New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Regression: Missing/black chromosome name in batch mode using MATE_CHROMOSOME grouping since 2.12.0 #1172
Comments
|
Could you provide a test script, it's o.k. if the bam file isn't reachable I'll substitute another. |
|
@cbrueffer ignore test request, I can reproduce it. Tangentially, I don't recall why we paint group labels (chromosome names in this case) with black background / white text, I think they should look as they do on screen. |
|
Glad to hear it, I was just putting the files together. I was wondering the same, at first I thought it was an Xvfb artifact. |
|
I tracked it down, its a java idiosyncrasy, found this in the java docs. I have switched from using "clearRect" to "fillRect" which seems to fix the problem. Fix will be in the next release, early August, and in the development snapshot within the next few minutes.
|
|
Thanks a lot for the quick fix! |
…nment track group labels. Now uses white background - black text to match onscreen appearance. Fixes #1172
First of all, thanks for all your work on this software!
Working version: up to and including 2.11.9
Broken version: from 2.12.0 to latest (2.13.2 as of now)
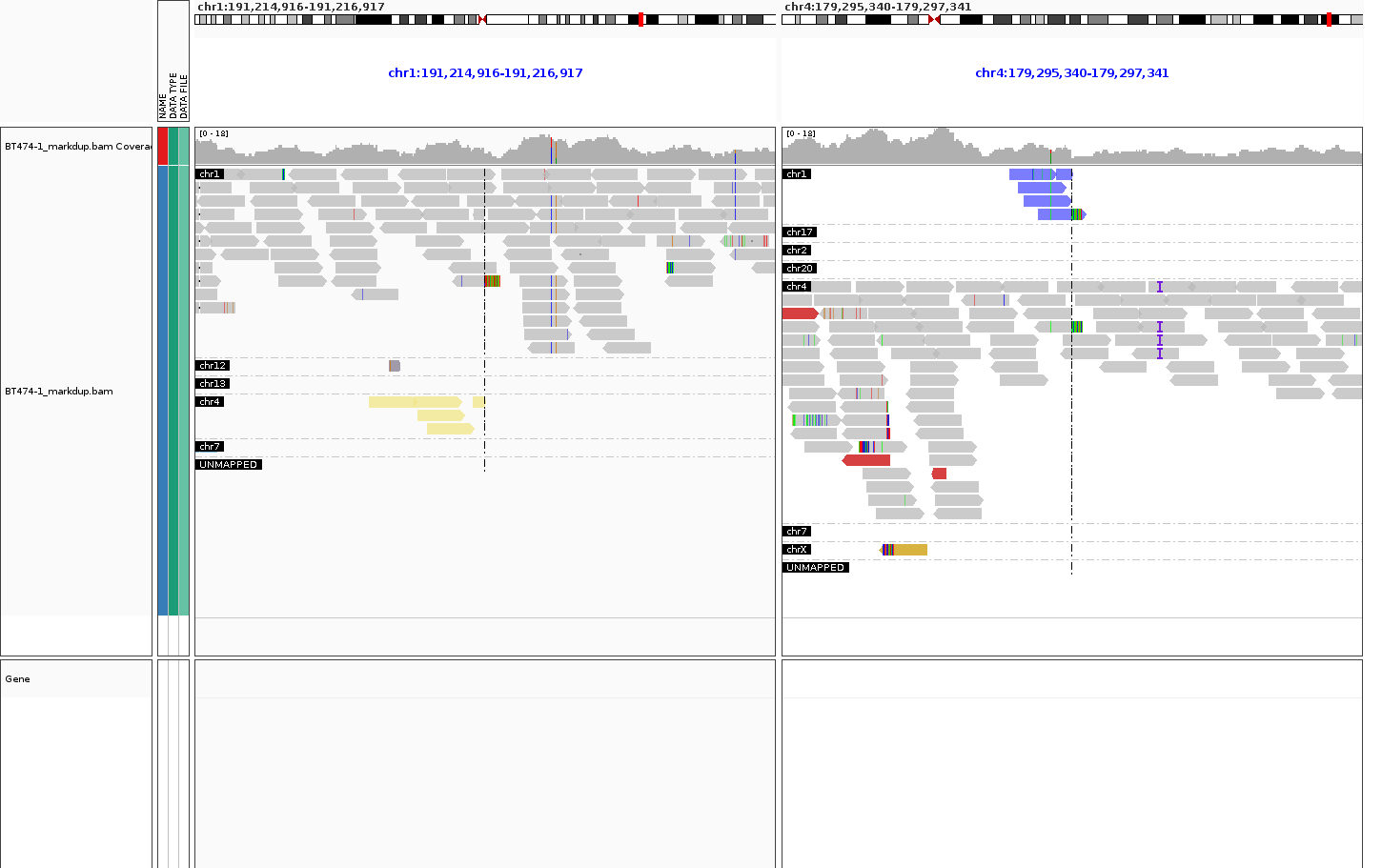
I'm using IGV in batch mode and headless using xvfb-run with option SAM.GROUP_OPTION=MATE_CHROMOSOME. This should show the chromosome name in each "read group" in the top-left corner of each grouping. In batch mode this worked up to and including version 2.11.9. Since version 2.12.0, including the latest 2.13.2, only a black box is being printed.
Environment:
The first (working) screenshot is from 2.11.9, the second (broken) from 2.12.0. Config, batch file etc are the same.
The text was updated successfully, but these errors were encountered: