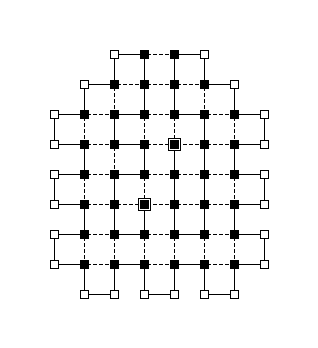
This program can be used to plot the structure of a protein in the 2D HP model. It takes as input the sequence of aminoacids, the direction of the edges between the aminoacids and the output file name. In the sequence, H stands for hydrophobic and P stands for polar. In the directions, R stands for right, L stands for left and S stands for straight.
Bellow there's an example of a figure plotted for the sequence HHHHHHHHHHHHPHPHPPHHPPHHPPHPPHHPPHHPPHPPHHPPHHPPHPHPHHHHHHHHHHHH and directions RLLSLRRLLRSRRLRLRRLLRRLLRRLRRLLRRLLRRLRRLLRRLLRRLRLRRSRLSSSLLSS.
The following projects can be used to predict the directions given the sequence
- gcc
- SDL2
How to install SDL2:
http://lazyfoo.net/tutorials/SDL/01_hello_SDL/linux/
If the link is broken, try this other one:
https://web.archive.org/web/20220706232201/http://lazyfoo.net/tutorials/SDL/01_hello_SDL/linux/
To compile, execute each one of the following commands from the root of the project:
gcc -Wall -O2 -c ./main.c -o ./main.o
gcc -o ./2dhp-plot ./main.o -lSDL2 -lm -s
After compiling it, you can run the 2dhp-plot binary by executing the following command from the root of the project.
./2dhp-plot "HHHHHHHHHHHHPHPHPPHHPPHHPPHPPHHPPHHPPHPPHHPPHHPPHPHPHHHHHHHHHHHH" "RLLSLRRLLRSRRLRLRRLLRRLLRRLRRLLRRLLRRLRRLLRRLLRRLRLRRSRLSSSLLSS" ./example_figure.bmp