-
-
Notifications
You must be signed in to change notification settings - Fork 1.7k
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Array fully loaded into memory when trying to store into a memmap array #5985
Comments
Are the results computed in parallel, leading to the high memory usage? |
|
@TomAugspurger - no I still see the issue if I put: at the top |
|
I think this might just be due to memory_profiler including the cache/buffer size in the memory usage. If I exclude cache size variations I think everything works fine. |
|
Could it be this? dask/distributed#3032
|
I believe that this is controllable with some of the keyword arguments to |
|
which kwarg? nothing i tried in the second example over in #5367 made a difference in the topology of the task graph. |
|
I don't recall unfortunately.
…On Sun, Mar 8, 2020 at 6:17 PM Davis Bennett ***@***.***> wrote:
which kwarg? nothing i tried in the second example over in #5367
<#5367> made a difference in the
topology of the task graph.
—
You are receiving this because you commented.
Reply to this email directly, view it on GitHub
<#5985?email_source=notifications&email_token=AACKZTDSHUB5GQEWBANGI4LRGQRRFA5CNFSM4LC6XLP2YY3PNVWWK3TUL52HS4DFVREXG43VMVBW63LNMVXHJKTDN5WW2ZLOORPWSZGOEOFDDXI#issuecomment-596259293>,
or unsubscribe
<https://github.com/notifications/unsubscribe-auth/AACKZTGRK5UEMHNOXUZHUZDRGQRRFANCNFSM4LC6XLPQ>
.
|
|
This seems different from dask/distributed#3032 I think. The size of the memmapped array should be large, right? |
|
What is the status of this? I have seen a related PR #6605 but I'm not sure if memmaps are supported (it seems not from the discussion). |
Current behaviorthis question on SO encounters this error and I see the same behavior on 2022.04.0 I'm not sure if the full workaround below is necessary, but I'm also not sure how to read the array directly from the .npy file without using map_blocks MRE: import numpy as np, dask.array as dda, xarray as xr, pandas as pd, dask.distributed
# save a large numpy array
np.save('myarr.npy', np.empty(shape=(47789, 310, 310), dtype=np.float32))
cluster = dask.distributed.LocalCluster()
client = dask.distributed.Client(cluster)
print(cluster.dashboard_link)
arr = np.load('myarr.npy', mmap_mode='r')
da = dda.from_array(arr).rechunk(chunks=(100, 310, 310))
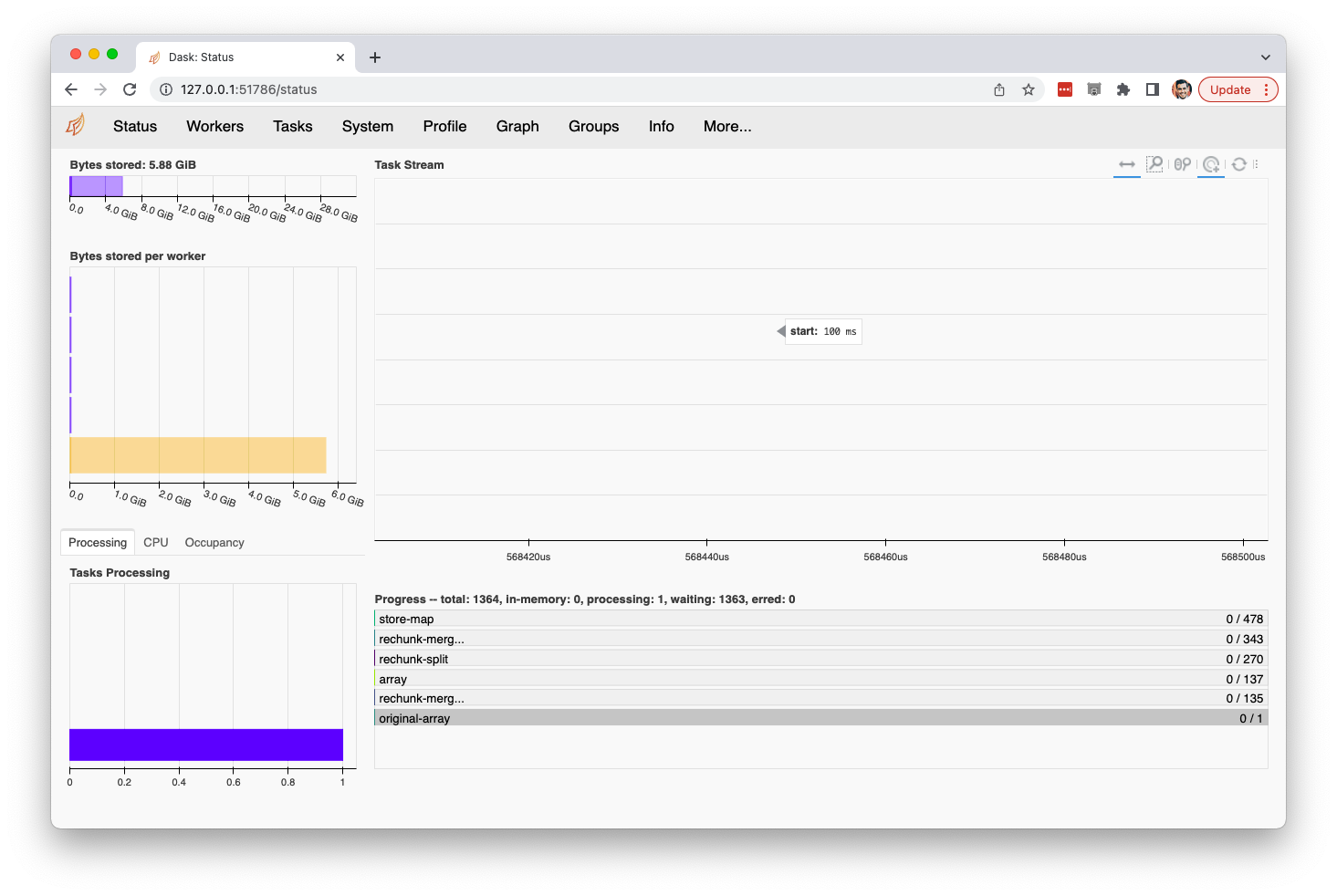
da.to_zarr('myarr.zarr', mode='w')This bottlenecks all tasks behind a giant read job "original-array" which consumes all system memory then dies workaroundI'm not sure if this is a bug or just points to a useful feature. For now, this works well: import numpy as np, dask.array as dda, xarray as xr, pandas as pd, dask.distributed
def load_npy_chunk(da, fp, block_info=None, mmap_mode='r'):
"""Load a slice of the .npy array, making use of the block_info kwarg"""
np_mmap = np.load(fp, mmap_mode=mmap_mode)
array_location = block_info[0]['array-location']
dim_slicer = tuple(list(map(lambda x: slice(*x), array_location)))
return np_mmap[dim_slicer]
def dask_read_npy(fp, chunks=None, mmap_mode='r'):
"""Read metadata by opening the mmap, then send the read job to workers"""
np_mmap = np.load(fp, mmap_mode=mmap_mode)
da = dda.empty_like(np_mmap, chunks=chunks)
return da.map_blocks(load_npy_chunk, fp=fp, mmap_mode=mmap_mode, meta=da)
# save a large numpy array
np.save('myarr.npy', np.empty(shape=(47789, 310, 310), dtype=np.float32))
cluster = dask.distributed.LocalCluster()
client = dask.distributed.Client(cluster)
da = dask_read_npy('myarr.npy', chunks=(300, -1, -1), mmap_mode='r')
da.to_zarr('myarr.zarr', mode='w') |
|
ok - in the above, using threads in the LocalCluster does solve the problem. This works: import numpy as np, dask.array as dda, xarray as xr, pandas as pd, dask.distributed
# save a large numpy array
np.save('myarr.npy', np.empty(shape=(47789, 310, 310), dtype=np.float32))
# use threads, not processes
cluster = dask.distributed.LocalCluster(processes=False)
client = dask.distributed.Client(cluster)
print(cluster.dashboard_link)
arr = np.load('myarr.npy', mmap_mode='r')
da = dda.from_array(arr).rechunk(chunks=(100, 310, 310))
da.to_zarr('myarr.zarr', mode='w')But it does seem like a read_npy function could be helpful for cases where workers can’t share memory? Happy to throw this into a PR if so, though I’d definitely need a hand getting it across the finish line as I imagine the test setup you’d want for a new reader would be pretty extensive. |
I am trying to load data from a memory-mapped array, do operations with it in dask, and store the result in another memory-mapped array. Currently this results in the entire array being loaded into memory when calling
Array.store. Here is an example:The memory profile obtained with the
memory_profilepackage is:I would have expected the array to be computed/written chunk by chunk to the output memory mapped array, resulting in a maximum memory usage of the size of a chunk, not that of the whole array. This seems like a possible bug?
The text was updated successfully, but these errors were encountered: