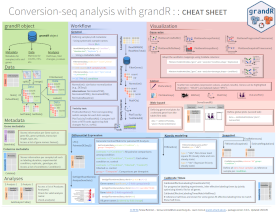
Nucleotide conversion sequencing experiments have been developed to add a temporal dimension to RNA-seq and single-cell RNA seq. Such experiments require specialized tools for primary processing such as GRAND-SLAM, and specialized tools for downstream analyses. grandR provides a comprehensive toolbox for quality control, kinetic modeling, differential gene expression analysis and visualization of such data.
grandR is available from CRAN. Install grandR using the following commands on the R console:
install.packages("grandR")
library(grandR)
You can also install the development version from github:
require("devtools")
devtools::install_github("erhard-lab/grandR")
library(grandR)
grandR should be compatible with Windows, Mac, and Linux operating systems, but we recommend using grandR on a Linux machine, where it has been extensively tested (Ubuntu 22.04). Due to restrictions of the parallel package, parallelization (SetParallel()) does not work under Windows. grandR runs on standard laptops (multi-core CPUs are recommended and memory requirements depend on the size of your data sets).
Installing it via install.packages or devtools::install_github will make sure that the following (standard) packages are available:
stats,Matrix,rlang,ggplot2,grDevices,patchwork,RCurl,plyr,parallel,reshape2,MASS,scales,cowplot,minpack.lm,lfc,labeling,methods,utils,numDeriv
Additional packages are optional and important for particular functions:
knitr, rmarkdown, circlize, Seurat, ComplexHeatmap, ggrepel, DESeq2, S4Vectors, data.table, clusterProfiler, biomaRt, msigdbr, fgsea, rclipboard, cubature, DT, RColorBrewer, gsl, htmltools, matrixStats, monocle, VGAM, quantreg, graphics, shiny, ggrastr, viridisLite
With all dependencies available, installation of grandR typically takes less than a minute.
First have a look at the Getting started vignette.
Then, go through the Differential expression or the Kinetic modeling vignette, which provide a comprehensive walk-through of the two main settings of nucleotide conversion experiments.
There are also additional vignettes:
- Loading data and working with grandR objects: Learn more about programming with grandR
- Working with data matrices and analysis results: Learn more about how to retrieve data from grandR objects
- Plotting: Learn about the plotting helper functions and the shiny web-interface of grandR
- Pulse-chase: Learn how to fit pulse-chase data with grandR
- Single cell: Learn how to load and process single cell data with grandR
- 4sU dropout: Learn how to perform analysis of 4sU dropout