I. Introduction
This topic originally stems from Kaggle (https://www.kaggle.com/navoneel/brain-mri-images-for-brain-tumor-detection).
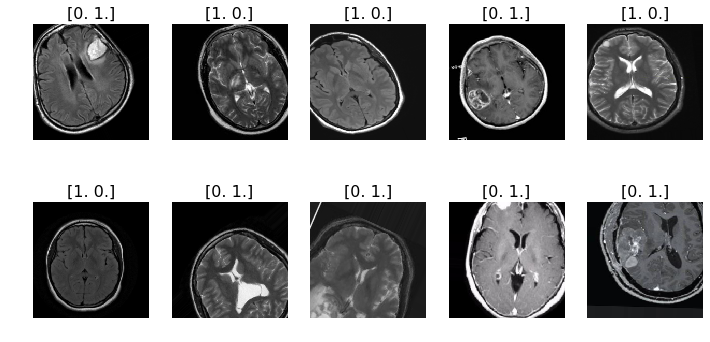
The purpose of this project is to help finding potential tumors in the MRI images of the patients brains. In the picture above we can nicely see the difference between tumors ([0. 1.]) and non-tumors ([1. 0.]). Finding tumors in an early stage could be the difference between life and death for the patient.
II. Objective
Image recognition has been implemented via convolutional neural networks (CNN), written in Python and using Keras (Tensorflow backend) as the framework. More specifically, my personal objective in this project is to have a simple comparison between self-made CNNs (build from scratch) and more famous CNNs (e.g. vgg16). !!(Please consider that the CNNs were just trained for a short time, as I just wanted to simply demonstrate 2 different approaches)!!
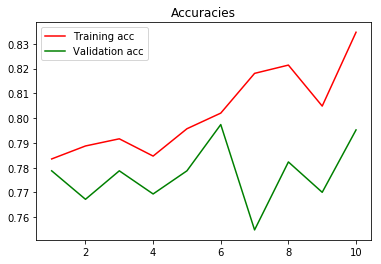
In the following we can see the Validation accuracy for my self-build CNN:
^ When we ignore the general fluctuation of the data, we can see that the mean Validation accuracy over the whole range is around 77-78%.
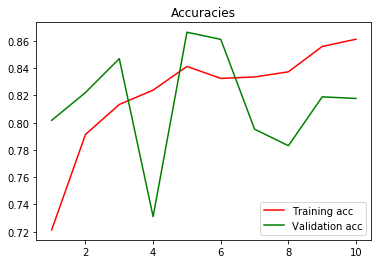
Now, next to the Validation accuracy from the pre-trained VGG16 CNN:
^ Although the data is far too noisy as well to have some concrete conclusions, it does seems that the overall Validation accuracy ranges higher than with the previous CNN (>80%). For the VGG16 (all layers untrainable), I only removed the last fully-connected layer, and added a new fully-conncted layer (trainable) with 2 outcomes (tumor and non-tumor).
III. Conclusion
The general conclusion has to be that both approaches yield a somewhat attractive prediction accuracy (~80%+). Considering that the training time was very short and the amount of images very little (~250), the accuracies should possibly be much higher when using more data and higher training times.