Author: Nicholas A. Battista, Ph.D.
Email: battistn[at]tcnj.edu
Website: http://battistn.pages.tcnj.edu
Department: Mathematics & Statistics (TCNJ MATH)
Institution: The College of New Jersey (TCNJ)
An easy to use immersed boundary method in 2D, with full implementations in MATLAB and Python that contains over 75 built-in examples, including multiple options for fiber-structure models and advection-diffusion, Boussinesq approximations, and/or artificial forcing.
If you use this software for research, educational, or recreational purposes, please let Nick Battista (battistn[at]tcnj[.]edu) know!
-
N.A. Battista, A.J. Baird, L.A. Miller, A mathematical model and MATLAB code for muscle-fluid-structure simulations, Integ. Comp. Biol. 55(5):901-911 (2015), LINK
-
N.A. Battista, W.C. Strickland, L.A. Miller, IB2d:a Python and MATLAB implementation of the immersed boundary method, Bioinspiration and Biomemetics 12(3): 036003 (2017), LINK
-
N.A. Battista, W.C. Strickland, A. Barrett, L.A. Miller, IB2d Reloaded: a more powerful Python and MATLAB implementation of the immersed boundary method, in press Math. Method. Appl. Sci. 41:8455-8480 (2018) LINK
-
Tutorial 1: https://youtu.be/PJyQA0vwbgU
An introduction to the immersed boundary method, fiber models, open source IB software, IB2d, and some FSI examples! -
Tutorial 2: https://youtu.be/jSwCKq0v84s
A tour of what comes with the IB2d software, how to download it, what Example subfolders contain and what input files are necessary to run a simulation -
Tutorial 3: https://youtu.be/I3TLpyEBXfE
An overview of how to construct immersed boundary geometries and create the input files (.vertex, .spring, etc.) for an IB2d simulation to run using the oscillating rubberband example from Tutorial 2 as a guide. -
Tutorial 4: https://youtu.be/4D4ruXbeCiQ
The basics of visualizing data using open source visualization software called VisIt (by Lawrence Livermore National Labs), visualizing the Lagrangian Points and Eulerian Data (colormaps for scalar data and vector fields for fluid velocity vectors)
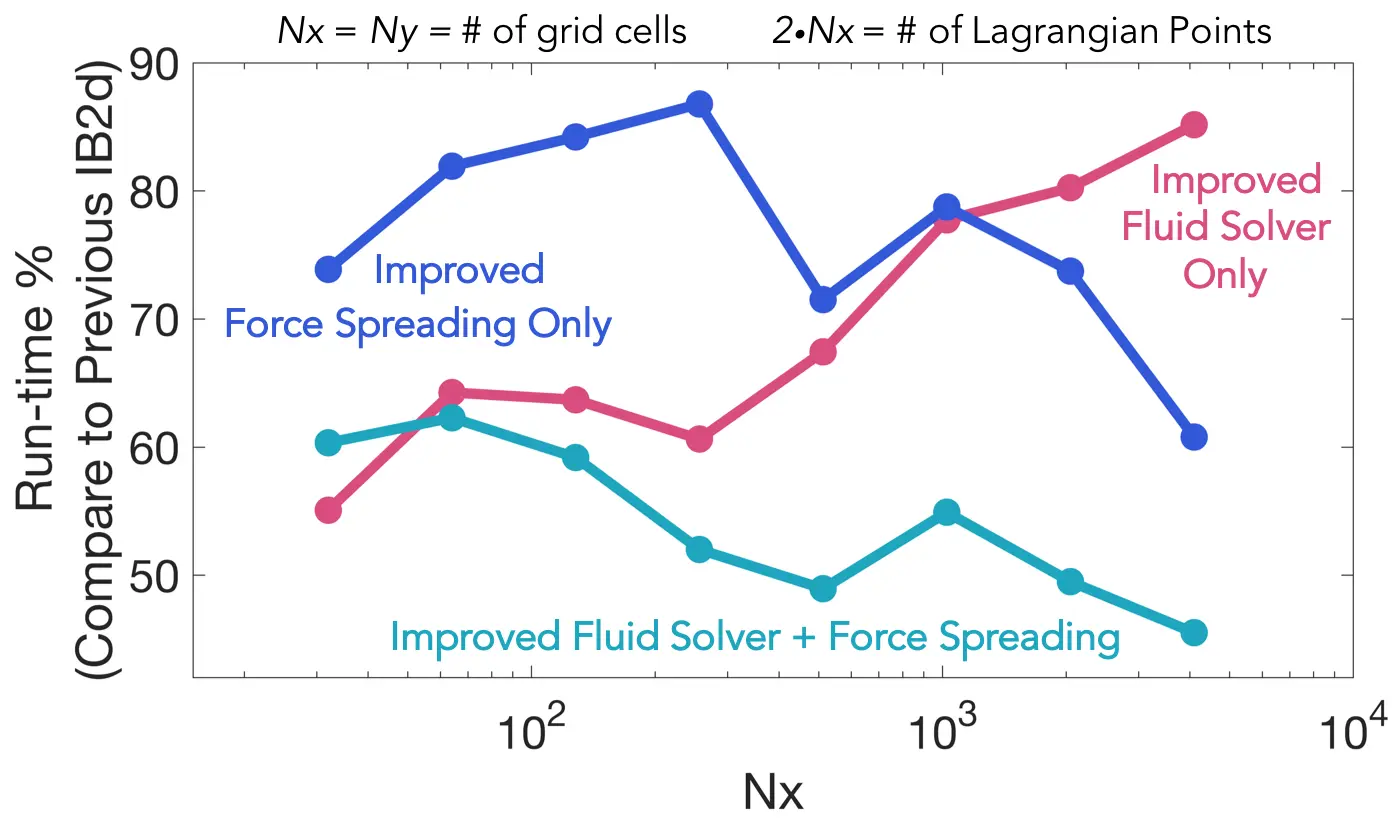
- Substantial speed-ups in performance for fluid solver and force spreading operations in MATLAB implementation. Speeds-ups in Python implementation forthcoming (February 2024):
Figure: Showing the percent runtimes when compared to IB2d's previous implementations, i.e., the
ratio of runtimes x 100. The simulation involved a square grid with an equal number of grid
cells (Nx) in the x- and y- directions. The corresponding # of Lagrangian points for each case was 2*Nx.
-
Our software release paper on the recenty added concentration dynamics infrastructure was published in Bioinspiration & Biomimetics. More information can be found at the following link:
M. Santiago, N.A. Battista, L.A. Miller, S. Khatri, Passive concentration dynamics incorporated into the library IB2d, a two-dimensional implementation of the immersed boundary method , Bioinspiration & Biomimetics 17: 036003 (2022)
-
Matea Santiago has updated the advection-diffusion solver to a third-order WENO scheme.
-
We have released a semi-automatic meshing tool, MeshmerizeMe , to help discretize Lagrangian geometries. More information can be found in our software release paper:
D.M. Senter, D.R. Douglas, W.C. Strickland, S. Thomas, A. Talkington, L.A. Miller, N.A. Battista, A Semi-Automated Finite Difference Mesh Creation Method for Use with Immersed Boundary Software IB2d and IBAMR , Bioinspiration and Biomimetics 16(1): 016008 (2021)
-
The MATLAB plotting routine is incompatible with MATLAB R2020a and R2020b. If you run into this issue, please consider visualizing the .vtk data with VisIt or ParaView instead.
--> Many VisIt users have gotten the following error when trying to open visualization files:
"The MetaData server running on localhost could not get the file list for the current directory"
--> To circumvent this error, try opening VisIt from the command line. Try the following:
- In your terminal, change directory until you're in the directory you want to open (eg - the viz_IB2d folder)
- In your terminal, type: /Applications/VisIt.app/Contents/Resources/bin/visit -gui (or the correct path based to where you installed VisIt)