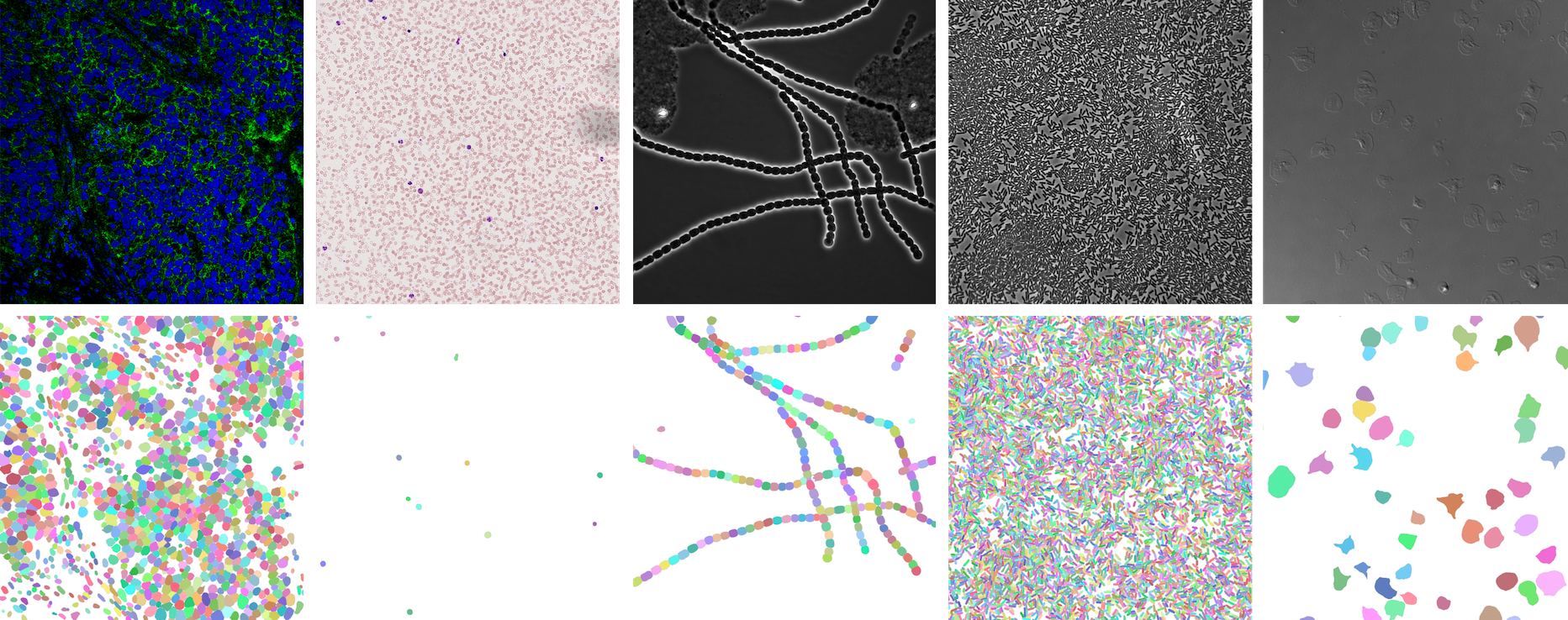
 https://openreview.net/forum?id=YtgRjBw-7GJ
https://openreview.net/forum?id=YtgRjBw-7GJ
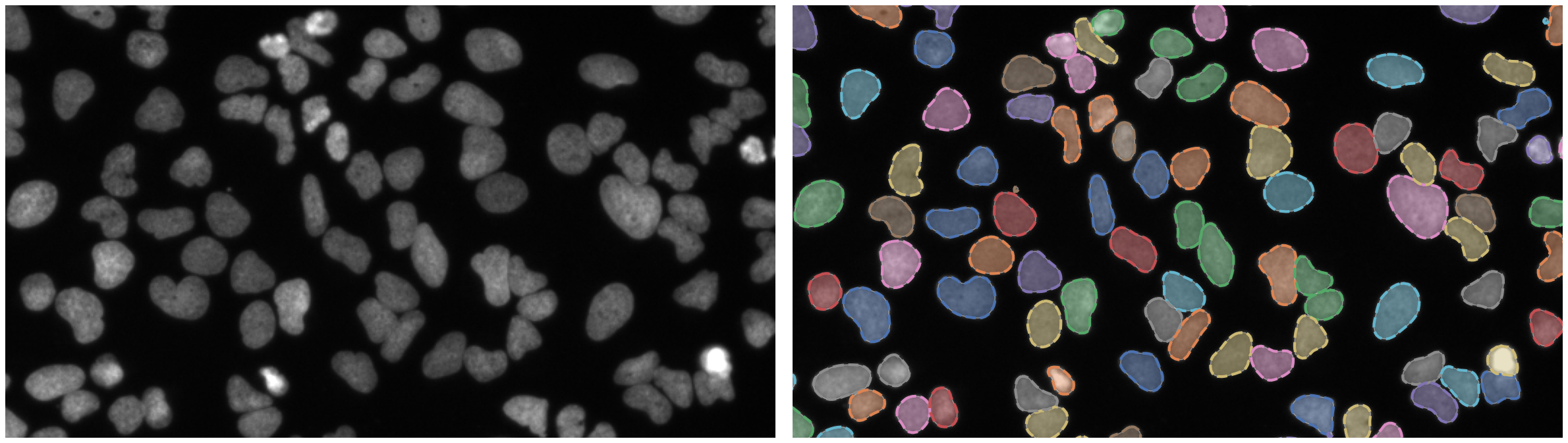 https://bbbc.broadinstitute.org/BBBC039 (CC0)
