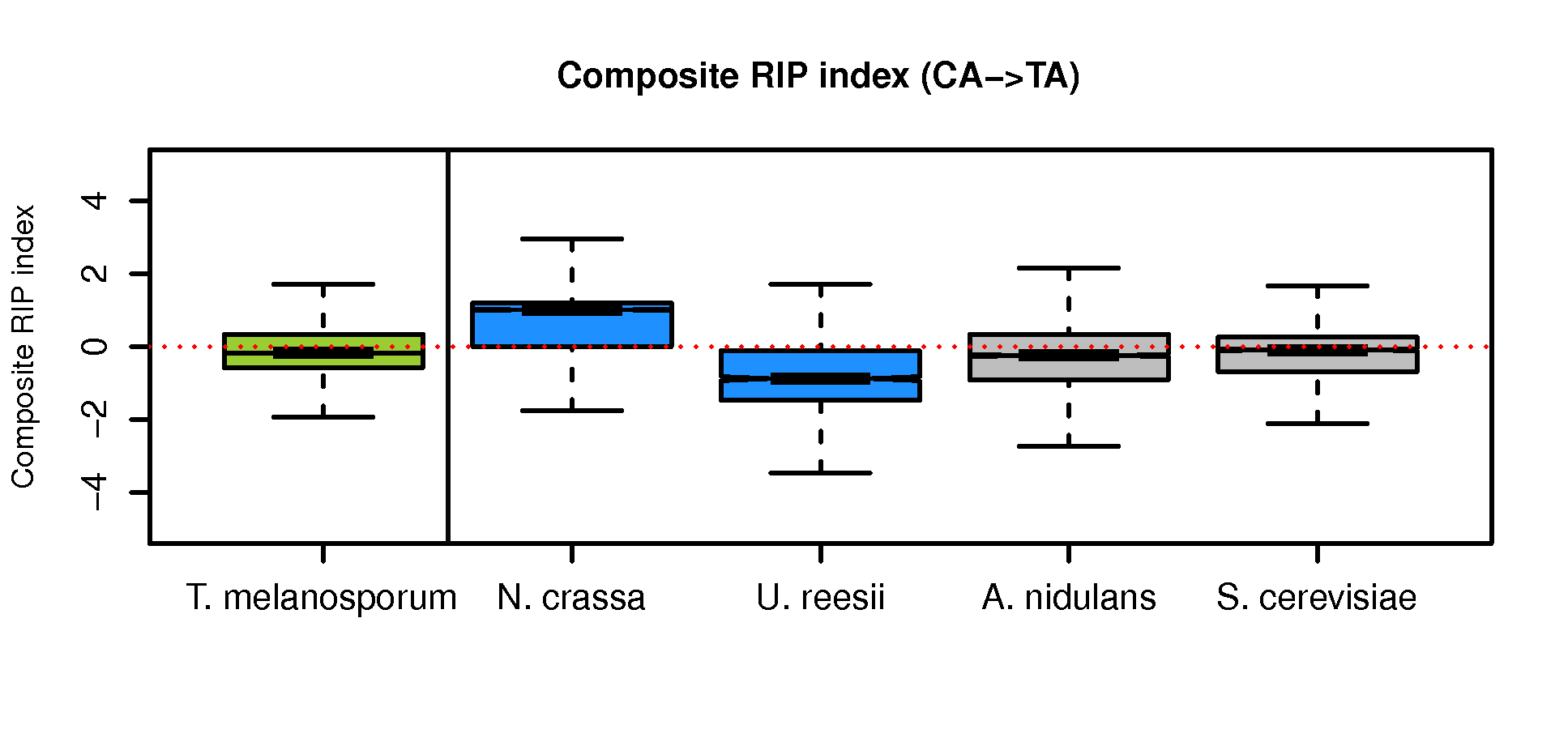
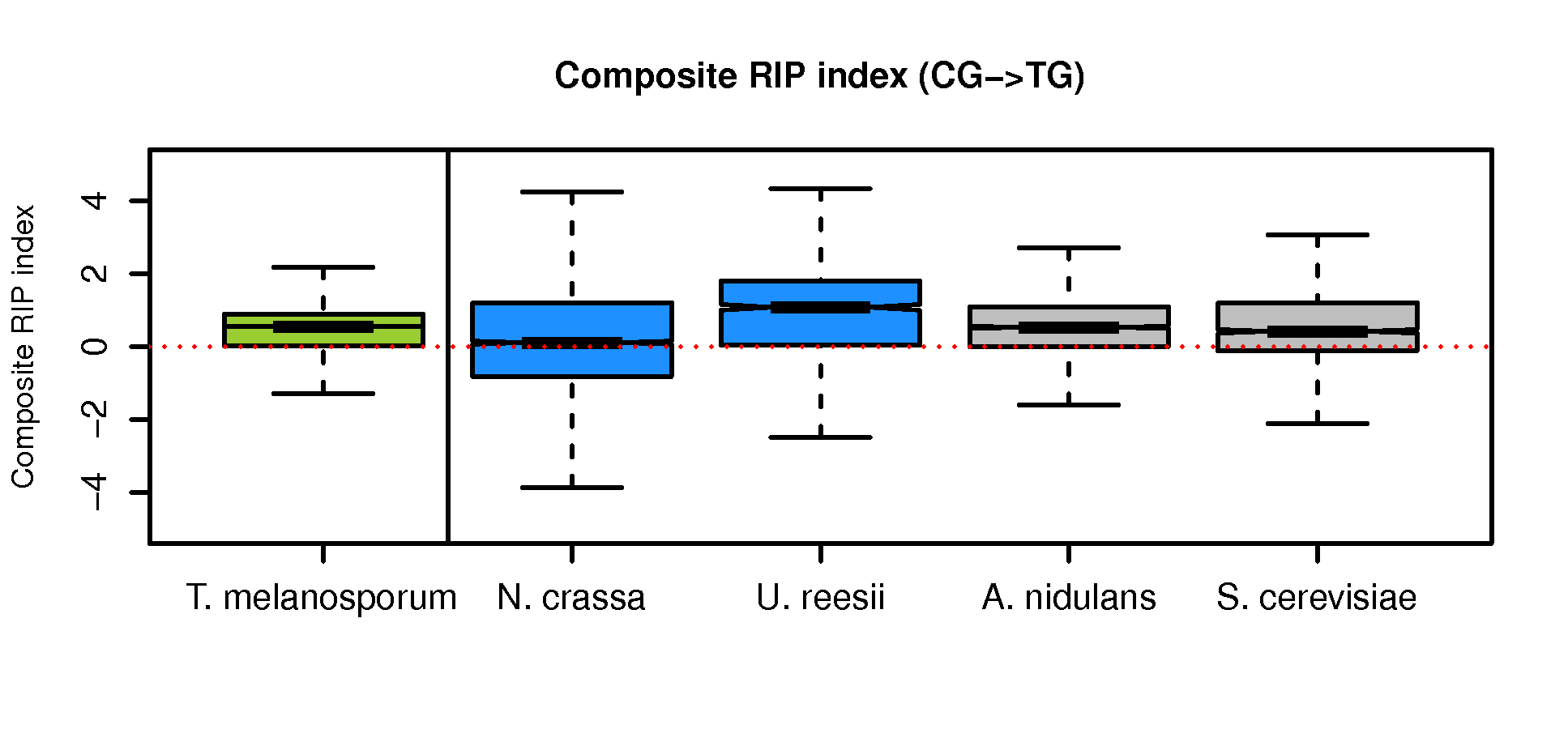
Calculate composite RIP index (CRI) score and generate box plots
Python 2.7 and R
MIT
- Neurospora_crassa.unmasked.fasta.seqs.txt.gz
- Saccharomyces_cerevisiae.EF4.66.dna.toplevel.fa.seqs.txt.gz
- Tuber_melanosporum_v1.0_unmasked.fa.seqs.txt.gz
- aspergillus_nidulans_fgsc_a4_1_contigs.fasta.seqs.txt.gz
- uncinocarpus_reesii_2_supercontigs.fasta.seqs.txt.gz
See other procedures in Methods for the CRI formula.
Example:
$ python CRI_fasta.py -f Tuber_melanosporum_v1.0_unmasked.fa.seqs.txt
Output files:
All outputs:
- CRI_CA_Neurospora_crassa_repeats.txt
- CRI_CA_aspergillus_nidulans_fgsc_a4_1_contigs_repeats.txt
- CRI_CA_Saccharomyces_cerevisiae_repeats.txt
- CRI_CA_Tuber_melanosporum_v1_repeats.txt
- CRI_CA_uncinocarpus_reesii_2_supercontigs_repeats.txt
- CRI_CG_aspergillus_nidulans_fgsc_a4_1_contigs_repeats.txt
- CRI_CG_Saccharomyces_cerevisiae_repeats.txt
- CRI_CG_Tuber_melanosporum_v1_repeats.txt
- CRI_CG_uncinocarpus_reesii_2_supercontigs_repeats.txt
- CRI_CG_Neurospora_crassa_repeats.txt
Run R program:
Boxplot-CRI-Tuber-vs-other4.R
Output files: