Discriminative Transcription factor Occupancy eXtraction
Predicting transcription factor binding remains challenging due to high false positive rates, cell type specific differences in DNA recognition, and experimental bias. We developed a motif-based discriminative method, dTOX (discriminative Transcription factor Occupancy eXtraction), to predict transcription factor binding based on a single data type—either PRO-seq, ATAC-seq, or DNase-I-seq.
dTOX is available to use in our online gateway for hg19 and mm10 or as a stand-alone R package for other genomes. Both implementations only require plus and minus strand bigWig files for PRO-seq, ATAC-seq, or DNase-I-seq data.
We provide a computational gateway to run dTOX on a GPU server. This gateway allows users to upload bigWig files and download the results, without installing any software, making it simple and easy to find transcription factor binding patterns.
Please click the link to try this site:
On the online service, dTOX can only be run on hg19 and mm10. If your studies are not limited to these species, you can install the dTOX pipeline and run your data locally.
Before you run your data on the dREG gateway, please check the server status here.
The Exchange email system might quarantine all emails including the word “password” or other sensitive things in links. (https://technet.microsoft.com/en-us/library/aa997692(v=exchg.160).aspx).
Unfortunately, some emails from dREG gateway are quarantined by this spam policy. Usually these quarantined emails are not delivered to the email box, so they can not be checked in any email folders, including junk, spam or inbox. If you find the emails from dREG gateway are not delivered into your email box, please conect the administrator of your email system. For the Cornell email, please check this link:
https://it.cornell.edu/spam-control/log-quarantine-management-spam-control
dTOX takes bigWig files with double strands for PRO-seq, ATAC-seq and DNase-I-seq, as the input.
-
Only positive values or only negative values in each strand, no mixture.
-
No normalization.
-
(PRO-seq only) Each read is mapped at 5’ (GRO-seq) or 3’ (PRO-seq) position (point mode) , not mapped to a continuous region starting from 5’ or 3’. This is different with the software Tfit.
To generate bigWig files from fastq data, please refer to https://github.com/Danko-Lab/proseq2.0/
To generate bigWig files from bam files, please refer to https://github.com/Danko-Lab/RunOnBamToBigWig/
To generate bigWig files from bam files, please refer to https://github.com/Danko-Lab/utils/tree/master/dnase/BamToBigWig
To generate bigWig files from bam files, please refer to https://github.com/Danko-Lab/utils/tree/master/atac/BamToBigWig
The source code and models of dTOX will be availiable on GitHub (https://github.com/Danko-Lab/dTOX).
Linux and Mac OSX are currently supported.
- bedops (http://bedops.readthedocs.org/en/latest/index.html)
- R (http://www.r-project.org/)
- bigWig R package (https://github.com/andrelmartins/bigWig; will be public very soon).
This software is already installed on many UNIX systems. Users can install the most appropriate version of these files for Ubuntu using:
sudo apt-get install r-base-core
sudo apt-get install libssl1.0.0 libssl-dev
Users who are not sure how to install the proper dependencies on their system should check with their system administrator for help.
dTOX also has several dependencies within R. These include data.table, e1071, mvtnorm, parallel, randomForest, rmutil, rphast, and snowfall. These packages are all availiable on the CRAN repository. For convenience, users can install these packages using the makefile:
make R_dependencies
If users run into any problems they should contact the package author for assistance.
Users should change to the directory containing this README.md file, and can then install dTOX by typing the following:
(1a) Install R dependencies
make R_dependencies
(1b) Install dTOX
make dTOX
(1c) Install Rgtsvm if you have GPU nodes.
make Rgtsvm
Pre-trained models that can be used to predict TF binding status across the genome are availiable in mammals. Download the newest models and TFBS data set from here: ftp://cbsuftp.tc.cornell.edu/danko/hub/dTOX/
dTOX provides a solution to identify TF binding status for 3 data types: PRO-seq, ATAC-seq, and DNase-I-seq.
Type:
bash run_dTOX.bsh genome_id seq_type plus_strand_bw minus_strand_bw out_prefix filter gpu_cores cpu_cores
Genome_id -- Currently only hg19 and mm10 by default, for other species, please refert to 3).
Sequeue Type -- ATAC-seq, DNase-1-seq or PRO-seq
plus_strand.bw -- Seqence data (plus strand). Read counts (not normalized) formatted as a bigWig file.
minus_strand.bw -- Seqence data (minus strand). Read counts (not normalized) formatted as a bigWig file.
out_prefix -- The prefix of the output file.
filter -- The filter is used or not, "TRUE" is used and "FALSE" is not.
gpu_cores -- [optional, default=1] indicating how many GPU cores can be used.
cpu_cores -- [optional, default=16] indicating how many CPU cores can be used.
For example, to run dTOX on the ATAC-seq data, use:
bash run_dTOX.bsh hg19 ATAC-seq atac.plus.bw atac.minus.bw atac.out TRUE 1 16
One file below is generated in this solution:
-
<out_prefix>.dTOX.bound.bed.gz:
reduced peak information, only including peak position, binding motif, max score, strand, predicted value, predict status.
Notice:
(1) That command takes 8~20 hours to execute on NVIDA GPU (K80, P100 or other types) using Rgtsvm package. Due to very long computational time, we don't suggest to run peak calling on CPU nodes, even in parallel mode.
The results generated by dTOX include all motifs define in CIS-BP library. If you only want to check the bound TFBS for specific TF, please use the following command:
Type:
bash extract_TF.bsh genome_id TF_name dTOX_reult out_file
genome_id -- hg19, mm10 or cisbp_zip file used in the usage 3).
TF_name -- TF Name, e.g. JUND, ELK1, MYC
dTOX_reult -- the result file generated by the 'run-dTOX' script. (Compressed bed file)
out_file -- the name of the output file. (Compressed bed file)
For example, to extract JUND from the results:
bash run_dTOX.bsh hg19 JUND atac.out.dTOX.bound.bed.gz JUND.out.bed.gz
Type:
bash run_rtfbsdb.bsh genome_id cisbp_zip genome_2bit TFBS_output_dir [cpu_cores]
genome_id -- any genome id rather than hg19 and mm10.
cisbp_zip -- The zip file including motifs list and PWM file. (downloaded from cis-bp).
genome_2bit -- The 2 bit file of genome
TFBS_output_dir -- The path to output the TFBS bed files for each motif define in cisbp_zip
cpu_cores -- [optional, default=1] indicating how many CPU cores can be used.
The pipeline perform the following tasks and take a long time to finish it (>=48 hours)
-
The script load all motifs define in the zip file (downloaded from CIS-BP)
-
scans the whole genome file(genome_2bit) to get all TFBS and write into the output folder indicated by 'TFBS_output_dir'.
-
get the summary information of RTFBSDB score around 25 bp at both side
-
write the TFBS region and summary information into TFBS folder within this package, named "TFBS/motif_rtfbsdb_{genome_id}_ext".
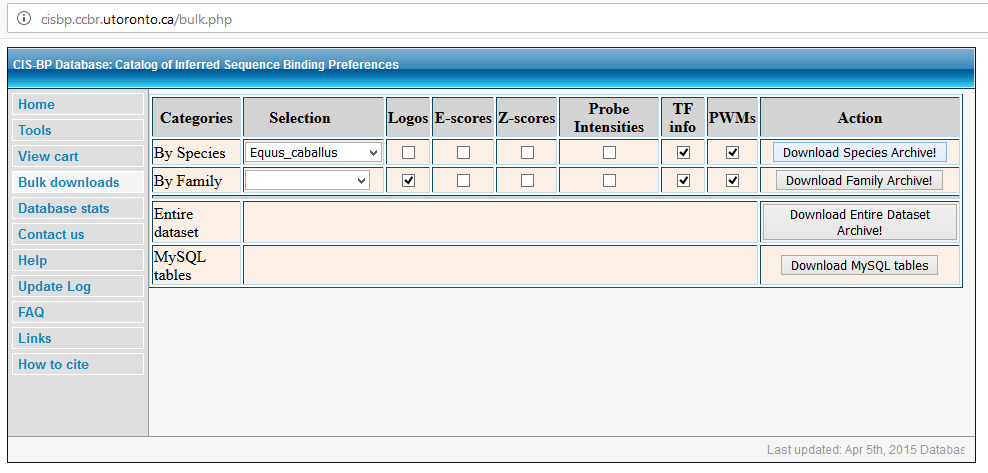
How to download motif information from the CIS-BP database (http://cisbp.ccbr.utoronto.ca/)
(1) Choate LA, Wang Z, and Danko CG. Identification of transcription factor binding patterns using genome-wide accessibility and transcription. 2019.