https://bbbc.broadinstitute.org/BBBC039 (CC0)
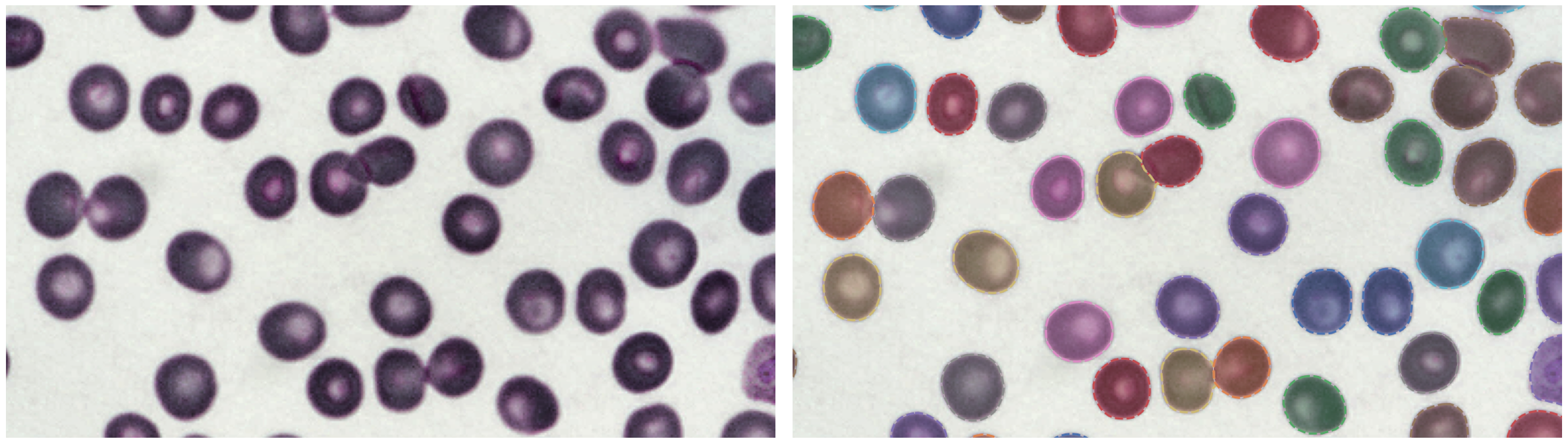 https://bbbc.broadinstitute.org/BBBC041 (CC BY-NC-SA 3.0)
https://bbbc.broadinstitute.org/BBBC041 (CC BY-NC-SA 3.0)
Make sure you have PyTorch installed.
pip install -U celldetection
pip install git+https://github.com/FZJ-INM1-BDA/celldetection.git
model = cd.fetch_model(model_name, check_hash=True)| model name | training data | link |
|---|---|---|
ginoro_CpnResNeXt101UNet-fbe875f1a3e5ce2c |
BBBC039, BBBC038, Omnipose, Cellpose, Sartorius - Cell Instance Segmentation, Livecell, NeurIPS 22 CellSeg Challenge | 🔗 |
Run a demo with a pretrained model
import torch, cv2, celldetection as cd
from skimage.data import coins
from matplotlib import pyplot as plt
# Load pretrained model
device = 'cuda' if torch.cuda.is_available() else 'cpu'
model = cd.fetch_model('ginoro_CpnResNeXt101UNet-fbe875f1a3e5ce2c', check_hash=True).to(device)
model.eval()
# Load input
img = coins()
img = cv2.cvtColor(img, cv2.COLOR_GRAY2RGB)
print(img.dtype, img.shape, (img.min(), img.max()))
# Run model
with torch.no_grad():
x = cd.to_tensor(img, transpose=True, device=device, dtype=torch.float32)
x = x / 255 # ensure 0..1 range
x = x[None] # add batch dimension: Tensor[3, h, w] -> Tensor[1, 3, h, w]
y = model(x)
# Show results for each batch item
contours = y['contours']
for n in range(len(x)):
cd.imshow_row(x[n], x[n], figsize=(16, 9), titles=('input', 'contours'))
cd.plot_contours(contours[n])
plt.show()import celldetection as cdContour Proposal Networks
cd.models.CPNcd.models.CpnU22cd.models.CPNCorecd.models.CpnResUNetcd.models.CpnSlimU22cd.models.CpnWideU22cd.models.CpnResNet18FPNcd.models.CpnResNet34FPNcd.models.CpnResNet50FPNcd.models.CpnResNeXt50FPNcd.models.CpnResNet101FPNcd.models.CpnResNet152FPNcd.models.CpnResNet18UNetcd.models.CpnResNet34UNetcd.models.CpnResNet50UNetcd.models.CpnResNeXt101FPNcd.models.CpnResNeXt152FPNcd.models.CpnResNeXt50UNetcd.models.CpnResNet101UNetcd.models.CpnResNet152UNetcd.models.CpnResNeXt101UNetcd.models.CpnResNeXt152UNetcd.models.CpnWideResNet50FPNcd.models.CpnWideResNet101FPNcd.models.CpnMobileNetV3LargeFPNcd.models.CpnMobileNetV3SmallFPN
PyTorch Image Models (timm)
Also have a look at Timm Documentation.
import timm
timm.list_models(filter='*') # explore available modelsSegmentation Models PyTorch (smp)
import segmentation_models_pytorch as smp
smp.encoders.get_encoder_names() # explore available modelsencoder = cd.models.SmpEncoder(encoder_name='mit_b5', pretrained='imagenet')Find a list of Smp Encoders in the smp documentation.
U-Nets
# U-Nets are available in 2D and 3D
import celldetection as cd
model = cd.models.ResNeXt50UNet(in_channels=3, out_channels=1, nd=3)cd.models.U22cd.models.U17cd.models.U12cd.models.UNetcd.models.WideU22cd.models.SlimU22cd.models.ResUNetcd.models.UNetEncodercd.models.ResNet50UNetcd.models.ResNet18UNetcd.models.ResNet34UNetcd.models.ResNet152UNetcd.models.ResNet101UNetcd.models.ResNeXt50UNetcd.models.ResNeXt152UNetcd.models.ResNeXt101UNetcd.models.WideResNet50UNetcd.models.WideResNet101UNetcd.models.MobileNetV3SmallUNetcd.models.MobileNetV3LargeUNet
MA-Nets
# Many MA-Nets are available in 2D and 3D
import celldetection as cd
encoder = cd.models.ConvNeXtSmall(in_channels=3, nd=3)
model = cd.models.MaNet(encoder, out_channels=1, nd=3)Feature Pyramid Networks
cd.models.FPNcd.models.ResNet18FPNcd.models.ResNet34FPNcd.models.ResNet50FPNcd.models.ResNeXt50FPNcd.models.ResNet101FPNcd.models.ResNet152FPNcd.models.ResNeXt101FPNcd.models.ResNeXt152FPNcd.models.WideResNet50FPNcd.models.WideResNet101FPNcd.models.MobileNetV3LargeFPNcd.models.MobileNetV3SmallFPN
ConvNeXt Networks
# ConvNeXt Networks are available in 2D and 3D
import celldetection as cd
model = cd.models.ConvNeXtSmall(in_channels=3, nd=3)Residual Networks
# Residual Networks are available in 2D and 3D
import celldetection as cd
model = cd.models.ResNet50(in_channels=3, nd=3)Mobile Networks
Find us on Docker Hub: https://hub.docker.com/r/ericup/celldetection
You can pull the latest version of celldetection via:
docker pull ericup/celldetection:latest
CPN inference via Docker with GPU
docker run --rm \
-v $PWD/docker/outputs:/outputs/ \
-v $PWD/docker/inputs/:/inputs/ \
-v $PWD/docker/models/:/models/ \
--gpus="device=0" \
celldetection:latest /bin/bash -c \
"python cpn_inference.py --tile_size=1024 --stride=768 --precision=32-true"
CPN inference via Docker with CPU
docker run --rm \
-v $PWD/docker/outputs:/outputs/ \
-v $PWD/docker/inputs/:/inputs/ \
-v $PWD/docker/models/:/models/ \
celldetection:latest /bin/bash -c \
"python cpn_inference.py --tile_size=1024 --stride=768 --precision=32-true --accelerator=cpu"
You can also pull our Docker images for the use with Apptainer (formerly Singularity) with this command:
apptainer pull --dir . --disable-cache docker://ericup/celldetection:latest
Find us on Hugging Face and upload your own images for segmentation: https://huggingface.co/spaces/ericup/celldetection
There's also an API (Python & JavaScript), allowing you to utilize community GPUs (currently Nvidia A100) remotely!
Hugging Face API
from gradio_client import Client
# Define inputs (local filename or URL)
inputs = 'https://raw.githubusercontent.com/scikit-image/scikit-image/main/skimage/data/coins.png'
# Set up client
client = Client("ericup/celldetection")
# Predict
overlay_filename, img_filename, h5_filename, csv_filename = client.predict(
inputs, # str: Local filepath or URL of your input image
# Model name
'ginoro_CpnResNeXt101UNet-fbe875f1a3e5ce2c',
# Custom Score Threshold (numeric value between 0 and 1)
False, .9, # bool: Whether to use custom setting; float: Custom setting
# Custom NMS Threshold
False, .3142, # bool: Whether to use custom setting; float: Custom setting
# Custom Number of Sample Points
False, 128, # bool: Whether to use custom setting; int: Custom setting
# Overlapping objects
True, # bool: Whether to allow overlapping objects
# API name (keep as is)
api_name="/predict"
)
# Example usage: Code below only shows how to use the results
from matplotlib import pyplot as plt
import celldetection as cd
import pandas as pd
# Read results from local temporary files
img = imread(img_filename)
overlay = imread(overlay_filename) # random colors per instance; transparent overlap
properties = pd.read_csv(csv_filename)
contours, scores, label_image = cd.from_h5(h5_filename, 'contours', 'scores', 'labels')
# Optionally display overlay
cd.imshow_row(img, img, figsize=(16, 9))
cd.imshow(overlay)
plt.show()
# Optionally display contours with text
cd.imshow_row(img, img, figsize=(16, 9))
cd.plot_contours(contours, texts=['score: %d%%\narea: %d' % s for s in zip((scores * 100).round(), properties.area)])
plt.show()import { client } from "@gradio/client";
