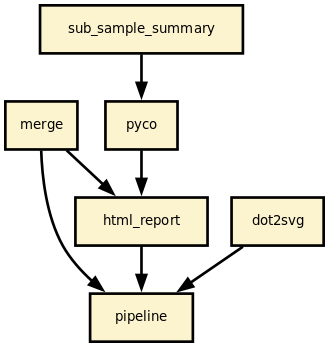
This is is the nanomerge pipeline from the Sequana project
- Overview
merge fastq files generated by Nanopore run and generates raw data QC.
- Input
individual fastq files generated by nanopore demultiplexing
- Output
merged fastq files for each barcode (or unique sample)
- Status
production
- Citation
Cokelaer et al, (2017), ‘Sequana’: a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI doi:10.21105/joss.00352
You can install the packages using pip:
pip install sequana_nanomerge --upgradeAn optional requirements is pycoQC, which can be install with conda/mamba using e.g.:
conda install pycoQCyou will also need graphviz installed.
sequana_nanomerge --helpIf you data is barcoded, they are usually in sub-directories barcoded/barcodeXY so you will need to use a pattern (--input-pattern) such as `/.gz`:
sequana_nanomerge --input-directory DATAPATH/barcoded --samplesheet samplesheet.csv
--summary summary.txt --input-pattern '*/*fastq.gz'otherwise all fastq files are in DATAPATH/ so the input pattern can just be `*.fastq.gz`:
sequana_nanomerge --input-directory DATAPATH --samplesheet samplesheet.csv
--summary summary.txt --input-pattern '*fastq.gz'The --summary is optional and takes as input the output of albacore/guppy demultiplexing. usually a file called sequencing_summary.txt
Note that the different between the two is the extra */ before the *.fastq.gz pattern since barcoded files are in individual subdirectories.
In both bases, the command creates a directory with the pipeline and configuration file. You will then need to execute the pipeline:
cd nanomerge
sh nanomerge.sh # for a local runThis launch a snakemake pipeline. If you are familiar with snakemake, you can retrieve the pipeline itself and its configuration files and then execute the pipeline yourself with specific parameters:
snakemake -s nanomerge.rules -c config.yaml --cores 4 --stats stats.txtOr use sequanix interface.
Concerning the sample sheet, whether your data is barcoded or not, it should be a CSV file :
barcode,project,sample
barcode01,main,A
barcode02,main,B
barcode03,main,CFor a non-barcoded run, you must provide a file where the barcode column can be set (empty):
barcode,project,sample
,main,Aor just removed:
project,sample
main,AWith apptainer, initiate the working directory as follows:
sequana_nanomerge --use-apptainerImages are downloaded in the working directory but you can store then in a directory globally (e.g.):
sequana_nanomerge --use-apptainer --apptainer-prefix ~/.sequana/apptainersand then:
cd nanomerge
sh nanomerge.shif you decide to use snakemake manually, do not forget to add apptainer options:
snakemake -s nanomerge.rules -c config.yaml --cores 4 --stats stats.txt --use-apptainer --apptainer-prefix ~/.sequana/apptainers --apptainer-args "-B /home:/home"This pipelines requires the following executable(s), which is optional:
- pycoQC
- dot
This pipeline runs nanomerge in parallel on the input fastq files (paired or not). A brief sequana summary report is also produced.
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
| Version | Description |
|---|---|
| 1.5.1 | * Fix wrappers tag |
| 1.5.0 | * refactoring to use Click |
1.4.0 |
|
1.3.0 |
|
1.2.0 |
|
1.1.0 |
|
1.0.1 |
|
| 1.0.0 | Stable release ready for production |
| 0.0.1 | First release. |