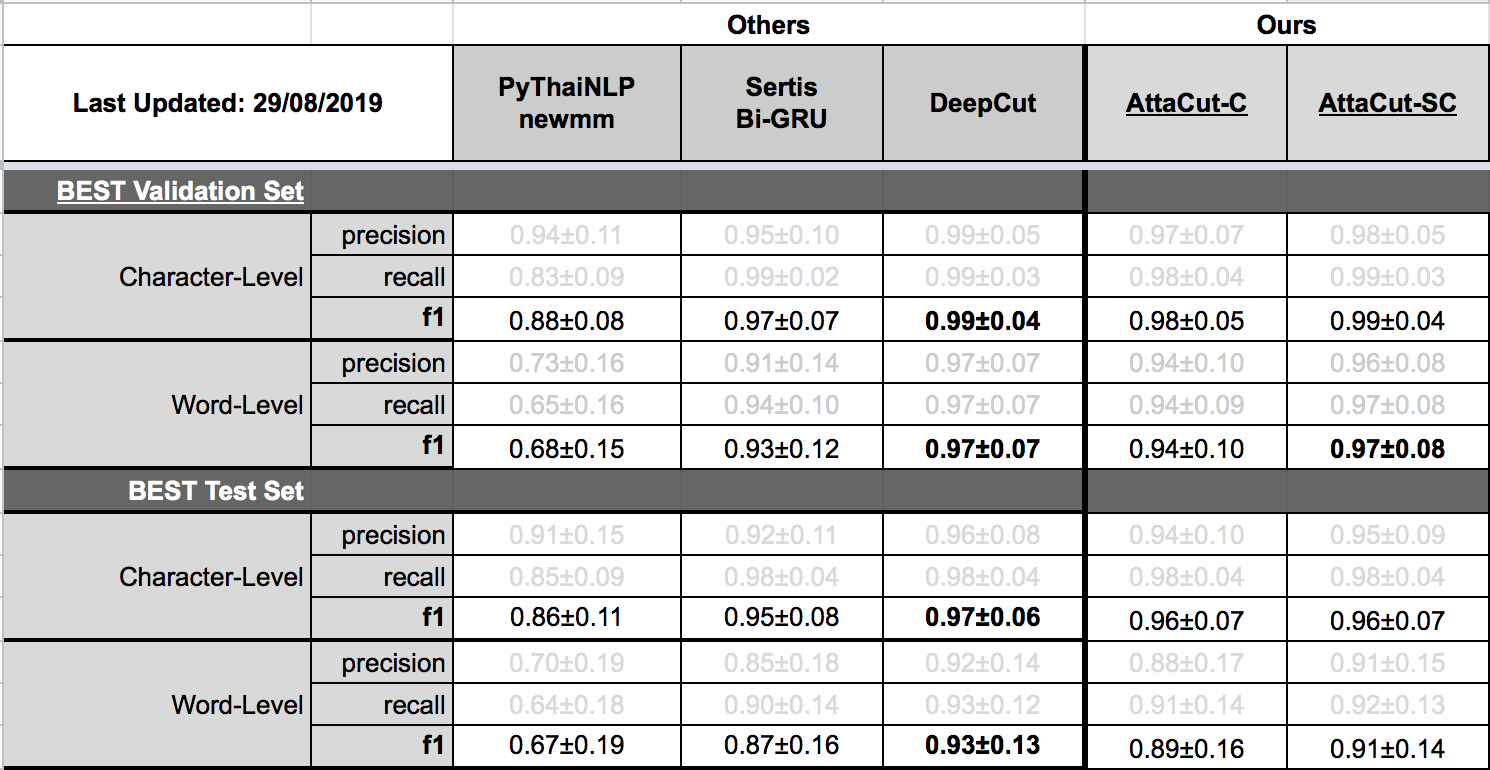
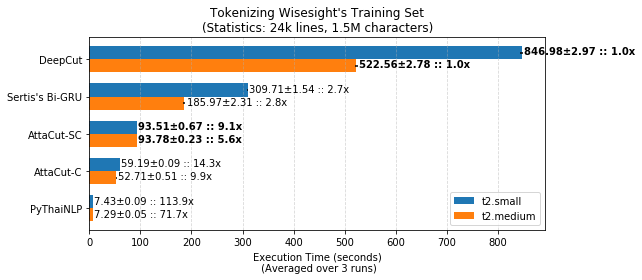
TL;DR: 3-Layer Dilated CNN on syllable and character features. It’s 6x faster than DeepCut (SOTA) while its WL-f1 on BEST is 91%, only 2% lower.
$ pip install attacut
Remarks: Windows users need to install PyTorch before the command above. Please consult PyTorch.org for more details.
$ attacut-cli -h
AttaCut: Fast and Reasonably Accurate Word Tokenizer for Thai
Usage:
attacut-cli <src> [--dest=<dest>] [--model=<model>]
attacut-cli [-v | --version]
attacut-cli [-h | --help]
Arguments:
<src> Path to input text file to be tokenized
Options:
-h --help Show this screen.
--model=<model> Model to be used [default: attacut-sc].
--dest=<dest> If not specified, it'll be <src>-tokenized-by-<model>.txt
-v --version Show version
from attacut import tokenize, Tokenizer
# tokenize `txt` using our best model `attacut-sc`
words = tokenize(txt)
# alternatively, an AttaCut tokenizer might be instantiated directly, allowing
# one to specify whether to use `attacut-sc` or `attacut-c`.
atta = Tokenizer(model="attacut-sc")
words = atta.tokenize(txt)
For better efficiency, we recommend using attacut-cli. Please consult our Google Colab tutorial for more detials.
Belows are brief summaries. More details can be found on our benchmarking page.
Please refer to our retraining page
This repository was initially done by Pattarawat Chormai, while interning at Dr. Attapol Thamrongrattanarit's NLP Lab, Chulalongkorn University, Bangkok, Thailand. Many people have involed in this project. Complete list of names can be found on Acknowledgement.