New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
PySAL 2.1.0 changing matplotlib backend (Jupyter) #1138
Comments
|
Could this be done through |
I don't think so, I don't specify the backend within the package, just for travis testing, but separately. And also splot has not changed much in functionality from 2.0.0 to 2.1.0. |
|
I’ll try reconstructing in two ways to see if I can narrow it down:
1. A minimal Jupyter + PySAL installation in anaconda
2. Messing about with the fuller install on which this problem appeared (will attach YAML file when back in front of my machine).
Could it be something from testing percolating through to the anaconda install (I’m using conda-forge) and those somehow becoming the default configuration?
Jon
…--
m: 07976987392
e: jon@reades.com
s: jreades
On 9 Aug 2019, 17:29 +0100, Stefanie Lumnitz ***@***.***>, wrote:
> Could this be done through splot @slumnitz ?
I don't think so, I don't specify the backend within the package, just for travis testing, but separately. And also splot has not changed much in functionality from 2.0.0 to 2.1.0.
—
You are receiving this because you authored the thread.
Reply to this email directly, view it on GitHub, or mute the thread.
|
|
I wonder how On this, it might be useful to provide a full |
|
As you know @darribas, I've been trying to switch over to Docker using your GDS image: I am seeing the same behaviour in that with my GSA (Geocomputation and Spatial Analysis) build on top of the |
|
Maybe it is a new or changed dependency that is causing the problem when importing |
|
Will do! Possibly tomorrow as 10:30pm here :-) Yes, a dependency issue seems reasonable, but wondering why my docker build on top of Dani's is triggering the error... |
|
I just did a fresh install of gds_env and I can't get passed the first cell of this example: So I'm wondering if you have some cross-environment stuff going on? Are you running this in a container? |
|
Ah sorry, I'd extended the environment with a new container as we use a few modules at King's that Dani doesn't at Liverpool; up to that point I'd assumed that it was something Mac-specific but the container is Ubuntu which is why I was posting an update. Taking @slumnitz's suggestion on board I went back to running conda directly on my Mac and created two fresh environments which differ only in having either PySAL 2.0.0 or 'latest' (i.e. 2.1.0) specified in the YAML file. The same issue recurs so the difference must be in either PySAL or one of the dependencies that changes between 2.0.0 and 2.1.0. I've diffed the output of Outputs from |
|
@sjsrey what is the |
|
OK, I think the https://github.com/matplotlib/jupyter-matplotlib A quick check confirms this: Is this something people use commonly? If so I could install in on the next version... |
|
@darribas that’s my code/fault, not @sjsrey's as it was stuff that had percolated through the work I was doing to modify your GDS container for use at King's — it gives you back interactive plots in jupyterlab if you install I’m inclined towards the dependency problem as being the most likely source of the issue, but the Of the libraries that I found to be different between installing PySAL 2.0.0 and 2.1.0, the only one that makes explicit reference to matplotlib (that I could find) was
Bizarrely, one of the developers of this package is @casperdcl who is currently working with @darribas and me on a cool Docker project! If you agree that my investigation has turned up the likely ‘culprit’ then I’ll get in touch with Casper. |
|
Given that you just @ mentioned me, you're already in touch :D |
|
I still can't replicate this with either development head, the For instance, using the source from pypi, this notebook (with # In[1]:
import matplotlib
# %matplotlib inline
matplotlib.get_backend()
# In[2]:
import pysal
pysal.__file__
# In[3]:
matplotlib.get_backend()
# In[4]:
from pysal import explore
# In[5]:
matplotlib.get_backend() |
|
Like, by distribute the docker image, I mean freeze & publish it in the broken state so we can check it out. |
|
P.S. @jreades in the unlikely event that you guys conclude this is an issue with |
|
I've got @darribas in town from the end of next week -- perhaps we can investigate further then... |
|
I'm 99% sure this is happening because of |



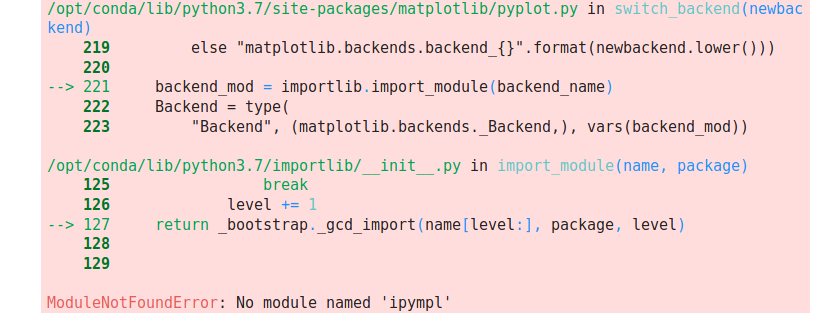


I've encountered an issue with PySAL 2.1.0 (possibly
condaspecific?) in which importing pysal into a Jupyter notebook changes the matplotlib rendering back-end from whatever I specify (TkAgg) toagg, which doesn't work and yields the error:The attached screenshot shows this behaviour in action in a notebook (I've also attached the notebook for testing purposes, though you'll need to change the extension back to
.ipynb).I've confirmed that this issue is specific to 2.1.0 by downgrading to 2.0.0 and the issue disappears (see second attached screenshot). Perhaps this is something specific to Jupyter/JupyterLab but the fact that downgrading PySAL makes the problem go away leaves me wondering...
jon
P.S. Can supply a full Conda YAML file if necessary, but this should be easily reproducible if it's actually something specific to PySAL 2.1.0.
PySAL Test.txt
The text was updated successfully, but these errors were encountered: