-
Notifications
You must be signed in to change notification settings - Fork 0
Home
Jean Ollion edited this page Dec 27, 2021
·
9 revisions
Segmentation of bacteria growing on agar-pads, imaged by de-focused transmitted light stacks
- Expected input images are stacks of 2D images, with Z-axis last : Image = [batch, Y, X, Z]. We advise stacks of 5 slices with a 0.2µm step, in range [-0.6µm, -1.4µm] (relatively to the focal plane)
- Segmentation is performed by regression of the Euclidean Distance Map (EDM).
- This repository does not include the downstream watershed step to obtain labeled images. It is included with bacmman software, see instructions below.
| Input transmitted-light Stack | Predicted EDM | Segmented Bacteria |
|---|---|---|
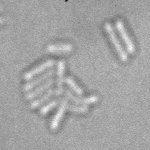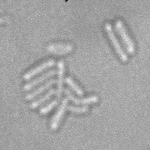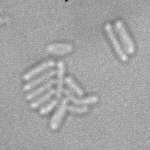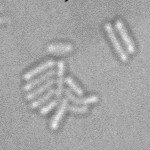 |
 |
 |
samples provided by Daniel Thédié, El Karoui Lab, University of Edinburg
- Based on U-net
- At first layer, Z-axis is both:
- Considered as channel axis and treated with with 2D convolutions
- Reduced using 3D convolutions and 3D max-pooling