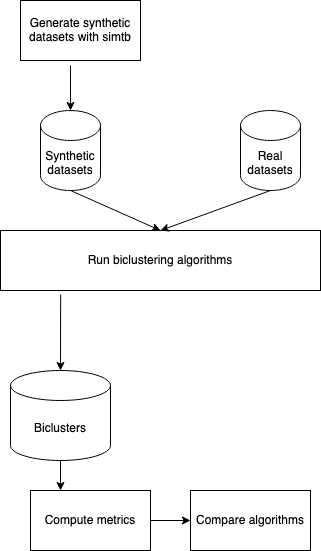
This repository aims to provide support to the associated paper. It acts as an repository to archive:
- The scripts that generate the synthetic datasets
- The datasets used in the paper
- Scripts to generate the biclusters
- Functions to read and analyse results
- Some generated biclusters to act as example
- SimTB Generators: Related with the process of generating synthetic datasets
- Data Collections: Data used for the analysis
- Biclustering Generators: Scripts for generating biclusters from the data
- Biclustering Analysis: Functions to compute the analysis metrics
The paper have three data colections.
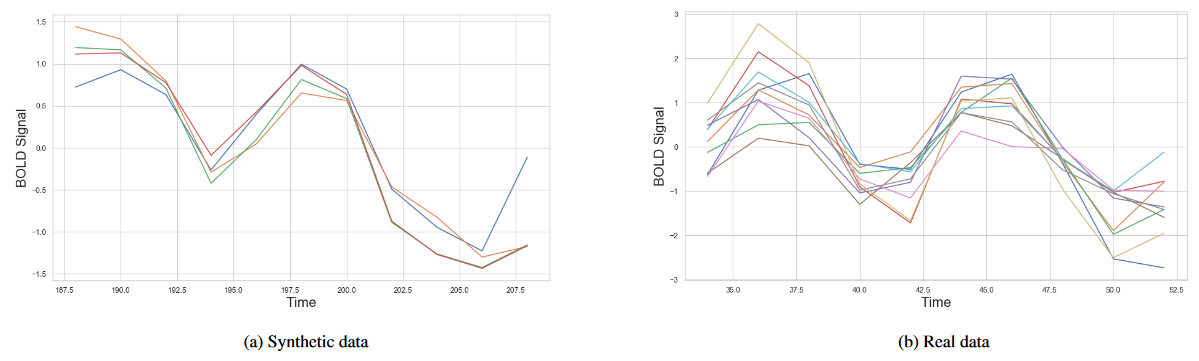
- First data collection: A single synthetic subject
- Second data collection: Twenty synthetic subjects
- Third data collection: Twenty real subjects
- ISA
- XMotifs
- FABIA
- Spectral
- Bimax
- CCC
- BicPAM
- K-Means
- Spectral
- Ward's
During the development of this paper some software was needed depending on the part of the analysis:
- MATLAB will be needed to run the SimTB fMRI data simulator, to generate the synthetic datasets.
- R will be needed for running most of the biclustering algorithms (FABIA, ISA, Bimax, XMotifs, Spectral).
- For FABIA, you need the fabia Package.
- For ISA, you need the isa2 Package.
- For the remaining algorithms, you need the biclust Package.
- Python will be used for running the clustering algorithms (Spectral, K-means, Ward's hierarchical method) and analysing the biclustering solutions.
- For the clustering algorithms, we used the implementations provided by scikit-learn.
- Other traditional python libraries are used during the scripts.
- BicPAM is implemented in the BicPAMS tool.
- Both an Desktop GUI and a Java API are provided.
- CCC is implemented as part of the BiGGEsTS software (GUI implementation).