-
Notifications
You must be signed in to change notification settings - Fork 2
Model Setup
The model code used is the Massachusetts Institute of Technology general circulation model (MITgcm) (Marshall 1997). The MITgcm is a finite volume primitive equation model solving the incompressible Navier-Stokes equations. The MITgcm can be downloaded here.
Best practice is to have a clean copy of the model to refer to. For each set up then have a folder with a code, buid and input directory along with an opt file. I like to even alias the genmake command so I can just leave that with the original model code.
alias genmake2='/home/hb1g13/MITgcm/MITgcm/tools/genmake2'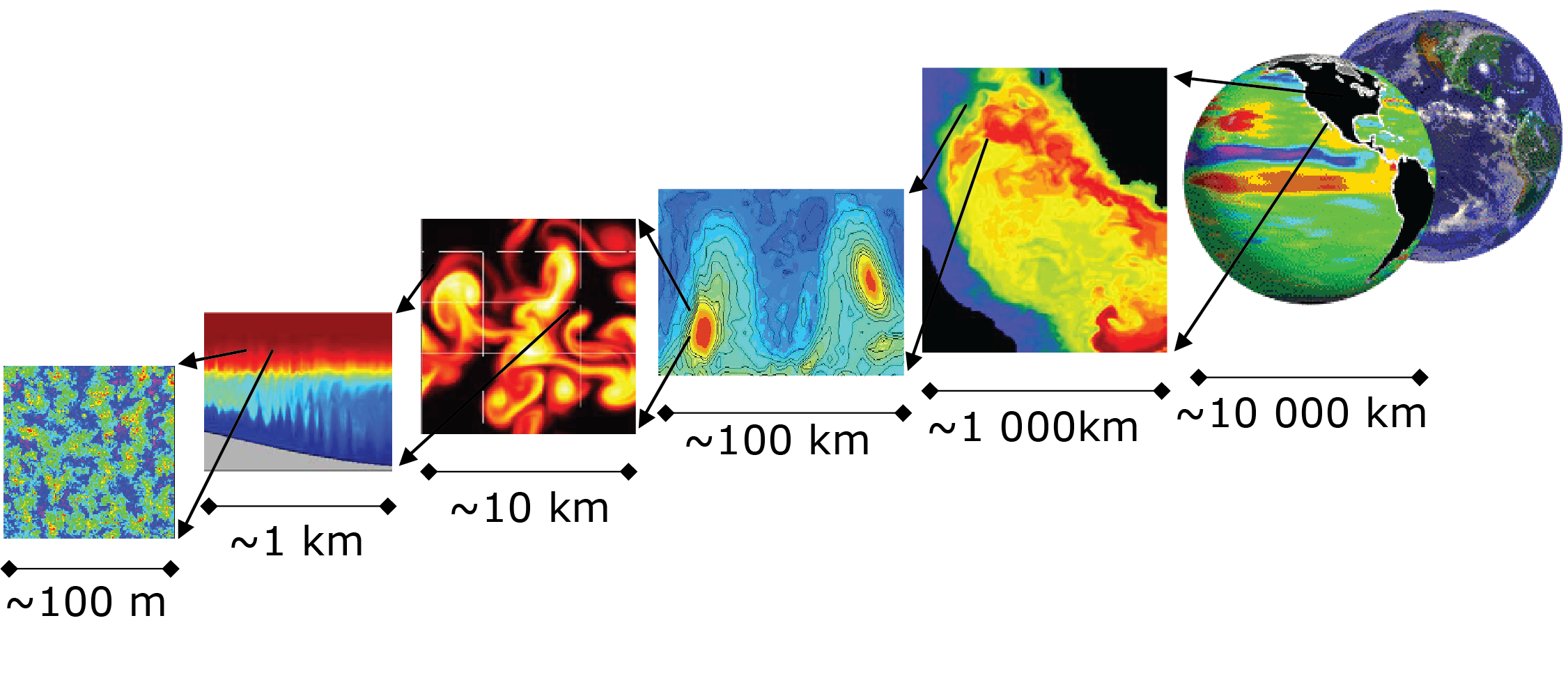
The southern ocean has complex non-linear dynamics which require A simple box ocean can then be used to run a high-resolution simulation to enable us to examine the basic physics controlling Southern Ocean diabatic eddies.