const response_0 = await fetch("https://raw.githubusercontent.com/scikit-image/scikit-image/main/skimage/data/coins.png");
const exampleImage = await response_0.blob();
const app = await client("ericup/celldetection");
const result = await app.predict("/predict", [
exampleImage, // blob: Your input image
// Model name (hosted model or URL)
"ginoro_CpnResNeXt101UNet-fbe875f1a3e5ce2c",
// Custom Score Threshold (numeric value between 0 and 1)
false, .9, // bool: Whether to use custom setting; float: Custom setting
// Custom NMS Threshold
false, .3142, // bool: Whether to use custom setting; float: Custom setting
// Custom Number of Sample Points
false, 128, // bool: Whether to use custom setting; int: Custom setting
// Overlapping objects
true, // bool: Whether to allow overlapping objects
// API name (keep as is)
api_name="/predict"
]);Find our Napari Plugin here: https://github.com/FZJ-INM1-BDA/celldetection-napari
Find out more about Napari here: https://napari.org
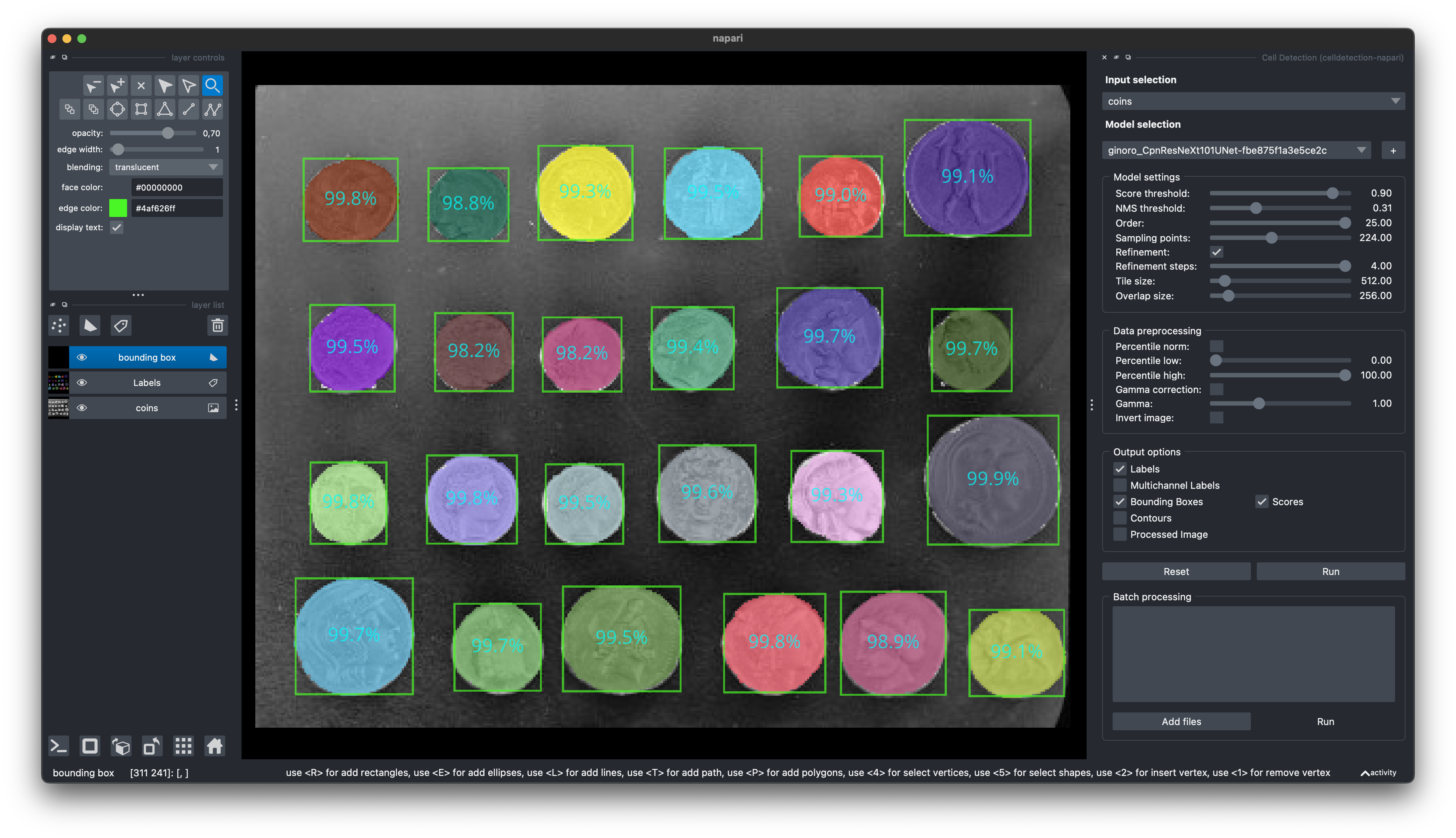 You can install it via pip:
You can install it via pip:
pip install git+https://github.com/FZJ-INM1-BDA/celldetection-napari.git
- NeurIPS 2022 Cell Segmentation Challenge: Winner Finalist Award
If you find this work useful, please consider giving a star ⭐️ and citation:
@article{UPSCHULTE2022102371,
title = {Contour proposal networks for biomedical instance segmentation},
journal = {Medical Image Analysis},
volume = {77},
pages = {102371},
year = {2022},
issn = {1361-8415},
doi = {https://doi.org/10.1016/j.media.2022.102371},
url = {https://www.sciencedirect.com/science/article/pii/S136184152200024X},
author = {Eric Upschulte and Stefan Harmeling and Katrin Amunts and Timo Dickscheid},
keywords = {Cell detection, Cell segmentation, Object detection, CPN},
}

