-
Notifications
You must be signed in to change notification settings - Fork 1
Estimate the acceptable minimum embedding dimension
Estimate the acceptable minimum embedding dimension.
Estimate the acceptable minimum embedding dimension using False Nearest Neighbors Algorithm [Kennel_1992].
Select appropriately from the embedded data with dimension
, find the closest tangent vector
, and set the distance to
. It is embedded from 1 to
D_max.
Calculate the closest distance for all data. The points that satisfy the following two conditions are counted. (
threshold_A) and (
threshold_R) are thresholds.
is the standard deviation obtained from the time series.
Let the acceptable minimum embedding dimension be the first dimension below 1% (threshold_percent) in the percentages obtained for each embedded dimension.
By default, the actual calculation result is displayed. The point actually adopted is red. If you want to turn off the display, give plot=False.
It is necessary to set the threshold appropriately, and it is difficult to adapt to unknown data. Different value of and
may lead to different result (
dimension). AFF Algorithm (est_dimension_w_afn) is devised.
from hundun.exploration import est_dimension_w_fnn- u_seq
- T
- D_max=10
- threshold_R=10
- threshold_A=2
- threshold_percent=1
- plot=True
- path_save_plot=None
- dimension
- percentages:
numpy.ndarray
Estimate the acceptable minimum embedding dimension for x in the Lorenz time series.
from hundun.equations import Lorenz
from hundun.exploration import est_dimension_w_fnn
x_seq = Lorenz.get_u_seq(25000)[::5, 0]
D, percentages = est_dimension_w_fnn(x_seq, T=10)
print(D)3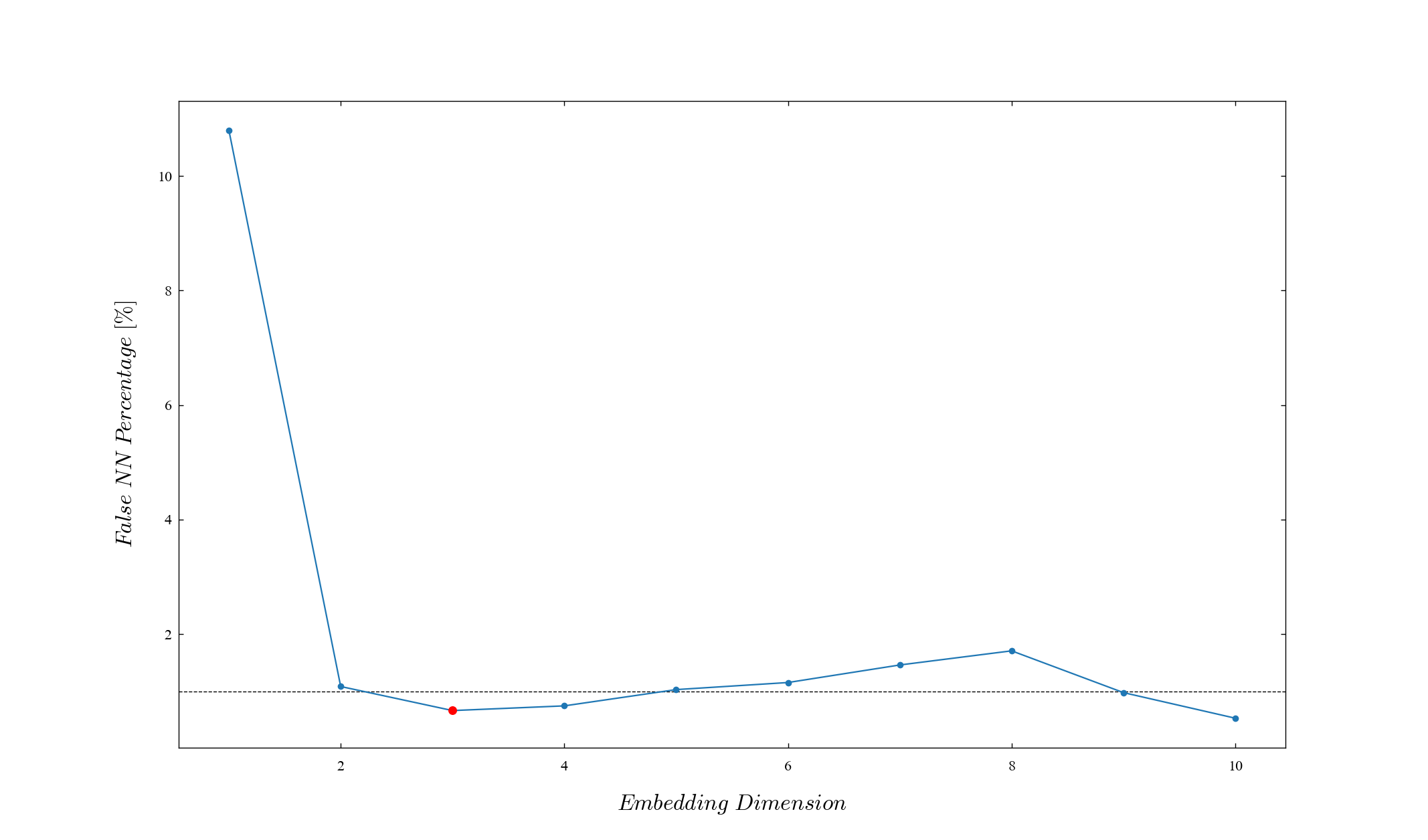
(1992) Matthew B. Kennel, Reggie Brown, and Henry D. I. Abarbanel
DOI: 10.1103/PhysRevA.45.3403
(1993) Abarbanel, Henry D. I. and Brown, Reggie and Sidorowich, John J. and Tsimring, Lev Sh
DOI: 10.1103/RevModPhys.65.1331
Estimate the acceptable minimum embedding dimension using Averaged False Neighbors Algorithm [Cao_1997].
Calculate the distance in the same way as FNN. However, it uses maximum norm instead of Euclidean distance.
Estimate the acceptable minimum embedding dimensions with reference to E1 and E2 obtained by calculation.
The first value at which E1 and E2 each exceed the threshold (threshold_E1=0.9, threshold_E2=1) is retrieved as the dimension.
from hundun.exploration import est_dimension_w_afn- u_seq
- T
- D_max=10
- threshold_E1=0.9
- threshold_E2=1
- plot=True
- path_save_plot=None
- dimension_list:
List[int] - E_list:
List[numpy.ndarray]
from hundun.equations import Lorenz
from hundun.exploration import est_dimension_w_afn
import numpy as np
x_seq = Lorenz.get_u_seq(25000)[::5, 0]
D_list, E_list = est_dimension_w_afn(x_seq, T=1)
print(D_list)[4, 3]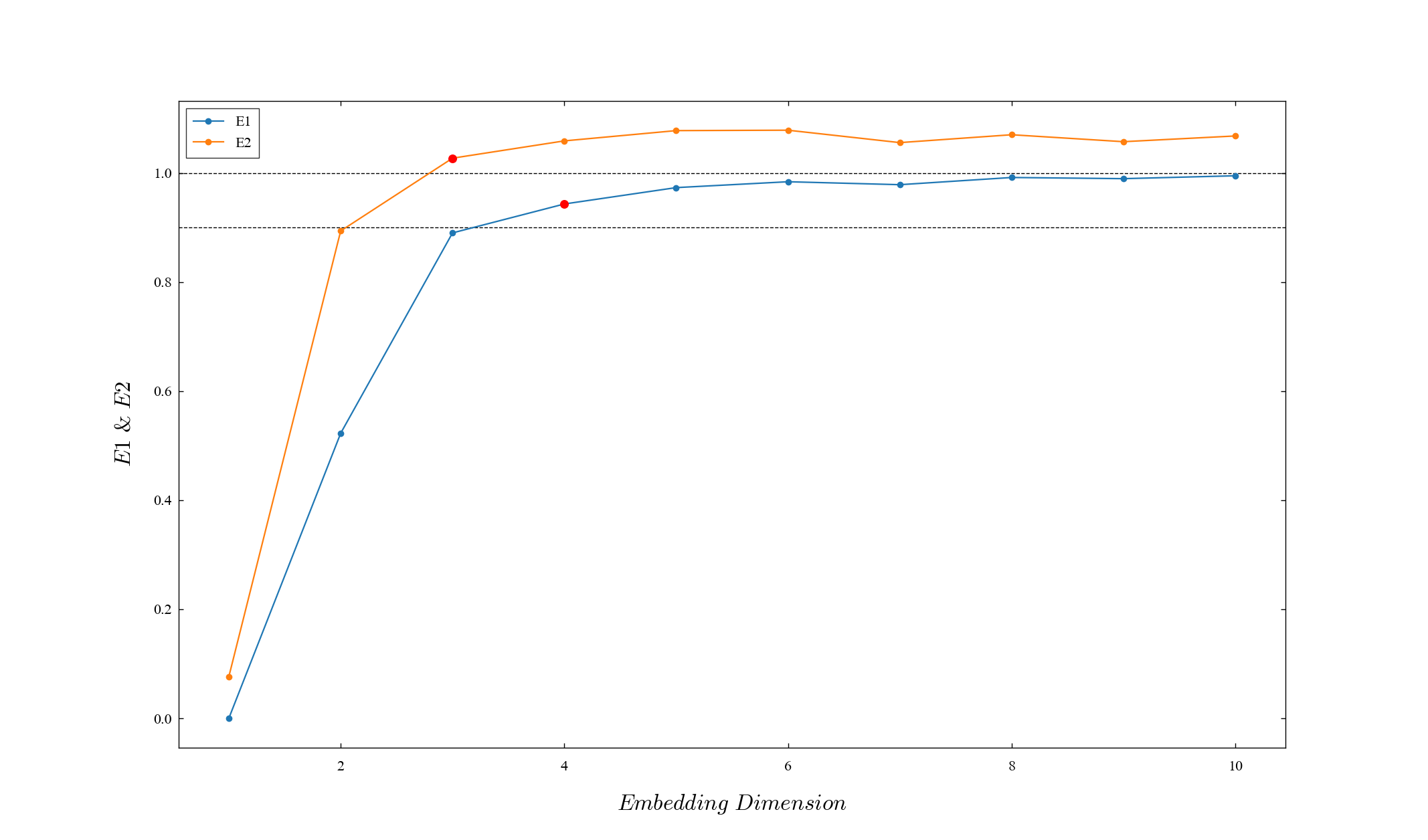
(1997) Liangyue Cao
DOI: 10.1016/S0167-2789(97)00118-8
Find (
K) closest tangent vectors for at a given time
.
These vectors are represented as
.
Using the vector after the
(
tau) step elapses, is defined as follows.
Calculate the variance of with the following equation.
Randomly calculate from
M=51 to find the median.
Do this
Q=10 times.
is calculated in each embedded dimension and the minimum value is adopted as the dimension.
from hundun.exploration import est_dimension_w_afn- u_seq
- T
- D_max=10
- tau=5
- K=4
- Q=10
- M=51
- reshape=True
- plot=True
- path_save_plot=None
- D
- median_e_trans_ave
from hundun.equations import Lorenz
from hundun.exploration import est_dimension_w_wayland
x_seq = Lorenz.get_u_seq(25000)[::5, 0]
D, median_e_trans_ave = est_dimension_w_wayland(x_seq, T=1)
print(D)3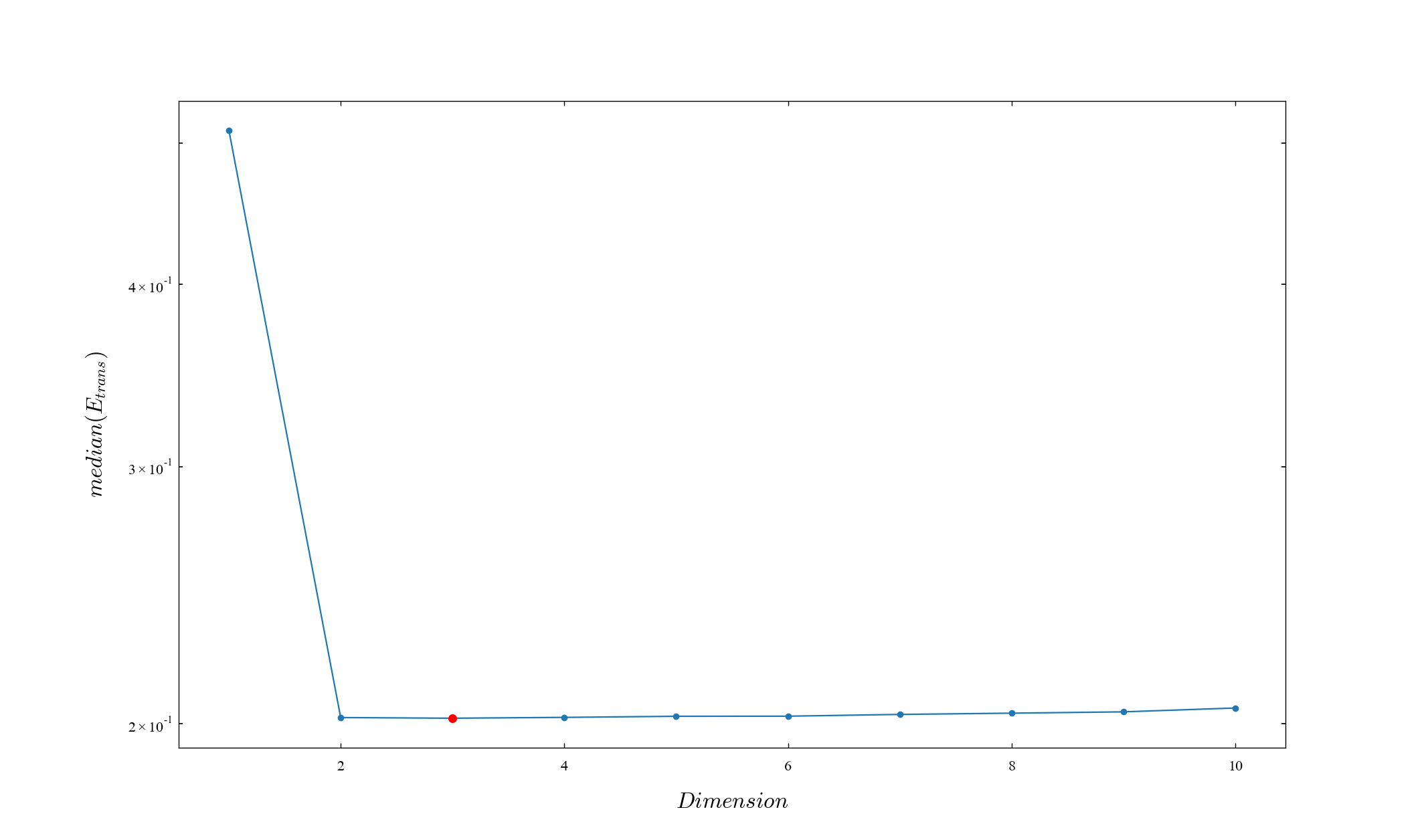
(1993) Wayland, Richard and Bromley, David and Pickett, Douglas and Passamante, Anthony
DOI: 10.1103/PhysRevLett.70.580
It will not be available after version 0.2. Use est_dimension_w_fnn instead.
It will not be available after version 0.2. Use est_dimension_w_afn instead.
