forked from Robbie1977/VFB2-draft
Commit
This commit does not belong to any branch on this repository, and may belong to a fork outside of the repository.
- Loading branch information
Showing
3 changed files
with
22 additions
and
0 deletions.
There are no files selected for viewing
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,22 @@ | ||
| --- | ||
| title: "Transcriptomics data available for cell types on VFB" | ||
| linkTitle: "Transcriptomics data" | ||
| date: 2024-04-10 | ||
| description: > | ||
| Transcriptomics data is now available on VFB! Find scRNAseq clusters from the Term Info pane for a cell type of interest. | ||
| --- | ||
|
|
||
| We now incorporate scRNAseq data from multiple studies, including the [Fly Cell Atlas](https://flycellatlas.org/) project and other datasets that identify nervous system cell types. These datasets can be found by searching for 'scRNAseq' and filtering to 'Dataset'. | ||
|
|
||
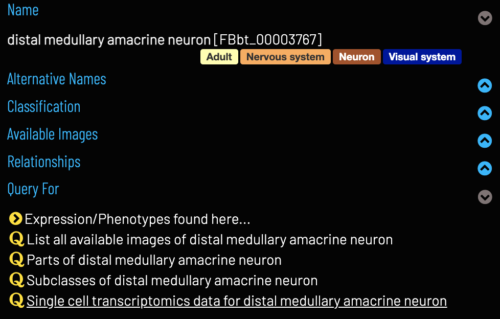
| Clusters can be found for particular cell types via the 'Single cell transcriptomics data for..' query on the cell type Term Info pane. | ||
|
|
||
|  | ||
|
|
||
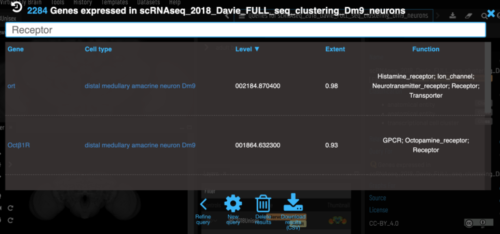
| Genes for a particular cluster can be filtered by function and sorted by expression level and extent (proportion of cells in cluster expressing the gene). As with other VFB search results, these can be exported as a csv. | ||
| _Note that we currently only include genes that have extent > 0.2 in a cluster._ | ||
|
|
||
|  | ||
|
|
||
| Data can also be retrieved using [VFB_connect](https://vfb-connect.readthedocs.io/en/stable/API_reference.html#transcriptomics-queries). | ||
|
|
||
| We pull scRNAseq data from [FlyBase](https://flybase.org/), which takes it from the [Single Cell Expression Atlas](https://www.ebi.ac.uk/gxa/sc/home). |
Sorry, something went wrong. Reload?
Sorry, we cannot display this file.
Sorry, this file is invalid so it cannot be displayed.
Sorry, something went wrong. Reload?
Sorry, we cannot display this file.
Sorry, this file is invalid so it cannot be displayed.