-
Notifications
You must be signed in to change notification settings - Fork 0
Commit
This commit does not belong to any branch on this repository, and may belong to a fork outside of the repository.
- Loading branch information
1 parent
c421566
commit c3e28f0
Showing
1 changed file
with
58 additions
and
3 deletions.
There are no files selected for viewing
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -1,9 +1,64 @@ | ||
| # GeneMap | ||
|
|
||
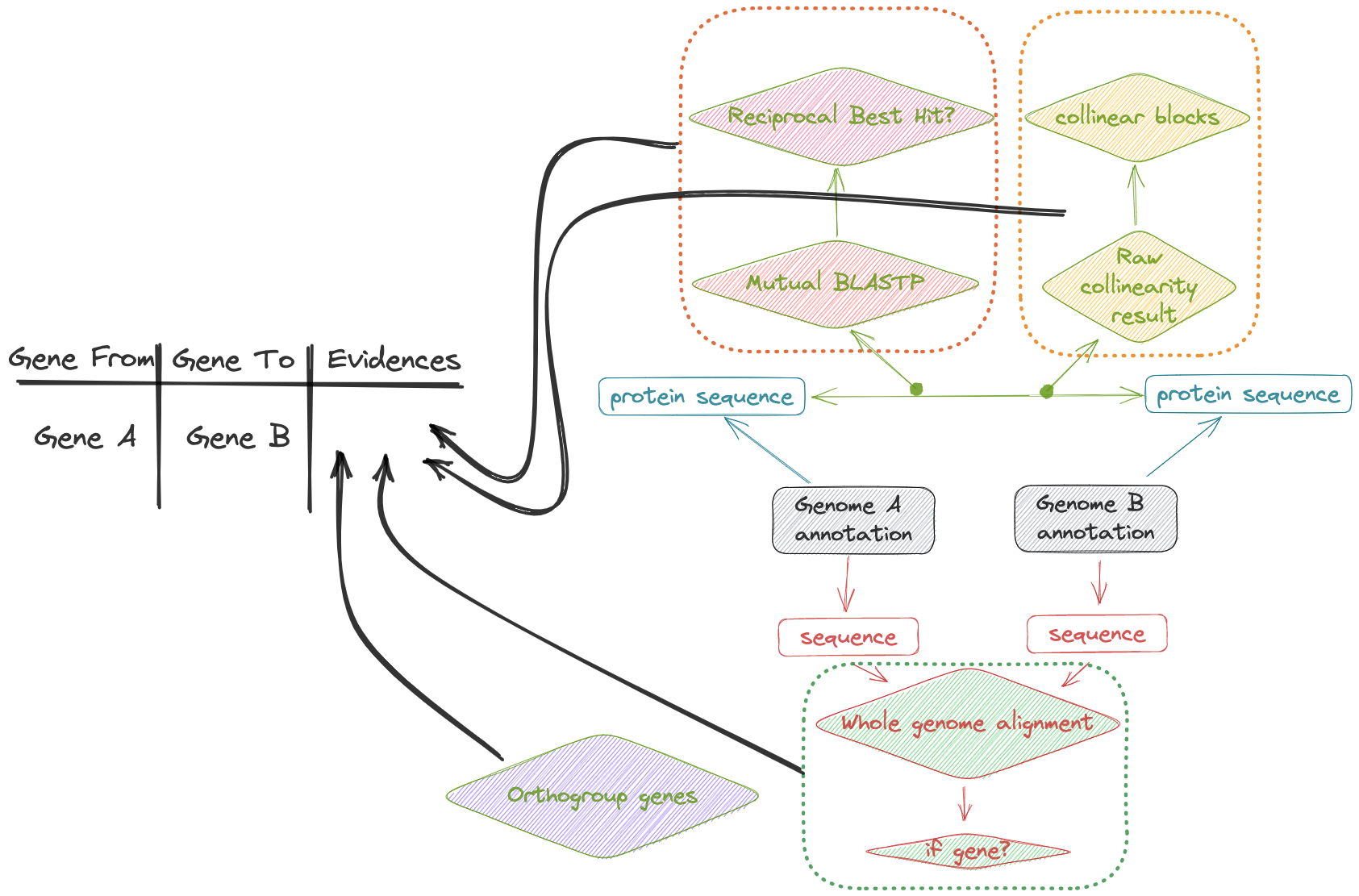
| GeneMap, A python tool for identifying the mapping of genes between two genomes by multi evidences. | ||
|  | ||
|
|
||
| ## Table of Contents | ||
|
|
||
| - [Background](#background) | ||
| - [Install](#install) | ||
| - [Usage](#usage) | ||
| - [Badge](#badge) | ||
| - [Maintainers](#maintainers) | ||
| - [Contributing](#contributing) | ||
| - [License](#license) | ||
|
|
||
| ## Background | ||
|
|
||
| When we have multiple genomes of the same species (or related species), we usually need to identify the gene mapping relationship between the two genomes(Gene A in genome a, mapped to gene B in genome B). We used four alternative evidence, including: | ||
|
|
||
| 1. RBH | ||
| **R**eciprocal **B**est BLAST **H**its | ||
|
|
||
| > Despite these and other complication, the identification of reciprocal best hits for gene products is a good first approximation to the identification of orthologues in two or more organisms. It forms the basis for many orthology-finding tools, such as MCL, OrthoMCL and OrthoFinder. It can be carried out by carrying out BLAST+ searches using a short program, and this will be illustrated below. | ||
| 2. ortholog | ||
| [OrthoFinder](https://github.com/davidemms/OrthoFinder) | ||
| 3. synteny block | ||
| [McScanX](<https://github.com/tanghaibao/jcvi/wiki/MCscan-(Python-version)>) | ||
| 4. crosssmap | ||
| [minimap](https://github.com/lh3/minimap2) | ||
| or | ||
| [AnchorWave](https://github.com/baoxingsong/AnchorWave) | ||
|
|
||
| ## Install | ||
|
|
||
| TODO | ||
|
|
||
| ```sh | ||
| $ pip3 install GeneMap | ||
| ``` | ||
|
|
||
| ## Usage | ||
|
|
||
| ```shell | ||
| python3 bin/GeneMap.py -l tests/Zm-B73v5.genelist -d . -q Zm-B73v5 -t Zv-TEO01 | ||
| ```sh | ||
| $ GeneMap.py -l list.txt -d work_dir -q GenomeA -t GenomeB -o output_prefix | ||
| ``` | ||
|
|
||
|  | ||
| ## Maintainers | ||
|
|
||
| [@wjwei-handsome](https://github.com/wjwei-handsome) | ||
|
|
||
| ## Contributing | ||
|
|
||
| Feel free to dive in! [Open an issue](https://github.com/wjwei-handsome/GeneMap/issues/new) or submit PRs. | ||
|
|
||
| Standard Readme follows the [Contributor Covenant](http://contributor-covenant.org/version/1/3/0/) Code of Conduct. | ||
|
|
||
| ### Contributors | ||
|
|
||
| Thanks [@songtaogui](https://github.com/songtaogui) for design ideas. | ||
|
|
||
| ## License | ||
|
|
||
| [GPL-3.0](LICENSE) © Weiwenjie |