New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Same prediction for every molecules with pretrained and finetuned model. #21
Comments
|
Hello, Best regards, |
|
Hello :) could you provide the rest of your code, please? Could you also explain how you run your code? Line by line from command-line, or with Jupyter notebook? Unfortunately, I am unable to deduce it from your screen :) |
|
Hello! thank you very much for the reply.
I expected featurizer creates different results but always same as you can see here. So I assume that problems come from 'featurizer = MatFeaturizer.from_pretrained('mat_masking_20M')'
|
|
Hmm, it's quite strange. Could you show me your conda environment, please? Moreover, could you try to reproduce this bug from a freshly cloned huggingmolecules repository? Cause I see you've modified the Btw: |
|
Yep, unfortunately, I cannot reproduce the bug :< I've tried on three different machines, but everything works. But looking at the above screenshots, I'm not sure if you tried to reinstall huggingmolecules once again, after installing pytorch 1.7.0. Could you please run Btw: in branch |
|
Hello, I created a new conda environment and started from the beginning. |
|
I have installed huggingmolecules and the additional requirements in the new conda environment, and everything works fine. |
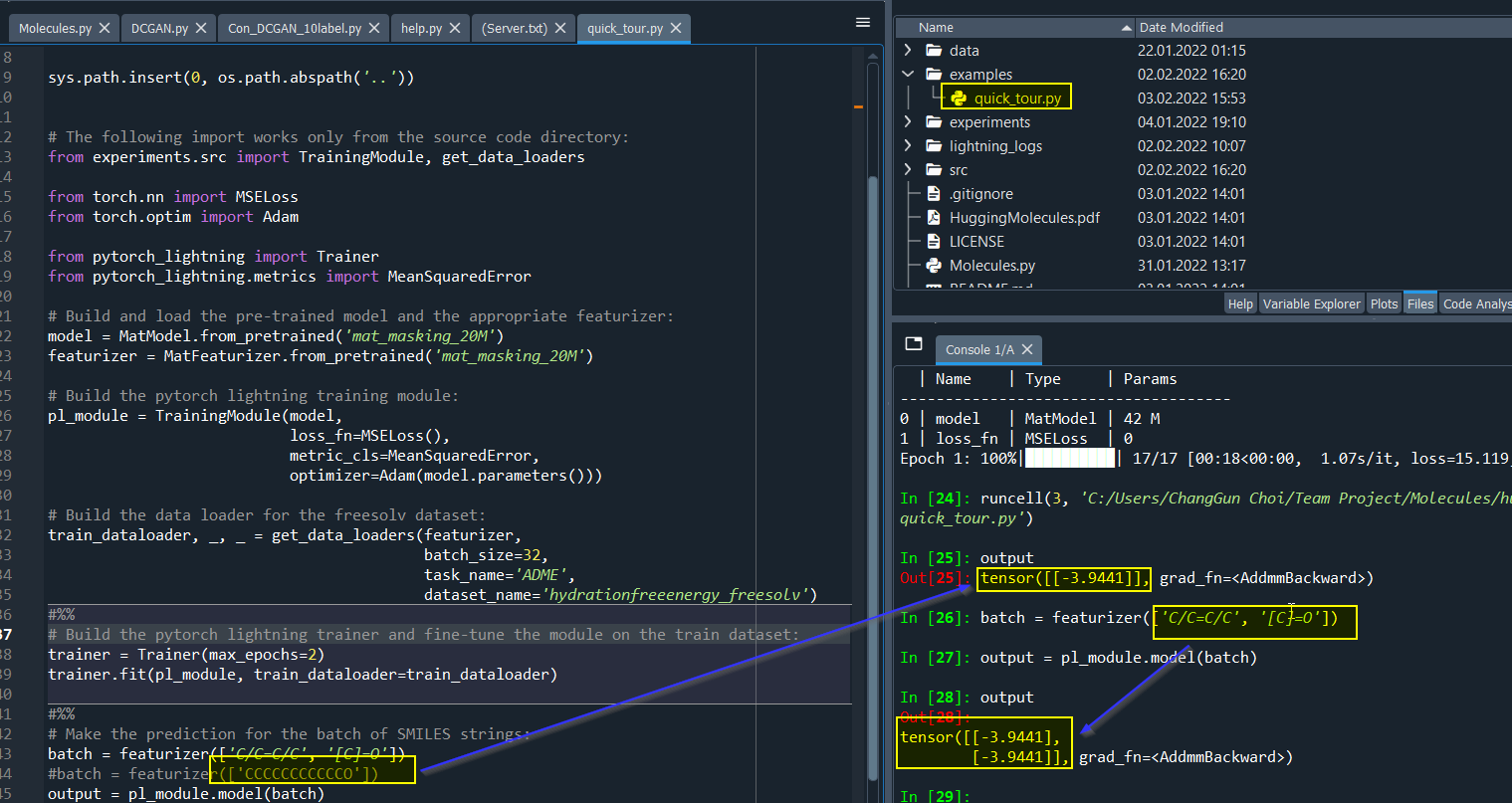
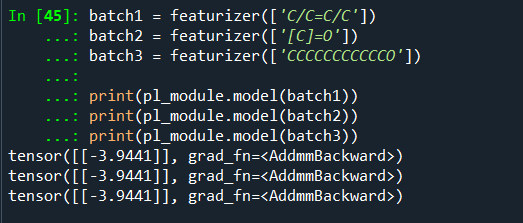
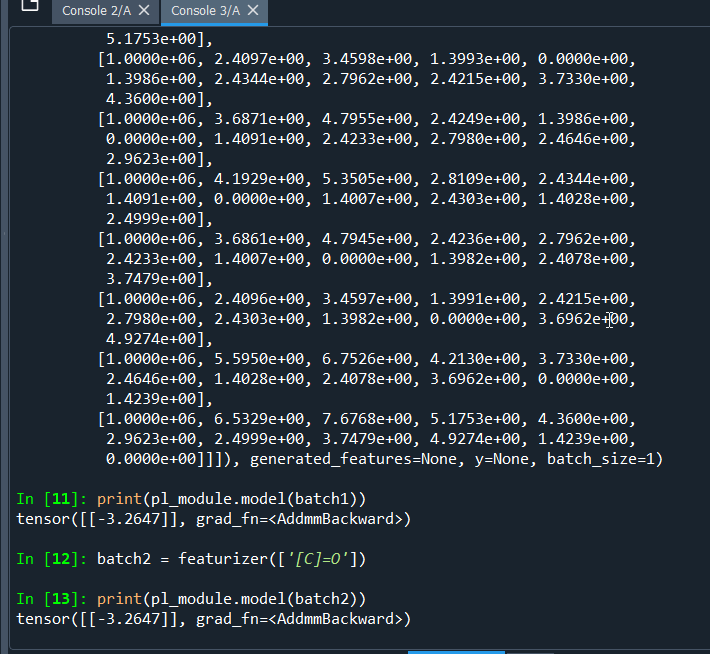


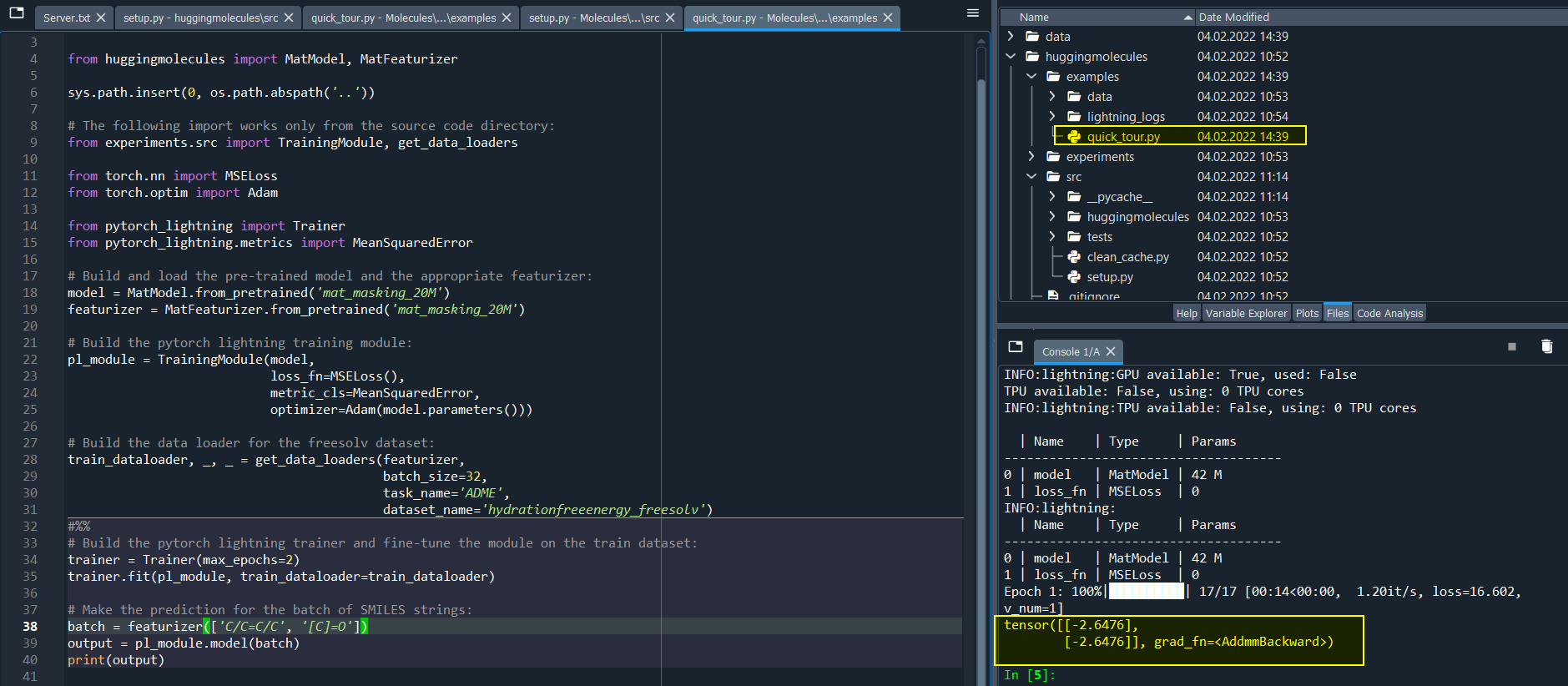
Hello,
By using quick_tour.py, results are always same for different molecules. I am confused about it.
The result is also same for the fine-tuned model.
The text was updated successfully, but these errors were encountered: