Alan is a simple, light command line alignment viewer, for viewing amino acid and DNA alignments in a linux terminal.
It can be used for viewing FASTA or Clustal format alignment files, and relies on just a few standard unix tools (less, awk, sed, grep among others).
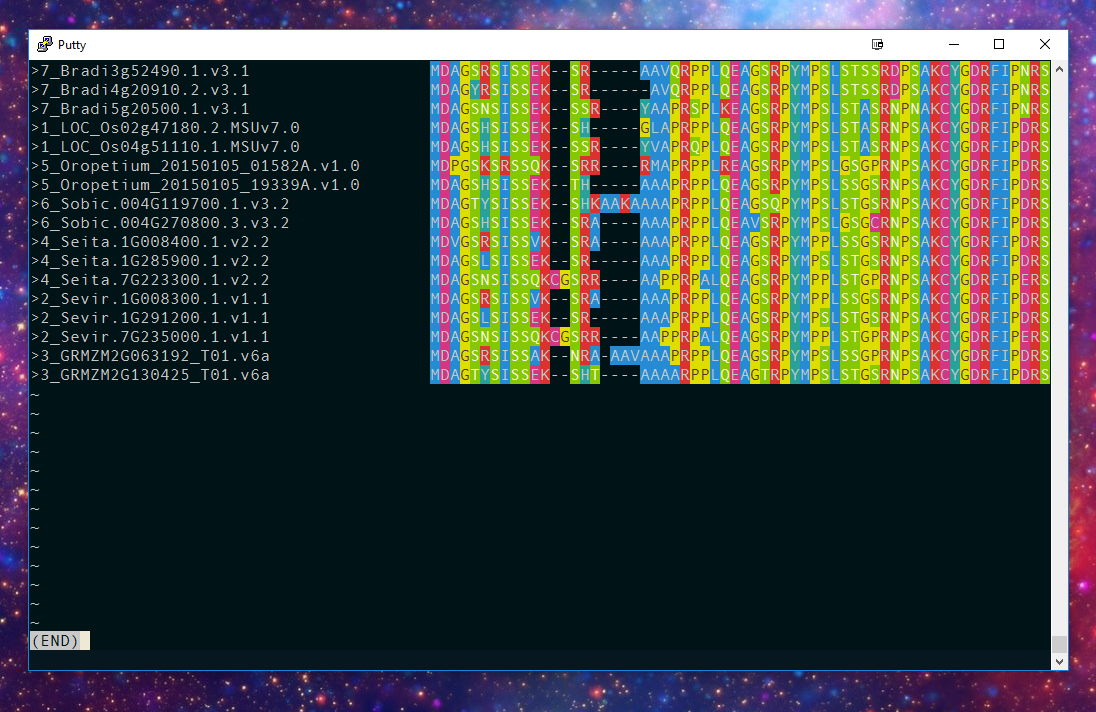
Here's an amino acid sequence alignment, viewed using Alan:
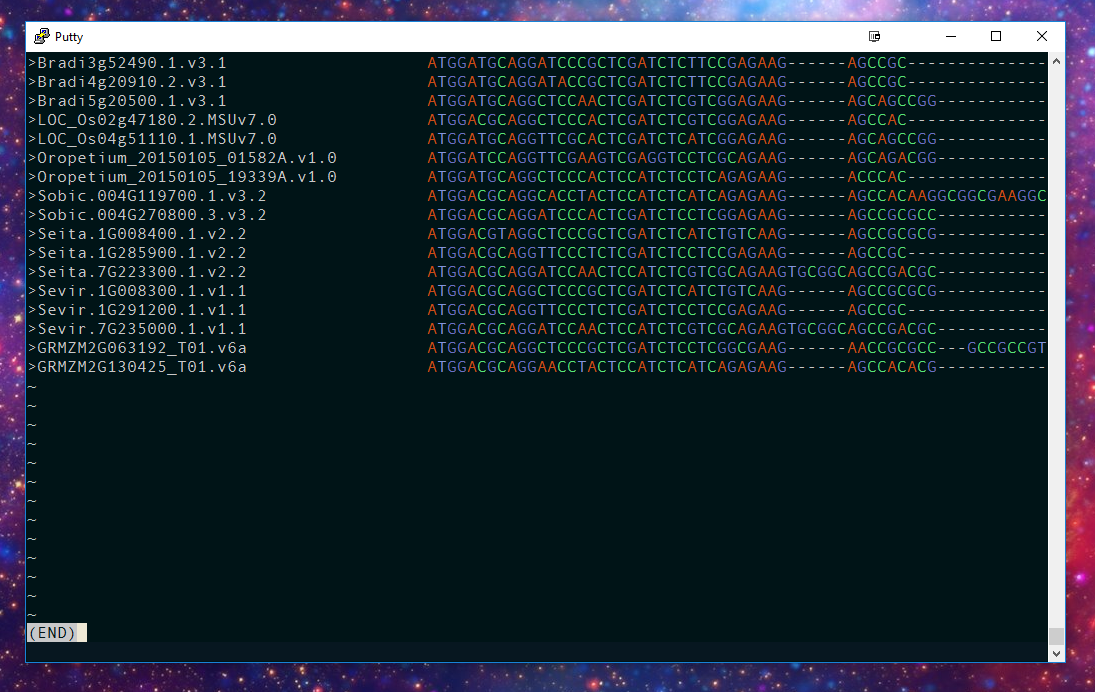
A DNA sequence alignment:
The alan command should be ready to use straight out of the box - no installation is required. Just type:
/path/to/alan your_alignment.faWhere /path/to/alan is wherever you've saved the alan file in this repo.
To make things even easier, you can "install" alan by adding the directory containing the alan executable to your $PATH, using:
# Ubuntu/etc:
echo 'PATH=$PATH:/path/to/alan_dir' > ~/.bashrc
# Use this one if you're on mac (probably):
#echo 'PATH=$PATH:/path/to/alan_dir' > ~/.zshrc
source ~/.bashrc
If you've done this, Alan can be run by simply typing alan your_alignment.fa.
Alan works on FASTA and Clustal format alignments. Any other alignment formats will confuse it. The choice of file format (FASTA or Clustal) is detected automatically.
If no information on molecule type is supplied, Alan will try to detect whether an alignment is composed of protein or nucleotide sequences, and will display them accordingly. To override this, use the options -n for nucleotides, or -p for protein sequences.
To navigate the alignment, use the keyboard arrow keys. The amount by which an alignment is shifted using the arrow keys can be set using the -s or --shift options, followed by a numeric value. The default is 10.
Alan uses less as its main viewer, and so any other in-viewer commands contained in less will also work in Alan. This includes searching (use /), and more: see https://en.wikipedia.org/wiki/Less_(Unix) for a few.
Alan sometimes struggles with larger alignments.
By default, Alan uses the clustal colour scheme as described here. The formatting is not context-dependent, and so always displays one colour per character. The formatting is designed to work on a black-screen terminal.
You can update the colour scheme used by Alan in two ways. Alan colour schemes are defined by colour files. The default one for protein alignments looks like this:
- 0;0
KkRr 01;41
SsTtQqNn 01;37;42
AaIiLlMmFfWwVvCc 01;44
Cc 01;105
EeDd 01;45
Gg 01;90;103
HhYy 01;46
Pp 01;90;43
The default one for nucleotide alignments looks like this:
-Nn 0;0
Aa 01;31
Gg 01;33
Cc 01;32
Tt 01;34
These are tab-delimited text files, and are embedded in the alan script itself. The first column defines the symbols to be coloured, the second defines the formatting and colouring using the codes from here. Different formatting instructions are separated by ;, so that for example 01;90;103 means 01 (bold) and 90 (dark grey text) and 103 (light yellow background). Finally, the colour file must not contain blank lines, and any instance of "-" must be at the start of a line.
To include a custom colour file, include a colp.csv or coln.csv file (for protein and nucleotides respectively) in the same directory as the alan file, the contents of which should be in the same format as the above. Alternatively, if you wish to use multiple colour schemes for different purposes, you may wish to explicitly refer to a colour scheme file using the --colp or --coln options. For example:
alan my_alignment.fa --colp my_colours.csv.
The original Alan command was a one-line modification of the less command. For its simplicity and portability, you may wish to use the orignal alan on your system. This and some derived commands are contained in the alan_museum directory, and they are loaded as functions, so type source alan_museum/alan_orig.sh to load them, and then type the individual commands to use them. The original Alan commands are:
| Command Name | Command | Description | Example |
|---|---|---|---|
| Alan Bennett | alanbennett |
Fasta, automatic | alanbennett alignment.fasta |
| Alan Davies | alandavies |
Fasta, protein | alandavies prot_alignment.fasta |
| Alan Rickman | alanrickman |
Fasta, nucleotide | alanrickman nuc_alignment.fasta |
| Alan Partridge | alanpartridge |
Clustal, automatic | alanpartridge alignment.fasta |
| Alan Menken | alanmenken |
Clustal, protein | alanmenken prot_alignment.fasta |
| Alan Shearer | alanshearer |
Clustal, nucleotide | alanshearer nuc_alignment.fasta |