRUE) :
Unknown SSL protocol error in connection to cactus.nci.nih.gov:443
sessionInfo()R version 3.5.1 (2018-07-02)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows >= 8 x64 (build 9200)
Matrix products: default
locale:
[1] LC_COLLATE=English_United Kingdom.1252 LC_CTYPE=English_United Kingdom.1252 LC_MONETARY=English_United Kingdom.1252
[4] LC_NUMERIC=C LC_TIME=English_United Kingdom.1252
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] RCurl_1.95-4.11 bitops_1.0-6 httr_1.3.1 curl_3.2 BiocInstaller_1.30.0
loaded via a namespace (and not attached):
[1] compiler_3.5.1 R6_2.3.0 tools_3.5.1 yaml_2.2.0
The text was updated successfully, but these errors were encountered: