-
Notifications
You must be signed in to change notification settings - Fork 29
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Should I do the channel location first? #20
Comments
|
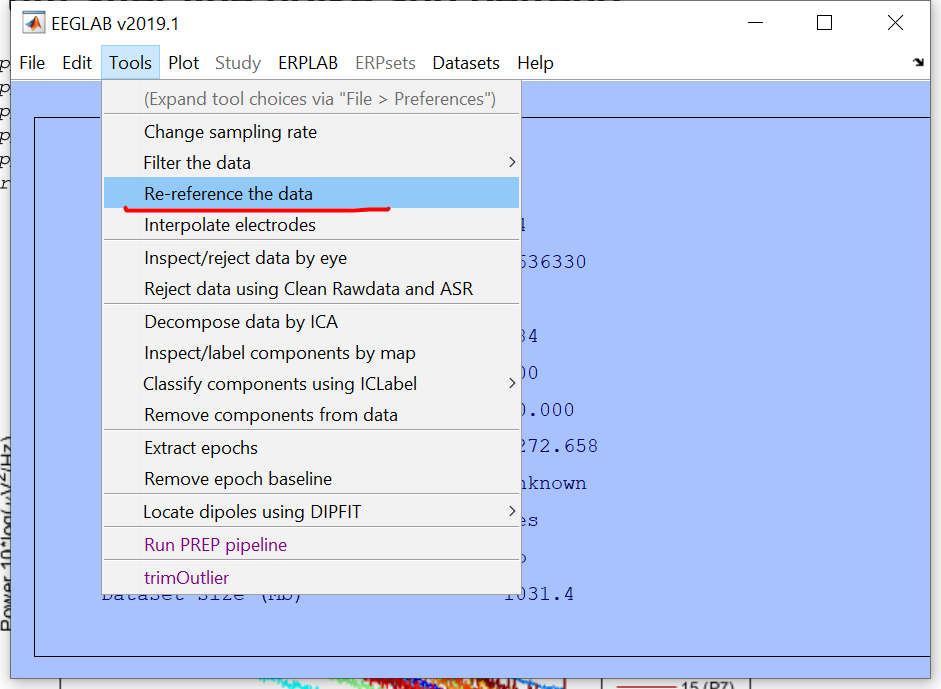
This is the EEGLAB menu, not the PREP pipeline menu. It looks like you have already loaded a dataset. Go to the Tools menu and select the menu item for the PREP pipeline. There will be a menu with buttons for each stage. You will need to set channel numbers in each stage. If you don't PREP uses them all. For line noise removal, you want to include EOG channels. For detrend channels, you probably want to include EOG. For doing the reference, there are actually three sets of channel numbers you need to set.
|
|
If you use the default settings for PREP is to high-pass filter at 1 Hz for calculating the robust reference only, but the postprocess keepFiltered parameter is false by default. You are not getting the filtered signal. If you want it to be high-pass filtered, you need to set keepFiltered to true. Beyond that, it would be good to go through the tutorials for EEGLAB. They have a recommended processing pipeline. You should read through their recommendations. before proceeding. You should also join the EEGLAB mailing list. They really focus on these types of user issues and are quick to respond with useful information. |
|
HERE are two alternative strategies for preprocessing.
These are mutually exclusive. PREP is a lighter touch strategy in that it tries to do as little as possible to the signal. ASR depends on the std parameter. After you do one (and only one of these) you should consider doing ICA and ICALabel to get rid of bad ICs. It is important that you read the documentation. I think the easiest for someone new to the field is to use clean_rawdata. You might want to take a look at our paper describing differences in results: https://pubmed.ncbi.nlm.nih.gov/32217478/. Again, you should join the EEGLAB mailing list and you can get some opinions on the list about the best way to proceed for your application. |
|











Dear PREP pipeline group
I have scaned your paper and read the introduction to prep, I appreciate what you have done, but I still felt confuse when I use this plugin on EEGlab. I check all the parameter setting choice, and find none directing to montage.
All in all, I want to want should I do with my data, before I use the tool. should I complete channel location first? should I discard the Veog channel. I cannot find any glue about that on your website.
After the process, I cannot see any change through this EEGlab interface. no channel location, no reference.
The text was updated successfully, but these errors were encountered: