Getting Started
-
The latest version of R can be downloaded and installed from the Comprehensive R Archive Network (CRAN).
-
ExpressionDB was built using ShinyApps, an extension of RStudio. A free, open-source copy of RStudio can be downloaded here.
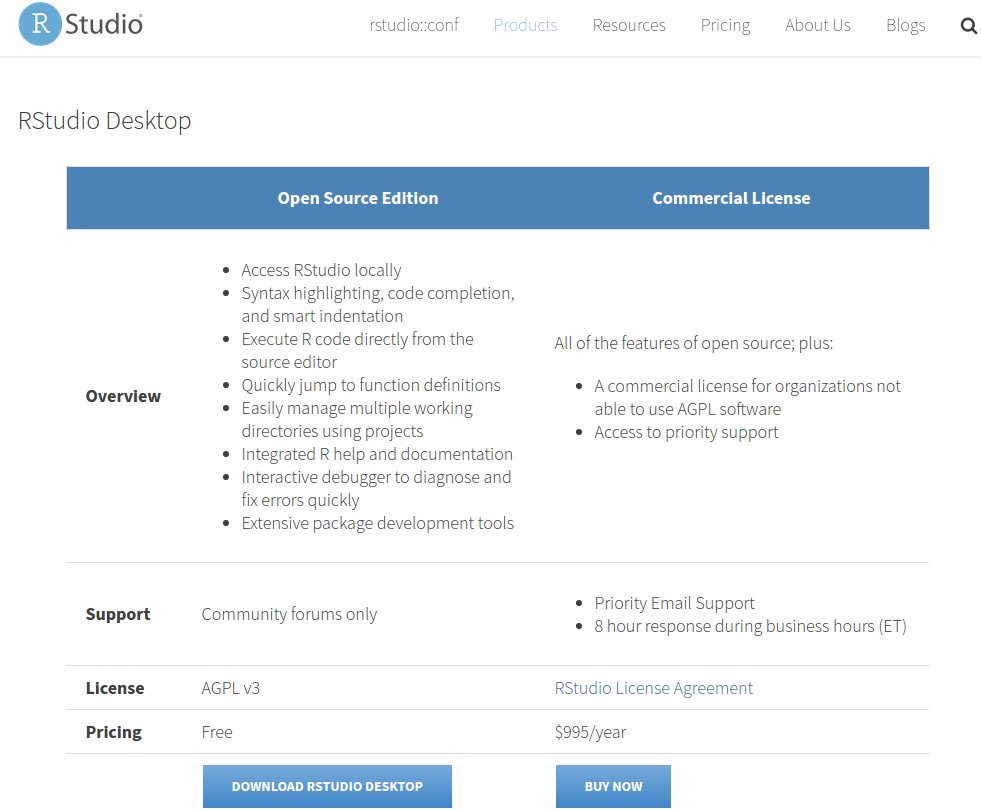
- Download the ExpressionDB package.
- The easiest way to launch ExpressionDB is through RStudio. Open RStudio and navigate to the folder containing your copy of ExpressionDB.

-
Open global.R

-
Click
Run Appgreen arrow from the top right side of the file window in RStudio.
If you have not installed the Shiny R package before, RStudio will ask you if you want to install it. Click yes.

Then two things will happen. First, all the necessary dependencies for ExpressionDB will install and load. This will take some time, but you only have to do this on the first instance. Second, a sample dataset will load, and statistics for the gene expression will be calculated.
-
The ExpressionDB will launch either in Chrome or internally within RStudio, allowing you to explore and visualize the data.
-
To stop the application, press
escat the command line console in RStudio. -
In the future (or if you've already installed the Shiny R package), you can launch the app by typing
shiny::runApp("<my_working_directory>")at the console prompt. Be sure to replace <my_working_directory> with the location of your directory, or set your working directory to the ExpressionDB folder.
See our User's Guide to see some examples of how you can use this code to build your own RNA-seq expression database.
- data.table version 1.10.4
- dplyr version 0.7.2
- DT version 0.2
- dtplyr version 0.0.2
- ggplot2 version 2.2.1
- heatmaply version 0.10.1
- RColorBrewer version 1.1-2
- shiny version 1.0.3
- shinydashboard version 0.6.1
- stringr version 1.2.0
- tidyr version 0.6.3
Creative Commons License: Hughes, Lewis, and Hughes, 2017