You signed in with another tab or window. Reload to refresh your session.You signed out in another tab or window. Reload to refresh your session.You switched accounts on another tab or window. Reload to refresh your session.Dismiss alert
I make extensive use of scatter_matrix and I find misalignments in the coordinates of the scatter plots (off-diagonal) and the histograms (on-diagonal) which seem undesirable to me.
Consider the picture
).
In this case, there is a clear misalignment (of about .02) in the x-coordinate between the subplot in position 1,2 and the subplot in position 2,2.
In other words, the peeks in subplots 1,2 and 2,2 should occur at the same x location
(at least, IMHO, this would be more meaningful behavior).
The text was updated successfully, but these errors were encountered:
a = - np.linspace(.4,.6,5000) + np.random.randn(5000)
b = np.linspace(.9,1,5000) + .1*np.random.randn(5000)
db = pd.DataFrame(zip(a,b),columns=list('ab'))
pd.scatter_matrix(db)
It seems that the x-coordinate increment is different in subplot 1,1 and subplot 2,1.
The offset (x_11 - x_21) ranges from 0 (at the center of the quadrants) to almost 1 (in this case) at the boundary.
The x-values are hand taken from the status bar.
I make extensive use of
scatter_matrixand I find misalignments in the coordinates of the scatter plots (off-diagonal) and the histograms (on-diagonal) which seem undesirable to me.Consider the picture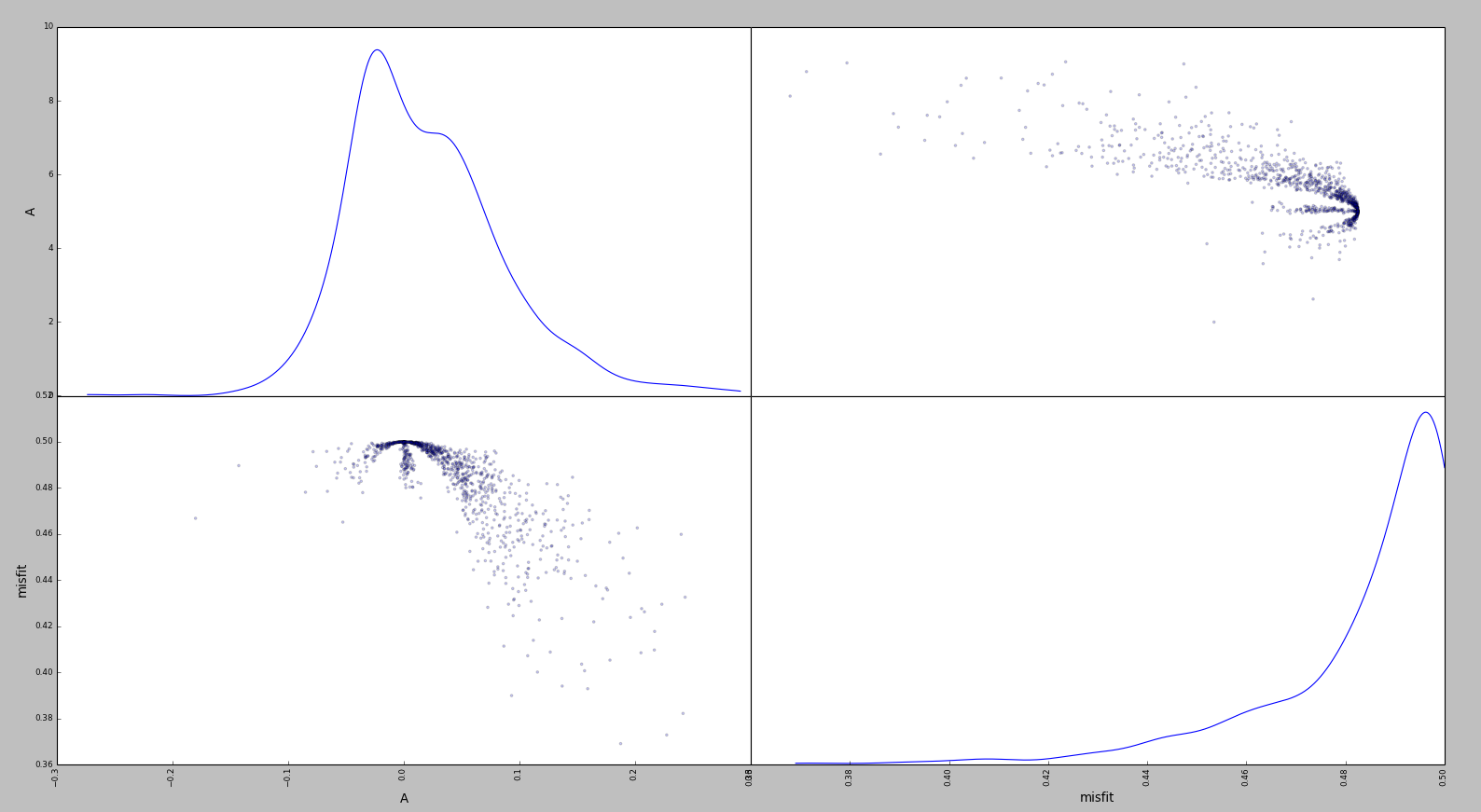
).
In this case, there is a clear misalignment (of about .02) in the x-coordinate between the subplot in position 1,2 and the subplot in position 2,2.
In other words, the peeks in subplots 1,2 and 2,2 should occur at the same x location
(at least, IMHO, this would be more meaningful behavior).
The text was updated successfully, but these errors were encountered: