New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Problems with reading some compressed FITS #349
Comments
|
After some crude experimentation, it seems the library does not implement the reversion of the floating-point data quantization described in §10.2 of the FITS standard. That is, to get an approximately expected result:
|
|
Hi @bogdanni! Thanks for getting this diagnosed! The compression staff is really not my doing, but based on your diagnosis I should be able to fix it just in time for the next update (1.17.1 around 12/15/22)... :-) |
|
@bogdanni As a shot in the dark, it looks like the decompression of GZIPed images was not using the tile options, which use the ZSCALE and ZZERO values from the compressed table. I added the options in. Let's see if that does the trick. You can try https://github.com/attipaci/nom-tam-fits/tree/issue-349 to see if that fixes it. (If not, I'll have to dive deeper...) |
|
Hi @attipaci, thanks for the rapid reaction! That change doesn't seem to fix it though... |
|
Well, I knew it was a long shot... But for sure ZSCALE and ZZERO were not used for GZIP'd images before. I'll have to learn more about what Ritchie was trying to do for compression, and hopefully track down where else the implementation is missing those. I might have more things for you to try in the next week or two... |
|
For AIJ we were looking to match LCO's lossy compression for floating point images, but ran into issues writing them, with similar banding to what is mentioned here; though it only showed in a subtraction of the original image and the compressed version. While comparing with CFITSIO's compression/decompression, we noticed that nom.tam's unpacking was giving different pixel values. Here is a test project, should print out There could be an issue with the writer as well, but its hard to tell when the reader is having issues as well. |
|
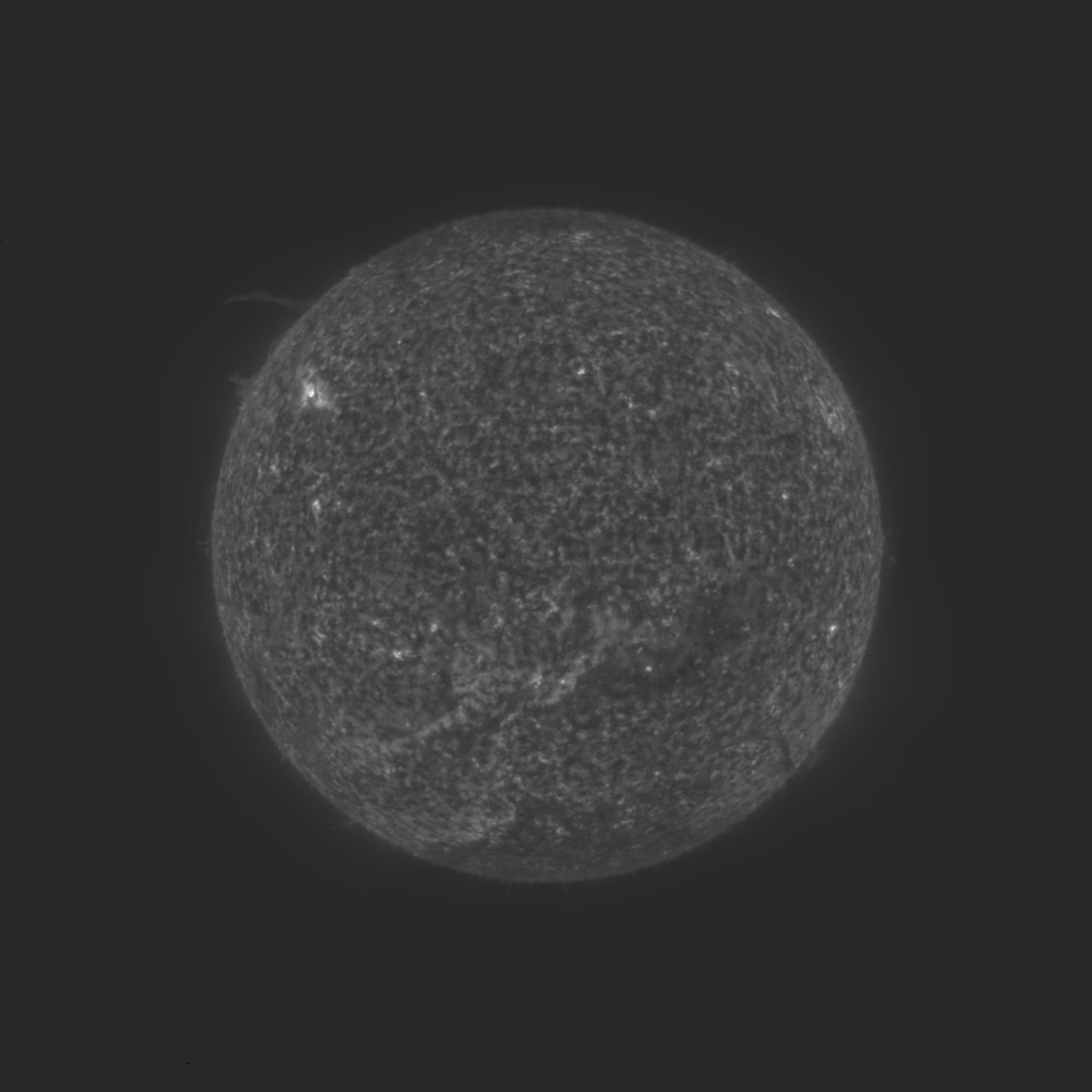
Good news at last. I have managed to finally decouple quantization from the compression so that quantization can be applied to all types of tile compression the same. It's been a nasty bug to fix, requiring many hundreds of lines to be changed over 42 source files. But, it looks like the compressed solar image decompresses fine at last, with no more visible stripes. Please check out the PR yourselves also (see #350). Once both of you are satisfied that it works as expected, I will merge it after the minor release of 1.17.1 (March 15), so that it will be in 1.18-SNAPSHOT thenceforth, and will eventually get included in the 1.18.0 release (expected around June 15). Thanks for all your help in advance. Here is that same compressed image from the top of the thread, now decompressed on the issue-349 branch (the PR): |
|
Thank you for fixing this! I confirm it looks good also in my tests. |
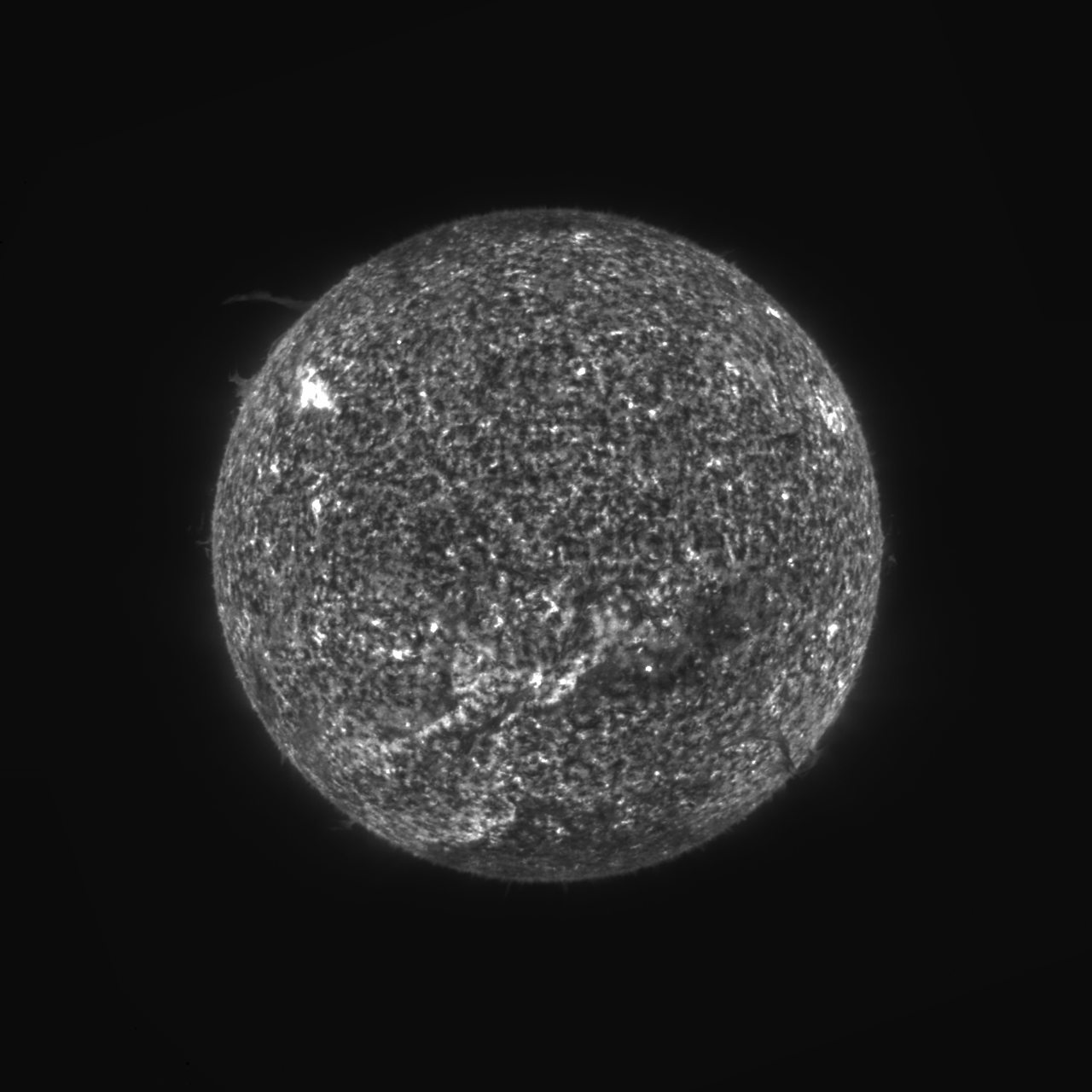
Having trouble decoding files like:
https://data.ngdc.noaa.gov/platforms/solar-space-observing-satellites/goes/goes16/l2/data/suvi-l2-ci304/2021/02/20/dr_suvi-l2-ci304_g16_s20210220T234800Z_e20210220T235200Z_v1-0-1.fits
SAOImage DS9 shows the image correctly:

Horizontal stripes like in the attached image show up when decoding the file with nom-tam-fits :
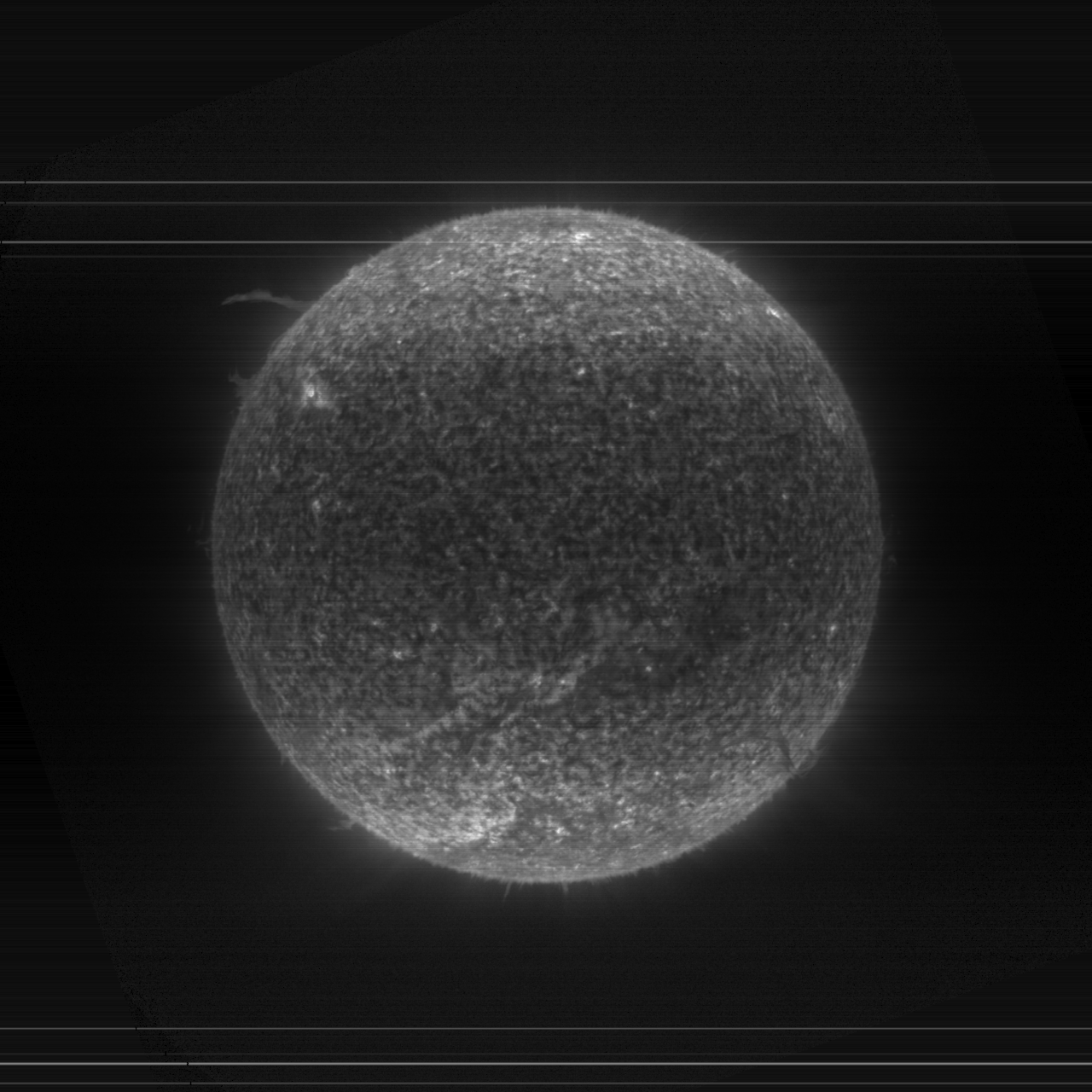
The image was produced by the code below:
The text was updated successfully, but these errors were encountered: