SBGNViz.as is a web application based on Cytoscape Web to visualize the pathway models represented by SBGN Process Description Notation. SBGNViz.as accepts the pathway models represented in SBGN-ML format.
SBGNViz.as was built by extending the web based data visualization tool CytoscapeWeb
to support the SBGN Process Description Notation. SBGNViz.as was
developed using Adobe Flash Builder and
[Flex]http://www.adobe.com/products/flex.html technologies.
A sample application using SBGNViz.as can be found here.
SBGNViz.as is distributed under GNU Lesser General Public License. Instructions for obtaining a working copy of the project can be found in README.txt and README_SBGNViz.txt files. SBGNViz.as works on every environment supporting Adobe Flash.
Below are some sample screenshots from SBGNViz.as, illustrating some of its capabilities. Please click on a figure to see it in full size. A User's Guide can be found here, documenting all features of the tool.
<td align="center">
<a href="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/layout_properties.png">
<img src="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/layout_properties_s.png"/>
</a>
<p>Parameters to configure automatic layout</p>
</td>
</tr>
<tr>
<td align="center">
<a href="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/before_layout.png">
<img src="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/before_layout_s.png"/>
</a>
<p>Sample pathway <em>before</em> automatic layout is applied</p>
</td>
<td align="center">
<a href="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/after_layout.png">
<img src="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/after_layout_s.png"/>
</a>
<p>Sample pathway <em>after</em> automatic layout is applied</p>
</td>
</tr>
<tr>
<td align="center">
<a href="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/highlight_processes.png">
<img src="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/highlight_processes_s.png"/>
</a>
<p>Highlighting all processes a selected node is involved in</p>
</td>
<td align="center">
<a href="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/inspector.png">
<img src="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/inspector_s.png"/>
</a>
<p>Inspecting detailed properties of a macromolecule</p>
</td>
</tr>
<tr>
<td align="center">
<a href="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/save.png">
<img src="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/save_s.png"/>
</a>
<p>Saving current process description diagram as PDF</p>
</td>
<td align="center">
<a href="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/load.png">
<img src="http://cs.bilkent.edu.tr/~ivis/sbgnviz-as/samples/load_s.png"/>
</a>
<p>Loading a process description diagram from an SBGN-ML file</p>
</td>
</tr>
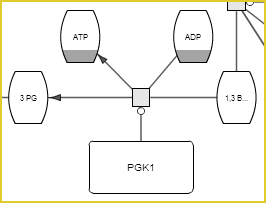
Glycolysis pathway visualized in SBGNViz.as |
- Istemi Bahceci, Ugur Dogrusoz, Mecit Sari, i-Vis at Bilkent University
- S.Onur Sumer, cBio at MSKCC