Higher resolution both increases the number of grid points and requires shorter time steps to satisfy the CFL condition (discussed separately later) therefore increases in resolution rapidly increasing computational cost. For the purpose of investigating eddy dynamics the coarsest resolution that is acceptable will be ``just about" eddy resolving. The Rossby radius of deformation () is calculated by:
where N is the buoyancy frequency typically , H is thickness scale (typically 1000 m) and f is the Coriolis parameter (
). Using those typical values we can estimate the Rossby radius 10-30 km. To resolve for eddies a minimum of 2 grid spacing per radius is required, therefore we can use 5km as our horizontal resolution.
Gendata.py is an example python script generating the model grid and forcing for ease of reading I have included an ipython notebook with the figures and equations in Gendata.ipynb
An example optimization file can be found here this is based off example opt files included in the MITgcm (outlined here) and years of experience coming from Jeff Blundell and then altered for the exact system used.
Derived from linux_amd64_ifort example but Should work fine on EM64T and other AMD64 compatible Intel systems
-
Processor-specific flags: * for more speed on Core2 processors replace -xW with -xT * for more speed on Pentium4 based EM64T processors replace -xW with -xP
-
For more speed, provided your data size doesn't exceed 2GB you can remove -fPIC which carries a performance penalty of 2-6%.
-
Provided that the libraries you link to are compiled with -fPIC this opt file should work.
-
You can replace -fPIC with -mcmodel=medium which may perform faster than -fPIC and still support data sizes over 2GB per process but all the libraries you link to must be compiled with -fPIC or -mcmodel=medium
-
Changed from -O3 to -O2 to avoid buggy Intel v.10 compilers. Speed impact appears to be minimal.
This version is intended to implement MPI parallelism.
In the code directory we will define the domanin size in SIZE.h:
PARAMETER (
& sNx = 50,
& sNy = 50,
& OLx = 4,
& OLy = 4,
& nSx = 1,
& nSy = 1,
& nPx = 16,
& nPy = 8,
& Nx = sNx*nSx*nPx,
& Ny = sNy*nSy*nPy,
& Nr = 30) c numer of dept lvlsWe've chosen an x,y,z domain of 800 x 400 x 30 grid points. We're going to run on 8 nodes with 16 cores ( 128 cores) The three-dimensional domain is comprised of nPxnSx blocks of size sNx along one axis nPynSy blocks of size sNy along another axis and one block of size Nz along the final axis. Blocks have overlap regions of size OLx and OLy along the along the dimensions that are subdivided.
packages.conf defined packages to be included:
This set up uses:
- gfd - This is predefined group of standard packagesgfd : mom_common mom_fluxform mom_vecinv generic_advdiff debug mdsio rw monitor
- mnc - net cdf output
- timeave - timeaveraging
- diagnostics - diagnostics package for file outputs
- cd_code - used to fix a computational mode which is sometimes excited when the Coriolis force is exactly aligned with a relatively low-resolution lat-lon grid.
- rbcs - northern boundary sponge layer
- layers - for calculation of isothermal streamfuction
You make need to edit the header files for these packages for example the layers package needs the number of layers defined here,
- Check code files like packages.conf have packages turned on etc
- Check code folder contains hear file expected
- Go to build directory
- Run the following commands
make CLEAN
genmake2 --mpi --mods=../code ../optfile=../linux_opt_file
make depend
makeA wee while late you have an executable mitgcmuv which you will need to move to your run directory symbolic link to.
Gendata_Nchannel.py generates binary and netCDF files of the model grid and forcing files (wind, heat flux etc) as well as the sponge layer mask. To see in detail the forcing generated and rationale behind it please refer to the ipython notebook Gendata_Nchannel.ipynb
Writebin.py is used in order to write the MITgcm binaries in the correct format.
** This script should be run in the run directory on a login node of HPC system to be used**
- data - Runtime flags
- eedata - Allows debugging
- data.pkg - Packages on/off
- data.mnc - netCDF options
- data.layers - layers options
- data.rbcs - Sponge layer options (mask file, relaxation time scale)
- data.diagnostics - Output of model diagnostics
In the run directory, you will have your input files alongside files in a format such as sample run dir as well as your executable mitgcmuv
Depending on your HPC system the model will have to be run in chained jobs, the system we used allowed us to run 12 hours at a time. This required the model to run in 20-year chunks. Writing all the data files on job submission files for this is tedious so I have included starter files in each directory and bash scripts to auto-generate the required files.
After auto-generating the required files. You can batch submit jobs e.g. for SLURM this bash script will autorun the first 20 jobs.
This submits each job file which will:
-
Calls the next files -
-
Runs the MITgcm models next run