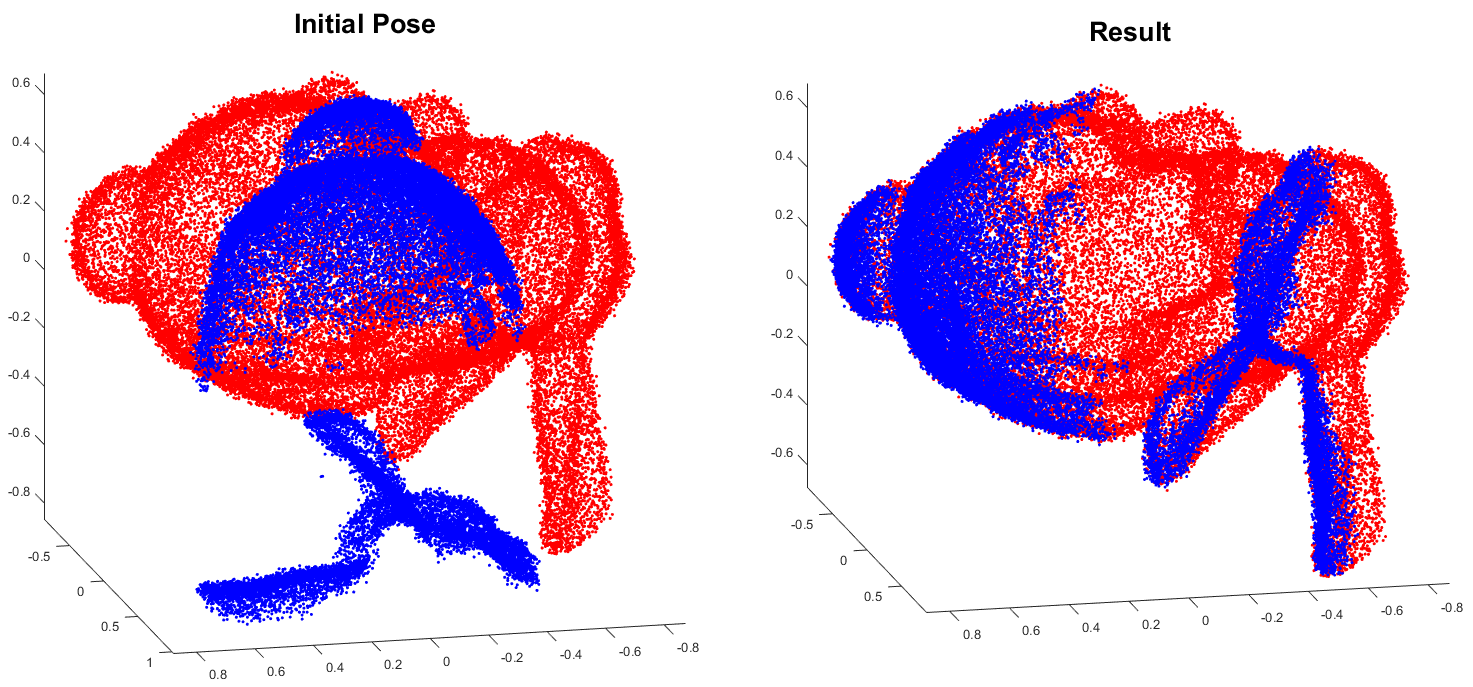
This is a cython version of the original go-icp project by yangjiaolong.
(A demo video can be found on here.)
This repository contains the Cythonized code for the Go-ICP algorithm (with trimming strategy for outlier handling). It is free software under the terms of the GNU General Public License (GPL) v3. Details of the Go-ICP algorithm can be found in the papers:
-
J. Yang, H. Li, Y. Jia, Go-ICP: Solving 3D Registration Efficiently and Globally Optimally, International Conference on Computer Vision (ICCV), 2013. PDF
-
J. Yang, H. Li, D. Campbell, Y. Jia, Go-ICP: A Globally Optimal Solution to 3D ICP Point-Set Registration, IEEE Transactions on Pattern Analysis and Machine Intelligence (TPAMI), 2016. PDF
Please read this file carefully prior to using the code. Some frequently-asked questions have answers here.
The cython module uses Autowrap to created the pyx and cpp files. Also changes involve removal of .cpp files and having only .hpp files. Although you don't have to do anything but run on the head folder the below command,
python setup.py build_ext --inplace
If you wish to generate the pyx files after modifying the source code in c++ (Adventure is waiting), then run the below command from the terminal inside the 'src' folder
autowrap --out py_goicp.pyx goicpcc.pxd
You need CYTHON installed on your python or virtualenv, or conda environment (whichever you use) to compile and use this library
The package if you have compiled it manually then it can be installed using the command
python setup.py install
Otherwise this is available on the pip package as well. The current version is 0.0.3 and can be installed with
pip install py-goicp (you need administrator, else use --user flag
Use the test.py lying in parallel to the setup.py file. This should teach you on how to use the code. For the sake of simplicity only the below classes have been wrapped to use,
- GoICP
- POINT3D
- ROTNODE
- TRANSNODE
A simple usage will be (after setting the parameters)
import numpy as np;
from py_goicp import GoICP, POINT3D, ROTNODE, TRANSNODE;
def loadPointCloud(filename):
pcloud = np.loadtxt(filename, skiprows=1);
plist = pcloud.tolist();
p3dlist = [];
for x,y,z in plist:
pt = POINT3D(x,y,z);
p3dlist.append(pt);
return pcloud.shape[0], p3dlist;
goicp = GoICP();
Nm, a_points = loadPointCloud('./test_data/model_bunny.txt');
Nd, b_points = loadPointCloud('./test_data/data_bunny.txt');
goicp.loadModelAndData(Nm, a_points, Nd, b_points);
goicp.setDTSizeAndFactor(300, 2.0);
goicp.BuildDT();
goicp.Register();
print(goicp.optimalRotation()); # A python list of 3x3 is returned with the optimal rotation
print(goicp.optimalTranslation());# A python list of 1x3 is returned with the optimal translation-
Make sure both model and data points are normalized to fit in [-1,1]3 prior to running (we recommend first independently centralizing the two point clouds to the origin then simultaneously scaling them). The default initial translation cube is [-0.5,0.5]3 (see “config_example.txt”).
-
The convergence threshold is set on the Sum of Squared Error (SSE) as in the code and the paper. For the ease of parameter setting for different data point numbers, we use Mean of Squared Error (MSE) in the configuration (see “config_example.txt”). We use MSE threshold of 0.001 for the demos. Try smaller ones if your registration results are not satisfactory.
-
Make sure you tune the trimming percentage in the configuration file properly, if there are some outliers in the data pointset (i.e., some regions that are not overlapped by the model pointset). Note that a small portion of outliers may lead to competely wrong result if no trimming is used. Refer to our TPAMI paper for more details.
-
Building 3D distance transform with (default) 300 discrete nodes in each dimension takes about 20-25s in our experiments. Using smaller values can reduce memory and building time costs, but it will also degrade the distance accuracy.
This implementation uses the nanoflann library, and a simple matrix library written by Andreas Geiger. The distance transform implementation is adapted from the code of Alexander Vasilevskiy.
This impelementation uses the c++ code of yangjiaolong. My sincere thanks to the author for this valuable contribution to the geomety processing society. May the blessings shower the author all the time...
V0.1 (15-January-2019)
First complete version for release