This repository contains the R package now hosted on
Bioconductor
and our stable and development GitHub versions.
(macOS users only: Ensure you have installed XQuartz first.)
Make sure you have the latest R version and the latest BiocManager
package installed following these
instructions (if you use legacy
R versions (<=3.5.0) refer to the instructions at the end of the
mentioned page).
## install BiocManager if not installed
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")Ensure the following returns TRUE, or follow the guidelines provided
by the output.
BiocManager::valid()For installation in R, see options a) and b). For Docker containers, see c).
You can then install mixOmics using the following code:
## install mixOmics
BiocManager::install('mixOmics')Install the latest stable version (see below for latest
development
version) of mixOmics from GitHub (as bug-free as it can be):
BiocManager::install("mixOmicsTeam/mixOmics") Check after installation that the following code does not throw any error (especially Mac users - refer to installation instructions) and that the welcome message confirms you have installed the latest version:
library(mixOmics)
#> Loaded mixOmics ?.?.?You can also install the development version for new features yet to be widely tested (see What’s New):
BiocManager::install("mixOmicsTeam/mixOmics@devel")Click to expand
Note: this requires root privileges
- Install Docker following instructions at https://docs.docker.com/docker-for-mac/install/
if your OS is not compatible with the latest version download an older version of Docker from the following link:
- MacOS: https://docs.docker.com/docker-for-mac/release-notes/
- Windows: https://docs.docker.com/docker-for-windows/release-notes/
Then open your system’s command line interface (e.g. Terminal for MacOS and Command Promot for Windows) for the following steps.
MacOS users only: you will need to launch Docker Desktop to activate your root privileges before running any docker commands from the command line.
- Pull mixOmics container
docker pull mixomicsteam/mixomics- Ensure it is installed
The following command lists the running images:
docker imagesThis lists the installed images. The output should be something similar to the following:
$ docker images
> REPOSITORY TAG IMAGE ID CREATED SIZE
> mixomicsteam/mixomics latest e755393ac247 2 weeks ago 4.38GB
- Active the container
Running the following command activates the container. You must change
your_password to a custom password of your own. You can also customise
ports (8787:8787) if desired/necessary. see
https://docs.docker.com/config/containers/container-networking/ for
details.
docker run -e PASSWORD=your_password --rm -p 8787:8787 mixomicsteam/mixomics- Run
In your web browser, go to http://localhost:8787/ (change port if
necessary) and login with the following credentials:
username: rstudio
password: (your_password set in step 4)
- Inspect/stop
The following command lists the running containers:
sudo docker psThe output should be something similar to the following:
$ sudo docker ps
> CONTAINER ID IMAGE COMMAND CREATED STATUS PORTS NAMES
> f14b0bc28326 mixomicsteam/mixomics "/init" 7 minutes ago Up 7 minutes 0.0.0.0:8787->8787/tcp compassionate_mestorfThe listed image ID can then be used to stop the container (here
f14b0bc28326)
docker stop f14b0bc28326We welcome community contributions concordant with our code of
conduct.
We strongly recommend adhering to Bioconductor’s coding
guide for
software consistency if you wish to contribute to mixOmics R codes.
To report a bug (or offer a solution for a bug!) visit: https://github.com/mixOmicsTeam/mixOmics/issues. We fully welcome and appreciate well-formatted and detailed pull requests. Preferably with tests on our datasets.
Set up development environment
- Install the latest version of R
- Install RStudio
- Clone this repo, checkout master branch, pull origin and then run:
install.packages("renv", Ncpus=4)
install.packages("devtools", Ncpus=4)
# restore the renv environment
renv::restore()
# or to initialise renv
# renv::init(bioconductor = TRUE)
# update the renv environment if needed
# renv::snapshot()
# test installation
devtools::install()
devtools::test()
# complete package check (takes a while)
devtools::check()We wish to make our discussions transparent so please direct your analysis questions to our discussion forum https://mixomics-users.discourse.group. This forum is aimed to host discussions on choices of multivariate analyses, as well as comments and suggestions to improve the package. We hope to create an active community of users, data analysts, developers and R programmers alike! Thank you!
mixOmics is collaborative project between Australia (Melbourne),
France (Toulouse), and Canada (Vancouver). The core team includes
Kim-Anh Lê Cao - https://lecao-lab.science.unimelb.edu.au (University
of Melbourne), Florian Rohart - http://florian.rohart.free.fr
(Toulouse) and Sébastien Déjean -
https://perso.math.univ-toulouse.fr/dejean/. We also have key
contributors, past (Benoît Gautier, François Bartolo) and present (Al
Abadi, University of Melbourne) and several collaborators including
Amrit Singh (University of British Columbia), Olivier Chapleur (IRSTEA,
Paris), Antoine Bodein (Universite de Laval) - it could be you too, if
you wish to be involved!.
The project started at the Institut de Mathématiques de Toulouse in
France, and has been fully implemented in Australia, at the University
of Queensland, Brisbane (2009 – 2016) and at the University of
Melbourne, Australia (from 2017). We focus on the development of
computational and statistical methods for biological data integration
and their implementation in mixOmics.
mixOmics offers a wide range of novel multivariate methods for the
exploration and integration of biological datasets with a particular
focus on variable selection. Single ’omics analysis does not provide
enough information to give a deep understanding of a biological system,
but we can obtain a more holistic view of a system by combining multiple
’omics analyses. Our mixOmics R package proposes a whole range of
multivariate methods that we developed and validated on many biological
studies to gain more insight into ’omics biological studies.
www.mixOmics.org (tutorials and resources)
Our latest bookdown vignette: https://mixomicsteam.github.io/Bookdown/.
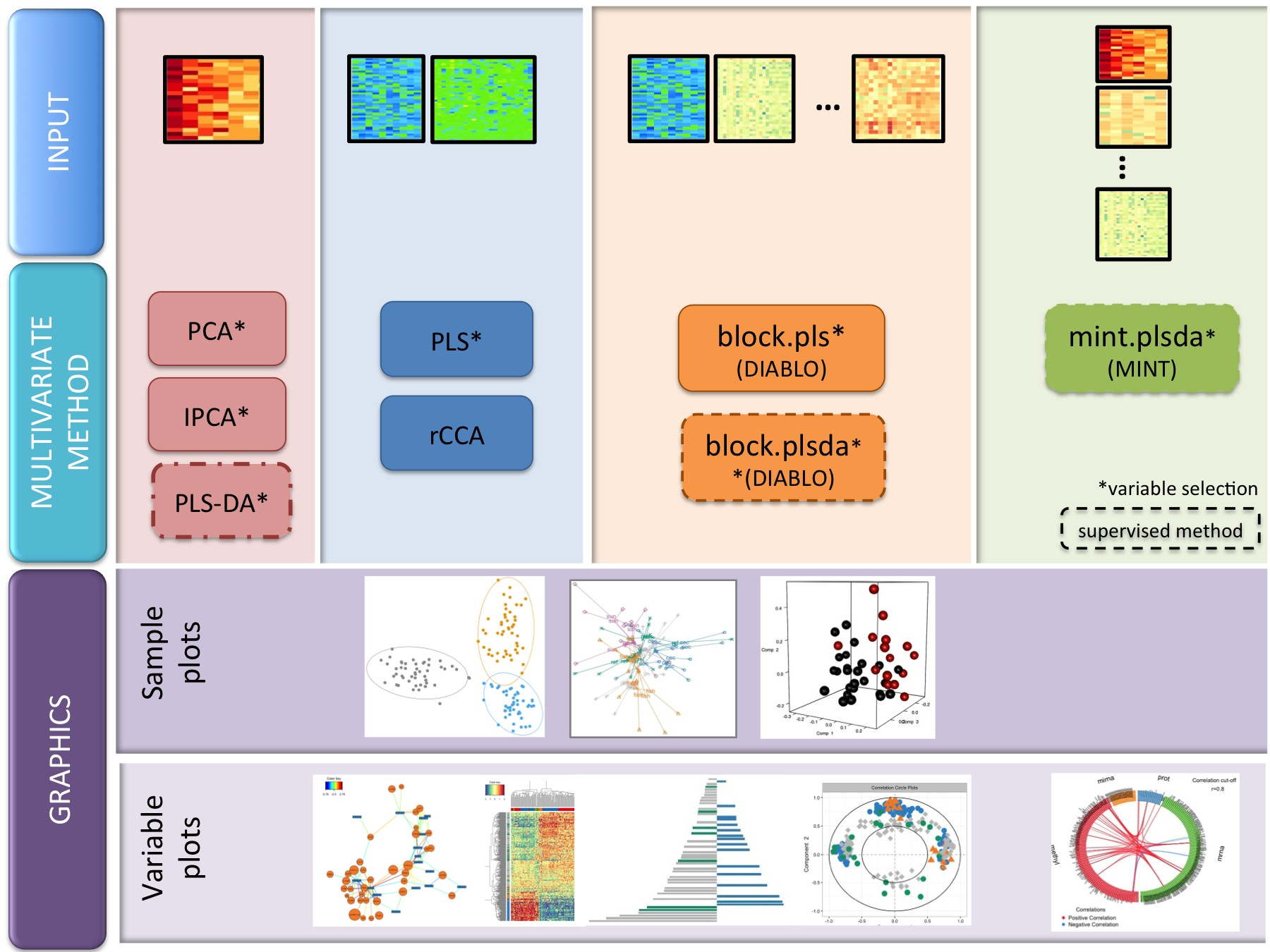
We have developed 17 novel multivariate methods (the package includes 19 methods in total). The names are full of acronyms, but are represented in this diagram. PLS stands for Projection to Latent Structures (also called Partial Least Squares, but not our preferred nomenclature), CCA for Canonical Correlation Analysis.
That’s it! Ready! Set! Go!
Thank you for using mixOmics!
- bug fix implemented for Issue
#196.
perf()can now handle features with a(s)plswhich have near zero variance. - bug fix implemented for Issue
#192.
predict()can now handle when the testing and training data have their columns in different orders. - bug fix implemented for Issue
#178. If the
indYparameter is used inblock.spls(),circosPlot()can now properly identify thedataframe.
- bug fix implemented for Issue
#172.
perf()now returns values for thechoice.ncompcomponent whennrepeatwhereas before it would just return
NAs. - bug fix implemented for Issue
#171.
cim()now can takepcaobjects as input. - bug fix implemented for Issue
#161.
tune.spca()can now handleNAvalues appropriately. - bug fix implemented for Issue
#150.
Provided users with a specific error message for when
plotArrow()is run on a(mint).(s)plsdaobject. - bug fix implemented for Issue
#122.
Provided users with a specific error message for when a
splsdaobject that has only one sample associated with a given class is passed toperf(). - bug fix implemented for Issue
#120.
plotLoadings()now returns the loading values for features from all dataframes rather than just the last one when operating on a(mint).(block).(s)plsdaobject. - bug fix implemented for Issue
#43.
Homogenised the way in which
tune.mint.splsda()andperf.mint.splsda()calculate balanced error rate (BER) as there was disparity between them. Also made the global BER a weighted average of BERs across each study. - enhancement implemented for Issue
#30/#34. The
parameter
verbose.callwas added to most of the methods. This parameter allows users to access the specific values input into the call of a function from its output. - bug fix implemented for Issue
#24.
background.predict()can now operate onmint.splsdaobjects and can be used as part ofplotIndiv().
- new function
plotMarkersto visualise the selected features in block analyses (see #134) tune.splsnow able to tune the selected variables on bothXandY. See?tune.spls- new function
impute.nipalsto impute missing values using the nipals algorithm - new function
tune.spcato tune the number of selected variables for pca components circosPlotnow has methods forblock.splsobjects. It can now handle similar feature names across blocks. It is also much more customisable. See advanced arguments in?circosPlot- new
biplotfunction forpcaandplsobjects. See?mixOmics::biplot plotDiablonow takescol.per.group(see #119)
- weighted consensus plots for DIABLO objects now consider per-component weights
plotIndivnow supports (weighted) consensus plots for block analyses. See the example in this issueplotIndiv(..., ind.names=FALSE)warning issue now fixed
perf.block.splsdanow supports calculation of combined AUCblock.splsdabug which could drop some classes withnear.zero.variance=TRUEnow fixed


-lightgrey.svg)