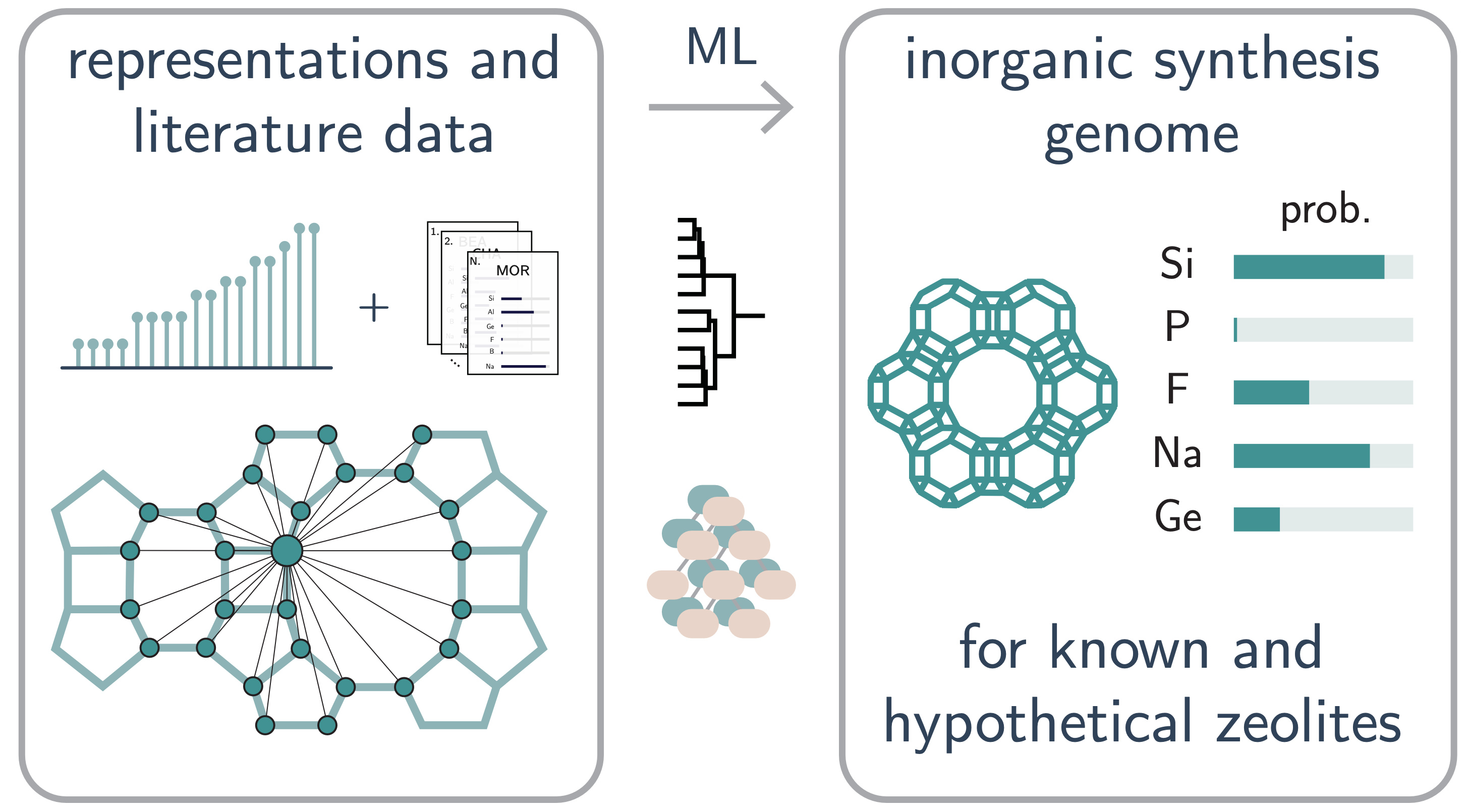
This repository contains all the data, plots, scripts, and notebooks to reproduce the manuscript:
D. Schwalbe-Koda et al. "Inorganic synthesis-structure maps in zeolites with machine learning and crystallographic distances". arXiv:2307.10935 (2023)
All the raw data for the computational analysis is found in the data folder. The scripts used to reproduce the plots is available at the zeo_amd folder. The Jupyter Notebooks in nbs contain all the code required to reproduce the analysis and the plots.
To reproduce the results from the manuscript, first create a new Python environment using your preferred virtual environment (e.g., venv or conda).
Then, clone this repository and install it with
git clone git@github.com:dskoda/Zeolites-AMD.git
cd Zeolites-AMD
pip install -e .This should install all dependencies (see pyproject.toml) and two scripts (zamd_compare and zamd_hyperopt).
Importantly, this also installs the average-minimum-distance package that compare crystals by their AMD or PDD.
For full reproducibility, all packages used when producing the results of this work are given in the environment.txt file.
To download the raw data that has all the results for this paper, simply run
chmod +x download.sh
./download.shin the root of the repository. While some of the data is already available in the repository, most of the raw data is too large for GitHub. Thus, part of the raw data that reproduces the paper is hosted on Zenodo for persistent storage (DOI: 10.5281/zenodo.8422372).
The raw data folder contains all results shown in the paper, including:
- Tabulated data for all losses, hyperparameters, and datasets analyzed in the manuscript.
- Distance matrices calculated with AMD for all known and hypothetical zeolites.
- Predictions of inorganic synthesis conditions for hypothetical zeolites.
- Synthesis data used to perform regression of the classifiers.
- Datasets of hypothetical and known zeolites.
- Final XGBoost models used in the work.
- All figures plotted in this work
For more details, see the data and dsets folders.
The Jupyter notebooks at nbs contain all information needed to reproduce the analysis of the manuscript.
Each notebook performs part of the analysis and replots the figures from the paper figs.
The code in zeo_amd simplifies the analysis in the notebooks by bundling relevant functions and scripts in the zeo_amd package.
They contain scripts that perform, for this specific work:
- Calculation of the distance matrix using the
amdcode - Plotting of the minimum spanning tree
- Clustering of the zeolite data
- Training and selection of classifiers
- Analysis of the results from hyperparameter optimization
This data has been produced for the following paper:
@article{SchwalbeKoda2023Inorganic,
title = {Inorganic synthesis-structure maps in zeolites with machine learning and crystallographic distances},
author = {Schwalbe-Koda, Daniel, Widdowson, Daniel E., Pham, Tuan Anh, Kurlin, Vitaliy E.},
year = {2023},
journal = {arXiv:2307.10935},
doi = {10.48550/arXiv.2307.10935},
url = {https://doi.org/10.48550/arXiv.2307.10935},
arxiv = {2307.10935},
}The data and all the content from this repository is distributed under the Creative Commons Attribution 4.0 (CC-BY 4.0).
This work was produced under the auspices of the U.S. Department of Energy by Lawrence Livermore National Laboratory under Contract DE-AC52-07NA27344.
Dataset released as: LLNL-MI-854709.