Information related to reproducible workflow format for IBEEM
This repository contains code and workflows for research projects within IBEEM (MSU SPG-sponsored Institute for Biodiversity, Ecology, Evolution, and Macrosystems).
These data are stored in a Google Shared Drive and the MSU HPCC IBEEM research space, accessible by collaborators. IBEEM research projects will publish open access data to accompany research papers.
All repositories should follow R packaging guidelines, even if they will not become official R packages. For questions, contact Pat Bills. If you work with other programming languages, include a separate directory structure for that (e.g., python/L0).
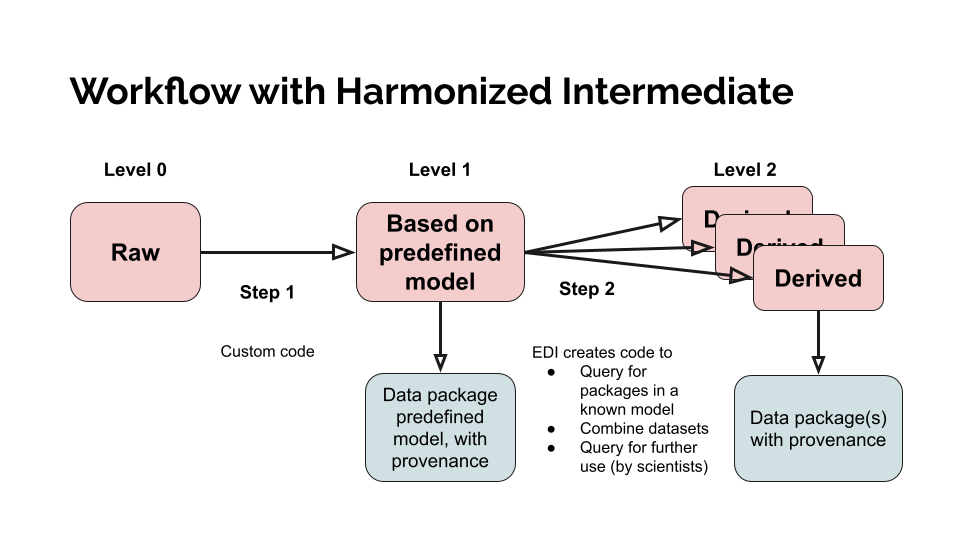
The workflow for this repository follows the guidelines set out by the Environmental Data Initiative (EDI). Briefly, this involves aligning with FAIR data practices, and employing a workflow that uses different levels for harmonization and derived data products. The overall workflow aligns with this EDI diagram:
- R: Main directory containing R scripts.
- /L0: Code & small text data files (includes R Markdown code & HTML/PDF output).
- /L1: Code to create L1 data products = cleaning & editing L0 data (includes R Markdown code & HTML/PDF output).
- /L2: Code to create derived products from L1 data (e.g., merging 2 or more L1 data products). Includes R Markdown code & HTML/PDF output.
- L0: raw, L0 data files, & metadata.
- L1: L1 data products produced from scripts that clean & edit L0 data.
- L2: derived data products from L1 data (e.g., merging 2 or more L1 data products, creating a summary variables from L1 data; e.g., diversity of species in L1 data product).
- documents supporting the data and analysis, such as manuscript outline, manuscript draft.
- meeting notes on the IBEEM research.
Funding is provided by Michigan State University Strategic Partnership Grant for IBEEM.
- Phoebe L. Zarnetske
- Pat Bills