| SEP | |
|---|---|
| Title | Inert DNA spacer |
| Authors | Jacob Beal (jakebeal@ieee.org), Shyam Bhakta, John Sexton |
| Editor | TBD |
| Type | Specification |
| SBOL Visual Version | 2.3 |
| Status | Accepted |
| Created | 7-Sept-2020 |
| Last modified | 2-Oct-2020 |
| Issue | #103 |
This SEP proposes to add a new glyph for inert DNA spacer sequences, generalizing on the concept of an insulator.
The insulator glyph is defined as SO:0000627, which is more biologically specific than the term is being used in practice. At the same time, the current "double box" insulator glyph has not found much favor, and has caused problems due to its design.
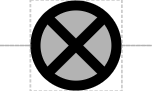
This SEP thus proposes to introduce a new "inert DNA spacer" glyph using the "circle-X" notation that has appeared in a number of published papers. While this has similarity to the Ori and OriT glyphs, it is still quite visually distinct.
The insulator glyph will also be deprecated, to be deleted at version 3.0.
SO:0002223 (Inert DNA Spacer)
The inert DNA spacer glyph is a circle with an X in its middle, suggesting the intent to cancel possible interactions:
Inserted 5' sequence intended to reduce effect of upstream genetic context on promoter behavior.
this section deliberately blank
Added to notes:
This glyph is deprecated in favor of inert DNA spacer.
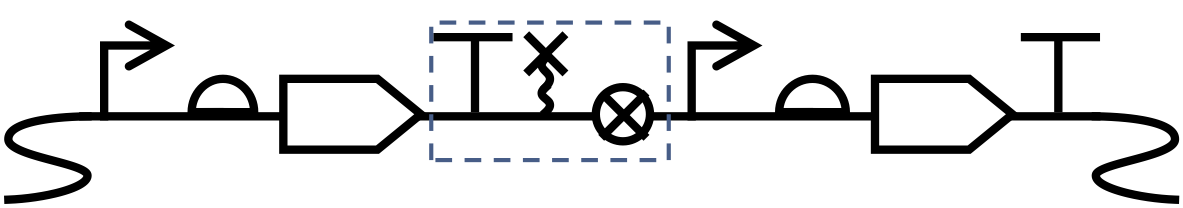
Ribozyme and spacer between two transcriptional units, forming a separation module with the preceding terminator (dashed box), with the whole construct integrated into the chromosome.
Addition of "Inert DNA Spacer" is backward compatible, as is deprecation of Insulator. The actual removal of Insulator will not be backward compatible, however, and thus must wait for 3.0.
A number of potential alternatives were discussed in the GiHub issue thread prior to creation of this SEP: #34

To the extent possible under law,
SBOL developers
has waived all copyright and related or neighboring rights to
SEP V022.
This work is published from:
United States.