-
Notifications
You must be signed in to change notification settings - Fork 0
/
enhance-viz.qmd
321 lines (267 loc) · 10.2 KB
/
enhance-viz.qmd
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221
222
223
224
225
226
227
228
229
230
231
232
233
234
235
236
237
238
239
240
241
242
243
244
245
246
247
248
249
250
251
252
253
254
255
256
257
258
259
260
261
262
263
264
265
266
267
268
269
270
271
272
273
274
275
276
277
278
279
280
281
282
283
284
285
286
287
288
289
290
291
292
293
294
295
296
297
298
299
300
301
302
303
304
305
306
307
308
309
310
311
312
313
314
315
316
317
318
319
320
321
# Enhancing Visualizations
```{r}
#| label: libraries1
#| include: false
library(igraph)
library(ggraph)
library(graphlayouts)
library(ggforce)
library(networkdata)
data("got")
gotS1 <- got[[1]]
got_palette <- c(
"#1A5878", "#C44237", "#AD8941", "#E99093",
"#50594B", "#8968CD", "#9ACD32"
)
## compute a clustering for node colors
V(gotS1)$clu <- as.character(membership(cluster_louvain(gotS1)))
## compute degree as node size
V(gotS1)$size <- degree(gotS1)
```
Everything that was covered so far should be enough to produce nice network visualizations, especially for scientific publications. However, `ggraph` has a lot more advanced functions/parameter settings to further enhance your visualization. If you are looking for something specific, it is always a good idea to read the documentation of the geoms.
## use the ggforce
The `ggforce` package works pretty nicely with `ggraph`. You can, for instance, use
the `geom_mark_*()` functions to highlight clusters.
```{r}
#| label: network-grps
set.seed(665)
## create network with a group structure
g <- sample_islands(9, 40, 0.4, 15)
g <- igraph::simplify(g)
V(g)$grp <- as.character(rep(1:9, each = 40))
bb <- layout_as_backbone(g, keep = 0.4)
E(g)$col <- F
E(g)$col[bb$backbone] <- T
```
```{r}
#| label: network-grps-sol
ggraph(g, #|
layout = "manual",
x = bb$xy[, 1],
y = bb$xy[, 2]
) +
geom_edge_link0(aes(col = col), width = 0.2) +
geom_node_point(aes(fill = grp), shape = 21, size = 3) +
geom_mark_hull(
aes(x, y, group = grp, fill = grp),
concavity = 4,
expand = unit(2, "mm"),
alpha = 0.25
) +
scale_color_brewer(palette = "Set1") +
scale_fill_brewer(palette = "Set1") +
scale_edge_color_manual(values = c(rgb(0, 0, 0, 0.3), rgb(0, 0, 0, 1))) +
theme_graph() +
theme(legend.position = "none")
```
Of course you can also add a label to your clusters.
```{r}
#| label: network-grps-label-sol
ggraph(g, #|
layout = "manual",
x = bb$xy[, 1],
y = bb$xy[, 2]
) +
geom_edge_link0(aes(col = col), width = 0.2) +
geom_node_point(aes(fill = grp), shape = 21, size = 3) +
geom_mark_hull(
aes(x, y, group = grp, fill = grp, label = grp),
concavity = 4,
expand = unit(2, "mm"),
alpha = 0.25
) +
scale_color_brewer(palette = "Set1") +
scale_fill_brewer(palette = "Set1") +
scale_edge_color_manual(values = c(rgb(0, 0, 0, 0.3), rgb(0, 0, 0, 1))) +
theme_graph() +
theme(legend.position = "none")
```
If you want to avoid node overlaps, you can use `geom_node_voronoi()`.
So this is actually already implemented in `ggraph`, but originates from `geom_voronoi_tile()`.
```{r}
#| label: network-voronoi
ggraph(g,
layout = "manual",
x = bb$xy[, 1],
y = bb$xy[, 2]
) +
geom_edge_link0(aes(filter = !col, col = col), width = 0.2) +
geom_node_voronoi(
aes(x, y, fill = grp),
max.radius = 0.4,
expand = unit(-0.5, "mm"),
colour = "black"
) +
scale_color_brewer(palette = "Set1") +
scale_fill_brewer(palette = "Set1") +
scale_edge_color_manual(values = c(rgb(0, 0, 0, 0.3), rgb(0, 0, 0, 1))) +
theme(
legend.position = "none",
panel.grid = element_blank(),
axis.ticks = element_blank(),
axis.text = element_blank()
) +
theme_graph() +
theme(legend.position = "none")
```
## Small tricks for common problems
> "How can I achieve that my directed edges stop at the node border, independent from the node size?"
This one has given me headaches for the longest time. No matter what I tried, I always ended up with
something like the below plot.
```{r}
#| label: arrow-size
## create a random network
set.seed(1071)
g <- sample_pa(30, 1)
V(g)$degree <- degree(g, mode = "in")
ggraph(g, "stress") +
geom_edge_link(
aes(end_cap = circle(node2.degree + 2, "pt")),
edge_colour = "black",
arrow = arrow(
angle = 10,
length = unit(0.15, "inches"),
ends = "last",
type = "closed"
)
) +
geom_node_point(aes(size = degree), col = "grey66", show.legend = FALSE) +
scale_size(range = c(3, 11)) +
theme_graph()
```
The overlap can be avoided by using the `I()` function from base R, which
treats the entries of a vector "as is". So we know that if a node has degree 5, it will be mapped to
a circle with radius (or diameter?) "5pt". Since this means, that you have no control over the scaling,
you need to do that beforehand.
```{r}
#| label: arrows-size-sol
## this function is borrowed from the ambient package
normalise <- function(x, from = range(x), to = c(0, 1)) {
x <- (x - from[1]) / (from[2] - from[1])
if (!identical(to, c(0, 1))) {
x <- x * (to[2] - to[1]) + to[1]
}
x
}
## map to the range you want
V(g)$degree <- normalise(V(g)$degree, to = c(3, 11))
ggraph(g, "stress") +
geom_edge_link(
aes(end_cap = circle(node2.degree + 2, "pt")),
edge_colour = "grey25",
arrow = arrow(
angle = 10,
length = unit(0.15, "inches"),
ends = "last",
type = "closed"
)
) +
geom_node_point(aes(size = I(degree)), col = "grey66") +
theme_graph()
```
I would not be surprised though if there is an even easier fix for this problem.
> "How can I lower the opacity of nodes without making edges visible underneath?"
One of the rules I try to follow is that edges should not be visible on top of nodes.
Usually that is easy to achieve by drawing the edges before the nodes. But if you
want to lower the opacity of nodes, they do become visible again.
```{r}
#| label: alpha-nodes
g <- sample_gnp(20, 0.5)
V(g)$degree <- degree(g)
ggraph(g, "stress") +
geom_edge_link(edge_colour = "grey66") +
geom_node_point(
size = 8,
aes(alpha = degree),
col = "red",
show.legend = FALSE
) +
theme_graph()
```
The solution is rather simple. Just add a node layer with the same aesthetics below with
`alpha=1` (default) and `color="white"` (or the background color of the plot).
```{r}
#| label: alpha-nodes-sol
ggraph(g, "stress") +
geom_edge_link(edge_colour = "grey66") +
geom_node_point(size = 8, col = "white") +
geom_node_point(
aes(alpha = degree),
size = 8,
col = "red",
show.legend = FALSE
) +
theme_graph()
```
Of course you could also use `start_cap` and `end_cap` here, but you may have to fiddle again as in the last example.
> "How can I enhance readability of node labels in hairball graphs?"
Sometimes it is really hard to make labels readable when the network is very cluttered
```{r}
#| label: unreadable-labels
#| fig-width: 8
#| fig-height: 8
g <- sample_gnp(50, 0.7)
V(g)$name <- sapply(1:50, function(x) paste0(sample(LETTERS, 4), collapse = ""))
E(g)$weight <- runif(ecount(g))
ggraph(g) +
geom_edge_link0(aes(edge_color = weight, edge_linewidth = weight), show.legend = FALSE) +
geom_node_point(size = 8, color = "#44a6c6") +
geom_node_text(aes(label = name), fontface = "bold") +
scale_edge_color_continuous(low = "grey66", high = "black") +
scale_edge_width(range = c(0.1, 0.5)) +
theme_graph() +
coord_fixed()
```
Here you can make use of the fact that the layout of the nodes are stored in a "hidden" data frame when a `ggraph` object is constructed. That means you can use other geoms from other packages. In this case, the `shadowtext` package as shown below.
```{r}
#| label: shadowtext-labels
#| fig-width: 8
#| fig-height: 8
ggraph(g, "stress") +
geom_edge_link0(aes(edge_color = weight, edge_linewidth = weight), show.legend = FALSE) +
geom_node_point(size = 8, color = "#44a6c6") +
shadowtext::geom_shadowtext(aes(x, y, label = name), color = "black", size = 4, bg.colour = "white") +
scale_edge_color_continuous(low = "grey66", high = "black") +
scale_edge_width(range = c(0.1, 0.5)) +
theme_graph() +
coord_fixed()
```
## Edge bundling
```{r}
#| label: fig-force-bundle
gr <- edgebundle::us_flights
states <- map_data("state")
ggraph(gr, x = longitude, y = latitude) +
geom_polygon(aes(long, lat, group = group), states, color = "white", linewidth = 0.2) +
coord_sf(crs = "NAD83", default_crs = sf::st_crs(4326)) +
geom_edge_bundle_force(color = "white", width = 0.05)
```
```{r}
#| label: fig-path-bundle
ggraph(gr, x = longitude, y = latitude) +
geom_polygon(aes(long, lat, group = group), states, color = "white", linewidth = 0.2) +
coord_sf(crs = "NAD83", default_crs = sf::st_crs(4326)) +
geom_edge_bundle_path(color = "white", width = 0.05)
```
## snahelper
Even with a lot of experience, it may still be a painful process to produce nice looking
figures by writing `ggraph` code. Enter the `snahelper`.
```r
install.packages("snahelper")
```
The `snahelper` is an RStudio addin which provides
you with a GUI to plot networks. Instead of writing code, you simply use drop-down menus to assign
attributes to aesthetics or change appearances globally. One great feature of the addin is that you
can adjust the position of nodes individually if you are not satisfied with their location.
Once you are done, you can either directly export the figure to png or automatically insert the code to produce the figure into your script. That way, you can review the code and hopefully learn something from it. Below if a demo that shows its functionality.
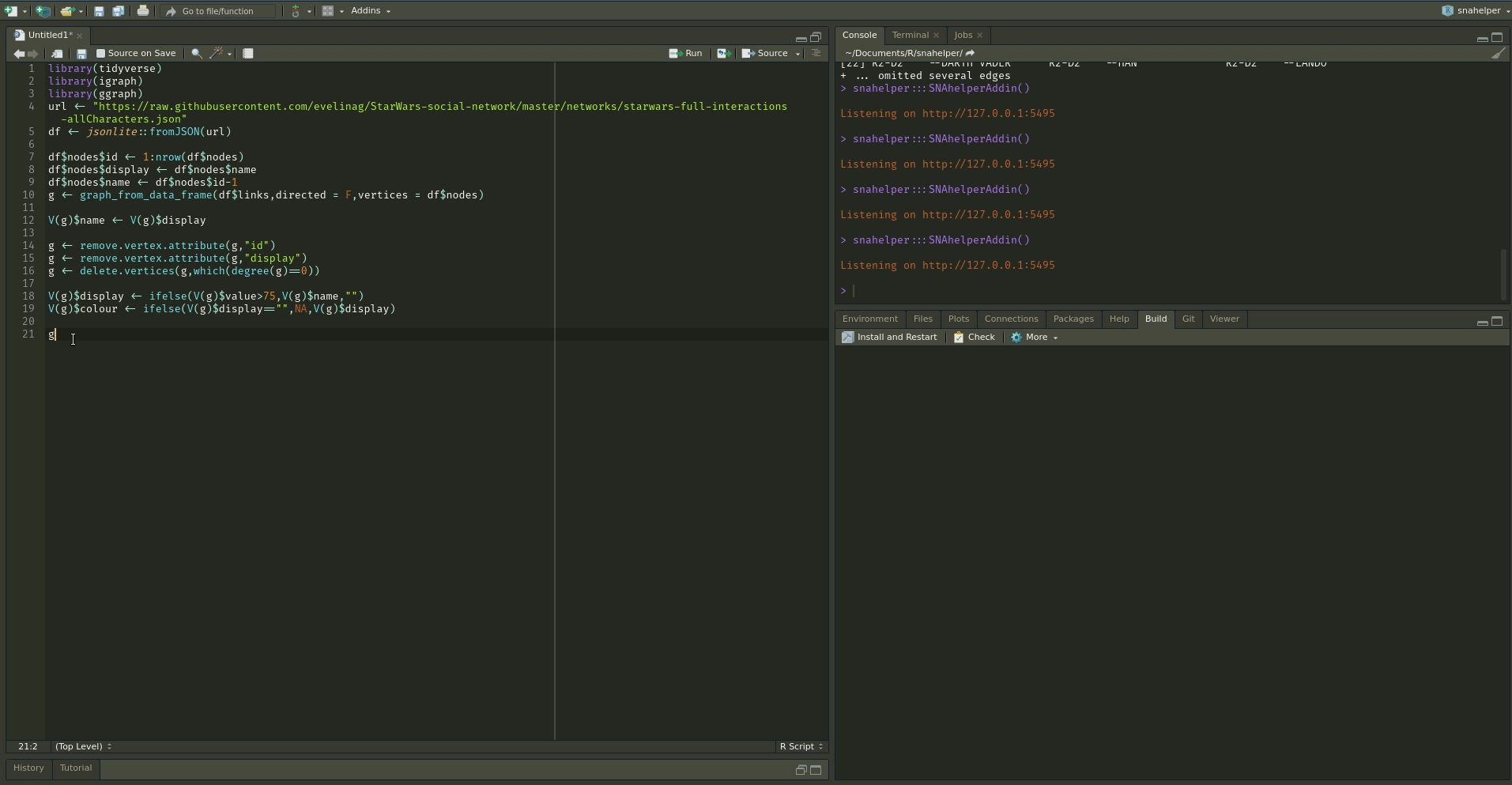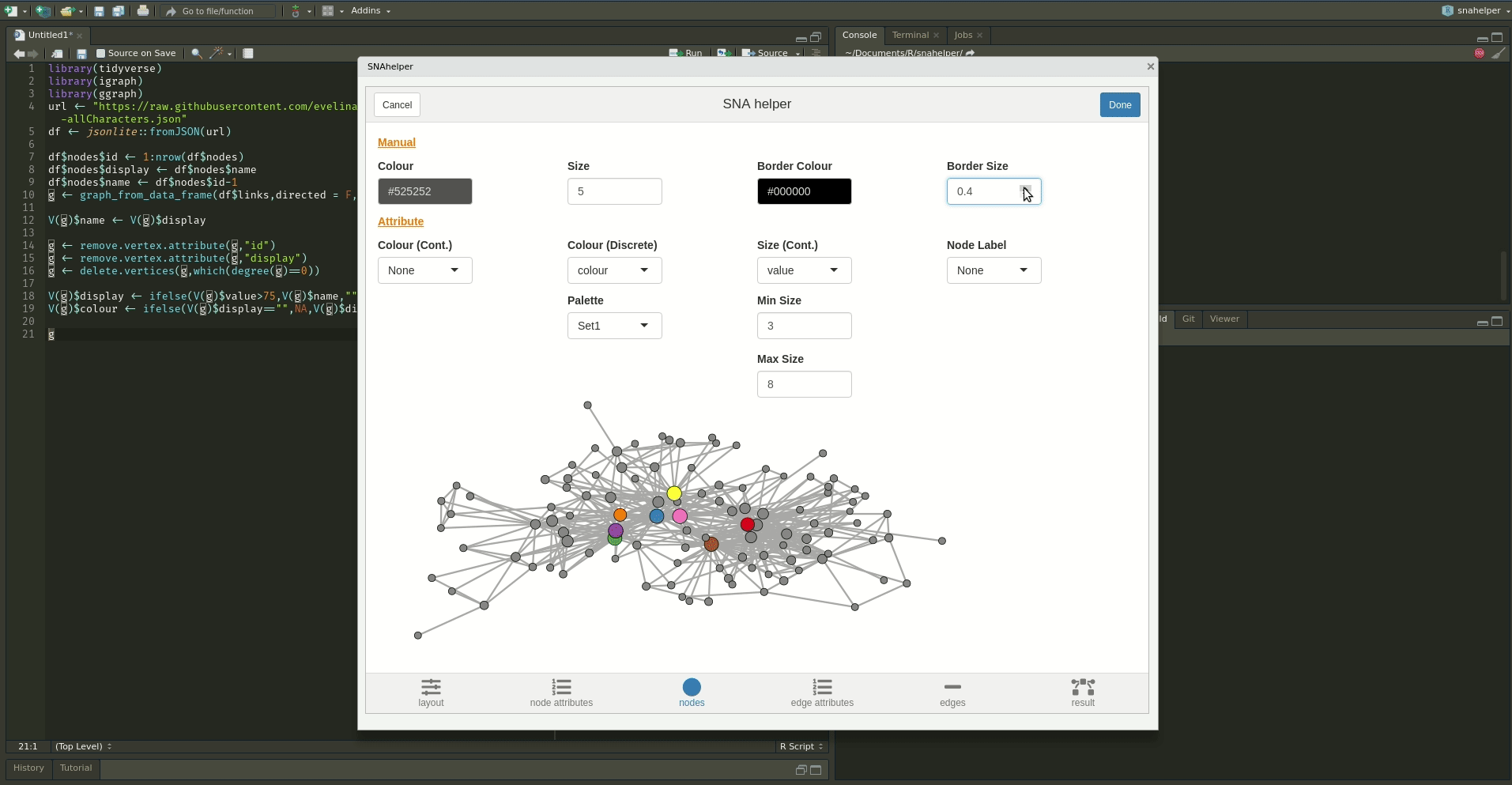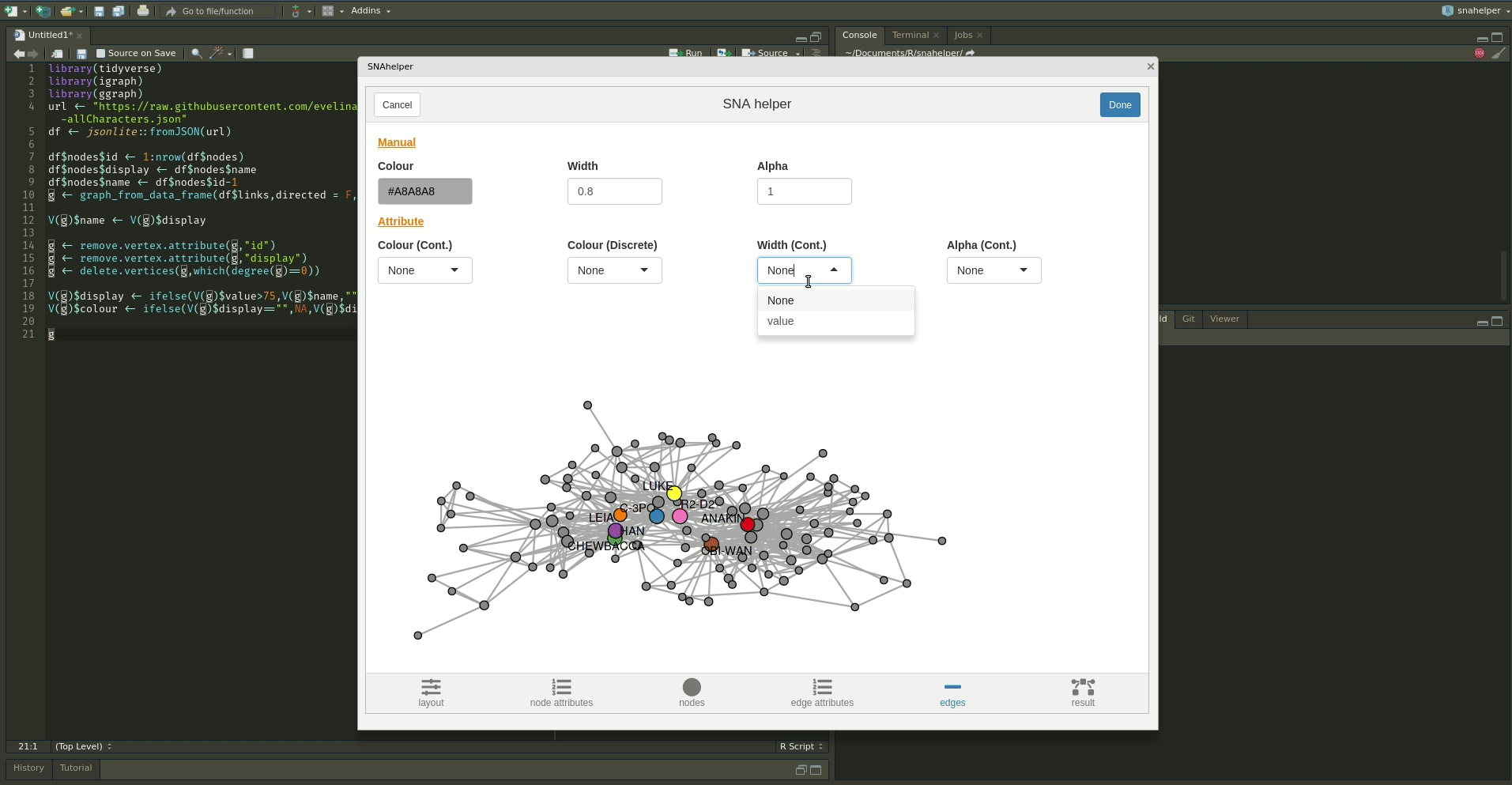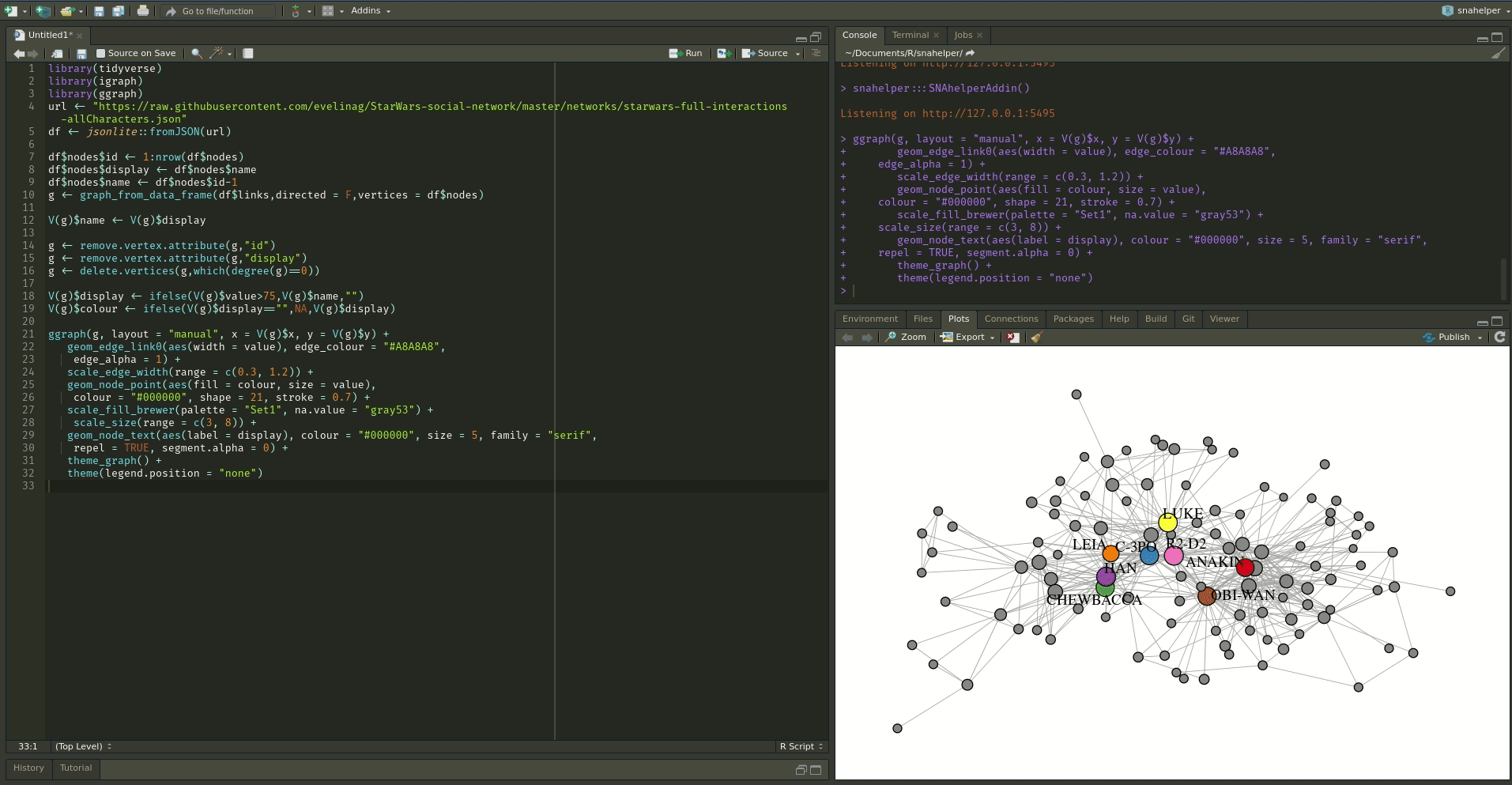
To use the addin, simply highlight the variable name of your network within an R script and
choose the SNAhelper from the Addins drop-down menu within RStudio. You can find more about the
Addin on its dedicated [pkgdown page](http://snahelper.schochastics.net)
## Misc
Some things that I frequently use are the following:
- change the `end_cap` in `geom_edge_link()` to end edges before reaching the node. This is helpful
for directed edges to not make the arrows disappear.
- `legend.position` in `theme()` controls all legends at once. If you don't want to show a specific legend,
use `guide = "none"` in the respective `scale_*` function.
- use `scale_color_viridis_c()` and `scale_color_viridis_d()`. The viridis colour palette makes plots easier to read by those with colorblindness and print well in grey scale.