Dataset and source code for paper "Enhancing Keyphrase Extraction from Academic Articles with their Reference Information".
The research content of this project is to analyze the impact of the introduction of reference title in scientific literature on the effect of keyword extraction. This project uses three datasets: SemEval-2010, PubMed and LIS-2000, which are located in the dataset folder. At the same time, we use two unsupervised methods: TF-IDF and TextRank, and three supervised learning methods: Naïve Bayes, CRF and BiLSTM-CRF. The first four are traditional keywords extraction methods, located in the folder ML, and the last one is deep learning method, located in the folder DL.
Keyphrase_Extraction Root directory ├─dl.bat Batch commands to run deep learning model ├─ml.bat Batch commands to run traditional models │ ├─Dataset Experimental datasets │ ├─SemEval-2010 Contains 244 scientific papers │ ├─PubMed Contains 1, 316 scientific papers │ └─LIS-2000 Contains 2, 000 scientific papers │ ├─DL Module of the deep learning model │ ├─build_path.py Create file paths for saving preprocessed data │ ├─crf.py Source code of CRF algorithm implementation (Use pytorch framework) │ ├─main.py The main function of running the program │ ├─model.py Source code of BiLSTM-CRF model │ ├─preprocess.py Source code of preprocessing function │ ├─textrank.py Source code of TextRank algorithm implementation. │ ├─tf_idf.py Source code of TF-IDF algorithm implementation. │ ├─utils.py Some auxiliary functions │ ├─models Parameter configuration of deep learning models │ └─datas │ └─tags Label settings for sequence labeling │ ├─ML Module of the traditional models │ ├─build_path.py Create file paths for saving preprocessed data │ ├─configs.py Path configuration file │ ├─crf.py Source code of CRF algorithm implementation(Use CRF++ Toolkit) │ ├─evaluate.py Surce code for result evaluation │ ├─naivebayes.py Source code of Naïve Bayes algorithm implementation(Use KEA-3.0 Toolkit) │ ├─preprocessing.py Source code of preprocessing function │ ├─textrank.py Source code of TextRank algorithm implementation │ ├─tf_idf.py Source code of TF-IDF algorithm implementation │ ├─utils.py Some auxiliary functions │ ├─CRF++ CRF++ Toolkit │ └─KEA-3.0 KEA-3.0 Toolkit │ └─README.md
The dataset includes the following three json files:
Each line of the json file includes:
In order to facilitate the reproduction of the experimental results, the project uses bat batch command to run the program uniformly (only in Windows Environment). The dl.bat file is the batch command to run the deep learning model, and the ml.bat file is the batch command to run the traditional algorithm.
In the Windows environment, use the key combination Win + R and enter cmd to open the DOS command box, and switch to the project's root directory (Keyphrase_Extraction). Then input dl.bat, that is, run deep learning model to get the result of keyword extraction; Enter ml.bat to run traditional algorithm to get keywords Extract the results.
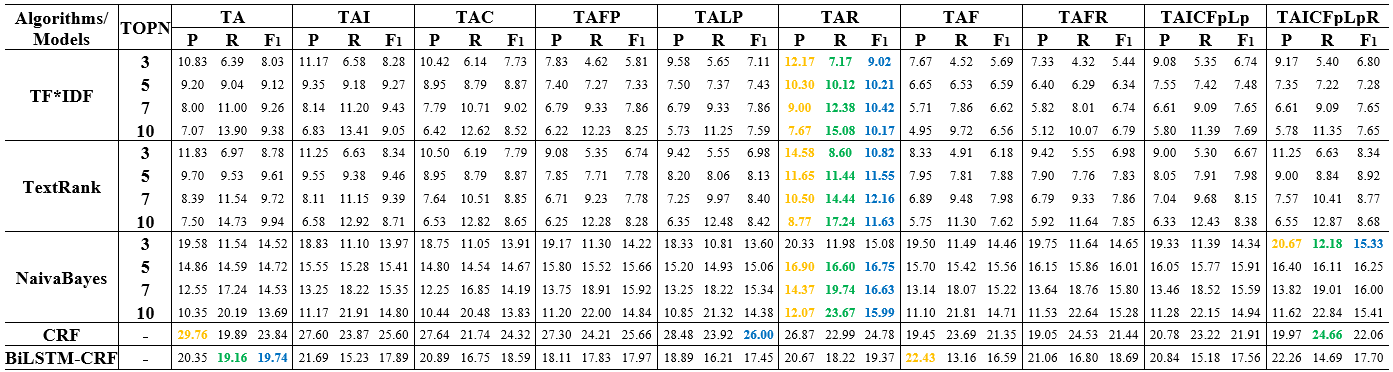
The following figures show that the influence of reference information on keyphrase extraction results of TF*IDF, TextRank, NB, CRF and BiLSTM-CRF.
Table 1: Keyphrase extraction performance of multiple corpora constructed using different logical structure texts on the dataset of SemEval-2010

Table 2: Keyphrase extraction performance of multiple corpora constructed using different logical structure texts on the dataset of PubMed
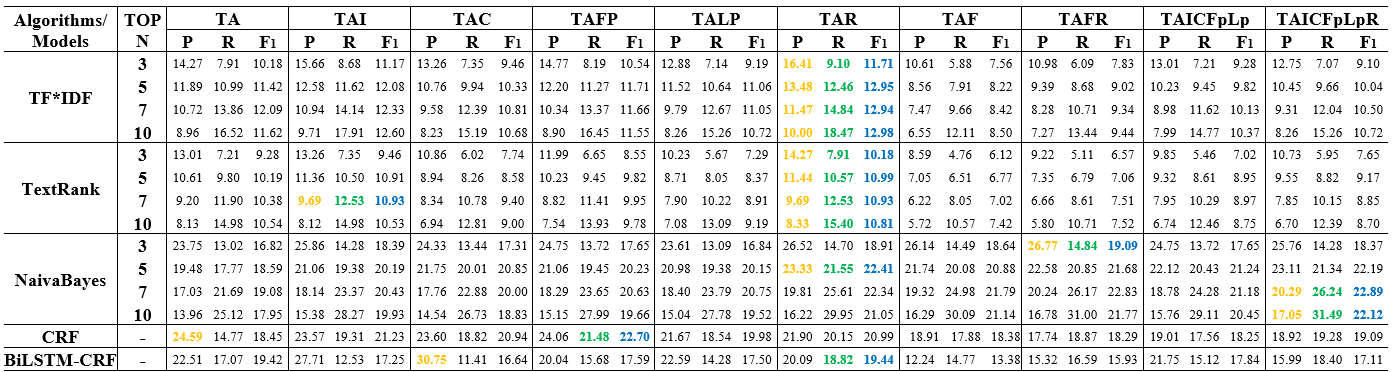
Table 3: Keyphrase extraction performance of multiple corpora constructed using different logical structure texts on the dataset of LIS-2000
Note: The yellow, green and blue bold fonts in the table represent the largest of the P, R and F1 value obtained from different corpora using the same model, respectively.
Before running this project, check that the following Python packages are included in your runtime environment.
Please cite the following paper if you use this code and dataset in your work.
Chengzhi Zhang, Lei Zhao, Mengyuan Zhao, Yingyi Zhang. Enhancing Keyphrase Extraction from Academic Articles with their Reference Information. Scientometrics, 2022, 127(2): 703–731. [doi] [arXiv]