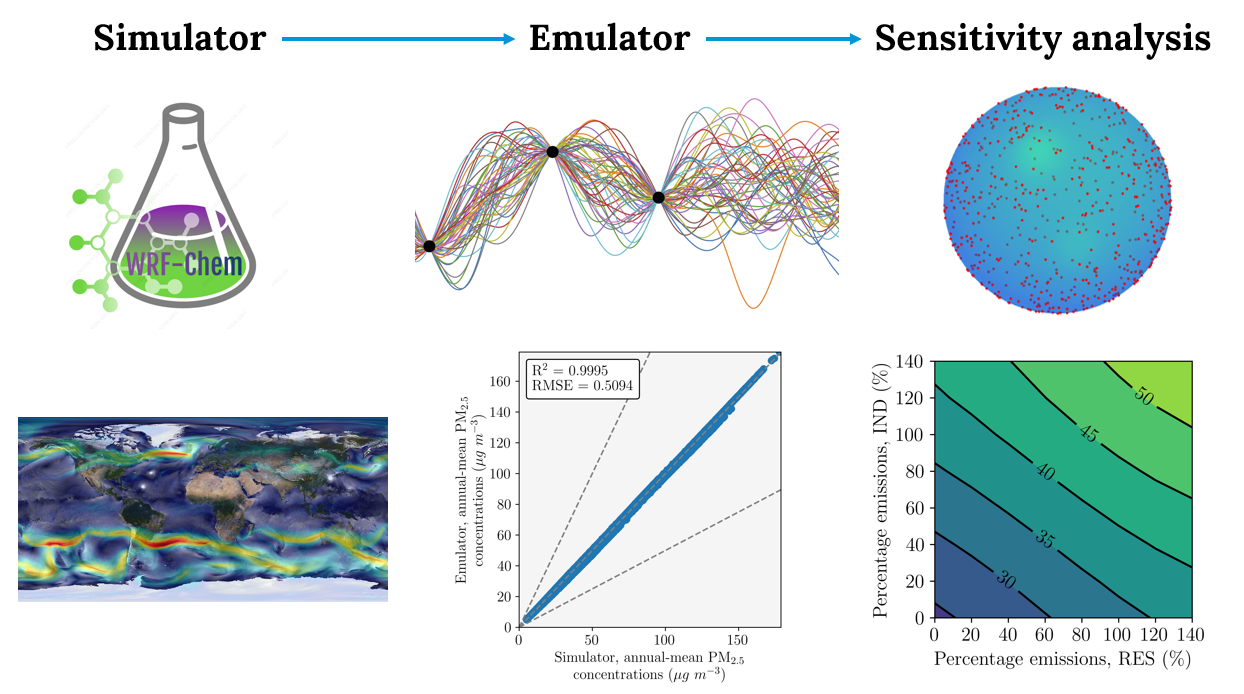
- For other scripts designing the emulator see other repository.
- Concatenate simulator data for the year (
concat_simulation_data.ipynb). - Verification plots of monthly simulation data (
check_simulation_data.ipynb). - Create ozone seasonal metric from simulator runs (
create_o3_metric.py). Submitted to HPC (create_o3_metric.bash) using Dask for workers viewing worker status on Jupyter Lab. - Create ozone seasonal metric for measurements (
create_o3_metric_measurements.pyandcreate_o3_metric_measurements.bash). - Create emulator input dictionaries (
create_emulator_inputs_outputs_df_crop). - Emulator cross-validation and sensitivity analysis (
emulator_creation.ipynb). Interactively computed on a HPC using Dask and Jupyter Lab following instructions here. - Emulator predictions for custom inputs (
emulator_predictions.py). Submitted to HPC (emulator_predictions.bash) using Dask for workers viewing worker status on Jupyter Lab. Can submit in batch mode (emulator_predictions_batch.bash). - Regrid custom outputs to population grid of the world (
regrid_to_popgrid.py). Submitted to HPC (regrid_to_popgrid.bash) using Dask for workers viewing worker status on Jupyter Lab. Can submit in batch mode (regrid_to_popgrid_batch.bash).- For PM2.5, may need
adjust_for_double_emissions.ipynband the variantadjusted_for_double_emissions_and_regrid_to_popgrid.pyinstead.
- For PM2.5, may need
- Create scaling metrics of evaluation against measurements (
wrfchem_evaluation_scaling.ipynb). - Scale predictions to measurements (
scale_to_measurements.py). Submitted to HPC (scale_to_measurements.bash) using Dask for workers viewing worker status on Jupyter Lab. Can submit in batch mode (scale_to_measurements.bash). - Crop population-weighted output predictions to region's shapefile (
popweighted_region.py). Submitted to HPC (popweighted_region.bash) using Dask for workers viewing worker status on Jupyter Lab. Uses cropping functions (cutshapefile.py). - Create shapefile clips for the each country, province, and prefecture used in the health impact assessment (
create_shapefile_clips.ipynb). - Long-term health impact assessment per configuration (
health_impacts_per_emission_configuration.py). Submitted to HPC (health_impacts_per_emission_configuration.bash) using Dask for workers viewing worker status on Jupyter Lab. Can submit in batch mode (health_impacts_per_emission_configuration.bash). - Bottom-up matching of emission configurations that match recent air quality trends (
find_emissions_that_caused_air_quality_change.pyandfind_emissions_that_caused_air_quality_change.bash). Can submit in batch mode (find_emissions_that_caused_air_quality_change_batch.bash). - Various emulator plots including emulator evaluation, sensitivity maps, prediction maps, and 2D contour pairs, (
emulator_plots.ipynbandfind_emissions_that_caused_air_quality_change.ipynb). - Plots for emissions (
emission_plots.ipynb).
- Create a conda environment with the required libraries from the config file (.yml) in the repository:
conda env create --name pangeo --file=pangeo_latest.yml
pip install salib dask_labextension pyarrow
jupyter labextension install dask-labextension
jupyter labextension install @jupyter-widgets/jupyterlab-manager
This code is currently licensed under the GPLv3 License, free of charge for non-commercial use.