-
Notifications
You must be signed in to change notification settings - Fork 2
Verticall summary
The verticall summary command is an optional step to summary the painting of one assembly relative to all other assemblies. It takes the TSV file made by verticall pairwise and produces either:
- a tab-delimited table (printed to stdout) which shows the number of vertical, horizontal and unaligned paintings.
- an interactive plot visualising the table data
To produce a table:
verticall summary -i verticall.tsv -a assemblies/abc.fastaTo produce an interactive plot:
verticall summary -i verticall.tsv -a assemblies/abc.fasta --plotHere's an example plot showing the summary for a Klebsiella variicola assembly in a collection of 210 assemblies spanning the Klebsiella pneumoniae species complex:
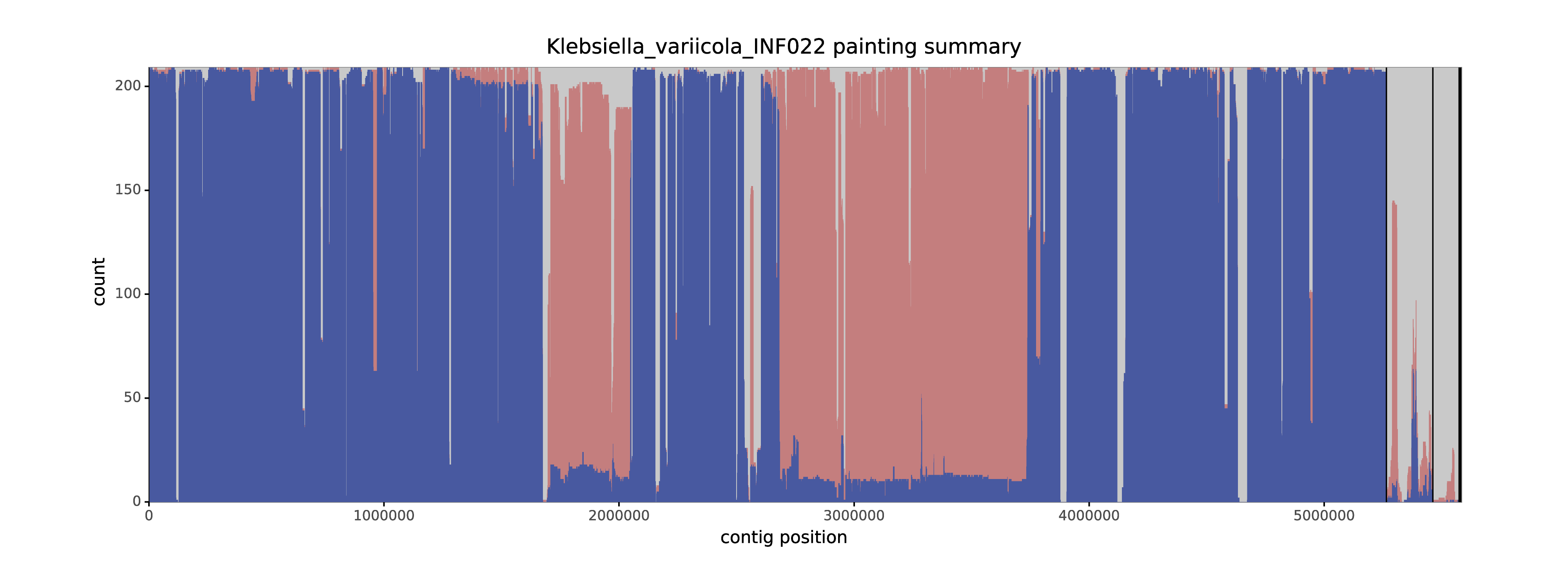
This was a completed assembly, which makes for a cleaner summary plot. Contigs are separated by solid vertical lines (most of the plot is the big chromosomal contig). The data is shown as a stacked area plot, where blue is vertical, red is horizontal and grey is unaligned. These three values total to 209 (the number of other assemblies) at each position. For most of the assembly, this genome is vertical compared to the other assemblies, indicated by the blue. However, two large pieces of the chromosome are mostly horizontal (this was a case of cross-species recombination). And a few pieces of the assembly (especially in the plasmid contigs) are mostly unaligned, indicating accessory genome.
usage: verticall summary -i IN_FILE -a ASSEMBLY [--all] [--plot] [--vertical_colour VERTICAL_COLOUR]
[--horizontal_colour HORIZONTAL_COLOUR] [--unaligned_colour UNALIGNED_COLOUR]
[-h] [--version]
summarise regions for one assembly
Required arguments:
-i IN_FILE, --in_file IN_FILE Filename of TSV created by vertical pairwise
-a ASSEMBLY, --assembly ASSEMBLY
Filename of assembly to be summarised
Settings:
--all Output one line for all assembly positions (default: omit redundant
adjacent lines)
--plot Instead of outputting a table, display an interactive plot (default:
do not display a plot)
Colours:
--vertical_colour VERTICAL_COLOUR
Hex colour for vertical inheritance (default: #4859a0)
--horizontal_colour HORIZONTAL_COLOUR
Hex colour for horizontal inheritance (default: #c47e7e)
--unaligned_colour UNALIGNED_COLOUR
Hex colour for unaligned inheritance (default: #c9c9c9)
Other:
-h, --help Show this help message and exit
--version Show program's version number and exit
- Home
- Software requirements and installation
- How to run Verticall
- Usage information
- Implementation details
- Other info