New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
ENH:stats: Add _sf and _isf methods to kappa3 #18822
Conversation
from scipy.stats import kappa3
from scipy.special import xlog1py, expm1
import numpy as np
import matplotlib.pyplot as plt
def _isf(q, a):
lg = xlog1py(-a, -q)
den = expm1(lg)
return (a / den)**(1.0 / a)
def plot_isf():
a = 2.0
q = np.logspace(-30, -1, 200)
plt.semilogx(q, kappa3.isf(q, a), label="main", ls="dashed")
plt.semilogx(q, _isf(q, a), label="pr", ls="dotted")
plt.legend()
plt.title("kappa3 inverse survival function")
plt.show() |
|
If the new round trip test doesn't pass with this distribution, can you add an |
I ran both test_continuous_basic and test_distributions on my local system and they seemed to pass. |
|
Please try it again after adding a case for this distribution to the new test. |
You are right we need to override the _sf method. I might need a bit of time to implement that. |
from scipy.stats import kappa3
import numpy as np
import matplotlib.pyplot as plt
from mpmath import mp
mp.dps = 200
def mpmath_sf(x, a):
x = mp.mpf(x)
a = mp.mpf(a)
return mp.one - x*(a + x**a)**(-mp.one/a)
def plot_sf():
a = 2.0
x = np.logspace(-5, 40, 200)
mp_values = np.array([mpmath_sf(_x, a) for _x in x], np.float64)
plt.loglog(x, kappa3.sf(x, a), label="main", ls="dashdot")
plt.semilogx(x, _sf(x, a), label="pr", ls="dashed")
plt.loglog(x, mp_values, label="mpmath", ls="dotted")
plt.legend()
plt.title("kappa3 survival function")
plt.show() |
|
I thought I'd share the code I used to find these cases. Start by defining a population of input parameters at which to test the SF and ISF functions. Hopefully, we'd have noticed already if arguments in the range 0.1 through 0.9 (e.g. log_min_p = -300
log_1m_max_p = -14
p0 = np.concatenate((10**rng.uniform(log_min_p, np.log10(0.5), size=n), # lots of points close to 0
1 - 10**rng.uniform(log_1m_max_p, np.log10(0.5), size=int(.05*n)))) # fewer points close to 1The values of the shape parameter are also log-uniformly distributed: log10_min_a = -10
log10_max_a = 10
a0 = 10**rng.uniform(log10_min_a, log10_max_a, size=len(p0))
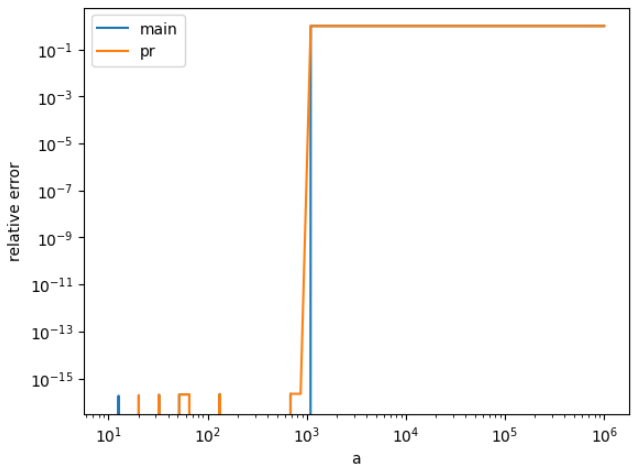
q = isf(p0, a0)Many of these The new implementation has much lower error for this population, but you can see there are some points for which it is not accurate. We can figure out which those are by plotting the error as a function of the arguments. So we see that large values of Note that the population we used was good at sniffing out inaccuracies, but if it is not representative of the population of arguments users will pass into the function, we can't use the histogram to conclude that the PR's implementation is better for SciPy. I don't know the answer to that question. import numpy as np
from scipy import special as sc
from scipy import stats
from mpmath import mp
mp.dps = 1000
import matplotlib.pyplot as plt
# Reference SF
@np.vectorize
def sf(x, a):
x, a = mp.mpf(x), mp.mpf(a)
return mp.one - x*(a + x**a)**(-mp.one/a)
# Reference ISF
@np.vectorize
def isf(x, a):
x, a = mp.mpf(x), mp.mpf(a)
y = mp.one - x
return (a/(y**-a - 1.0))**(1.0/a)
# Determine whether the value can be represented in float64
@np.vectorize
def is_float_representable(x):
finfo = np.finfo(np.float64)
return abs(x) > finfo.tiny and finfo.min < x < finfo.max
# Calculate the log10 of the relative error, clipping to relevant range
def log_relative_error(res, ref):
err = abs((res - ref)/ref)
err = err.astype(np.float64)
# we probably don't care much about the magnitude of relative error
# beyond this range
err = np.clip(err, 1e-20, 1)
return np.log10(err)
n = 1000
# Define population of arguments at which to test the ISF
log_min_p = -300
log_1m_max_p = -14
rng = np.random.default_rng(32492342965356)
p0 = np.concatenate((10**rng.uniform(log_min_p, np.log10(0.5), size=n),
1 - 10**rng.uniform(log_1m_max_p, np.log10(0.5), size=int(.05*n))))
p0 = np.sort(p0)
log10_min_a = -10
log10_max_a = 10
a0 = 10**rng.uniform(log10_min_a, log10_max_a, size=len(p0))
# Compute the ISF using the reference implementation
q = isf(p0, a0)
# Filter out cases for which the result can't be represented in float64
i = is_float_representable(q)
# these are the inputs we'll use for tests of isf
a_isf = a0[i].astype(np.float64)
p_isf = p0[i].astype(np.float64)
# these are the reference values
isf_mp = q[i].astype(np.float64)
# Now we'll use the float64-representable ISF values as input to the SF
# Compute the SF using the reference implementation
sf_mp = sf(isf_mp, a_isf)
# Filter out cases for which the result can't be represented in float64
i = is_float_representable(sf_mp)
# (All should be if start with float64-representable population)
assert np.all(i)
# these are the inputs we'll use for tests of sf
a_sf = a_isf[i]
q_sf = isf_mp[i]
# these are the reference values
sf_mp = sf_mp[i]
### SF Error Histogram ###
# PR SF implementation
def _sf(x, a):
lg = sc.xlog1py(-1.0 / a, a * x**-a)
return -sc.expm1(lg)
# Compute and plot the SF errors
sf_pr = _sf(q_sf, a_sf)
sf_main = stats.kappa3.sf(q_sf, a_sf)
err_pr = log_relative_error(sf_pr, sf_mp)
err_main = log_relative_error(sf_main, sf_mp)
bins = np.arange(-20, 0.5, 0.5)
plt.hist(err_pr, alpha=0.5, label='PR', bins=bins)
plt.hist(err_main, alpha=0.5, label='main', bins=bins)
plt.xlabel('log10 of error')
plt.title('SF method comparison')
plt.legend()
### SF scatter plot ###
x = np.log10(q_sf)
y = np.log10(a_sf)
plt.scatter(x, y, c=err_pr)
cbar = plt.colorbar()
plt.xlabel('q')
plt.ylabel('a')
cbar.set_label('log10 of error')
plt.title('The effect of arguments on kappa.sf error (PR)')
### ISF Error Histogram ###
def _isf(q, a):
lg = sc.xlog1py(-a, -q)
denom = sc.expm1(lg)
return (a / denom)**(1.0 / a)
# Test ISF
isf_pr = _isf(p_isf, a_isf)
isf_main = stats.kappa3.isf(p_isf, a_isf)
err_pr = log_relative_error(isf_pr, isf_mp)
err_main = log_relative_error(isf_main, isf_mp)
bins = np.arange(-20, 0.5, 0.5)
plt.hist(err_pr, alpha=0.5, label='PR', bins=bins)
plt.hist(err_main, alpha=0.5, label='main', bins=bins)
plt.xlabel('log10 of error')
plt.title('ISF method comparison')
plt.legend()
### ISF Scatter Plot
x = np.log10(p_isf)
y = np.log10(a_isf)
plt.scatter(x, y, c=err_pr)
cbar = plt.colorbar()
plt.xlabel('p')
plt.ylabel('a')
cbar.set_label('log10 of error')
plt.title('The effect of arguments on kappa.isf error (PR)')While I support gh-17832 and gh-18093 in principle, I'd suggest that we slow way down. I've reviewed dozens of PRs toward these two issue in the past few months because after bringing the PR count in I've mentioned that I am working on gh-15928 this summer. gh-18829, gh-18811, gh-17719, and gh-18650 are all toward that end, and there are many more on the way. This effort is going to make the default implementations of methods faster and more accurate, and it is going to make it much easier to see where the problems lie (e.g. by generalizing the code above and by supporting the use of |
|
@mdhaber This is a very nice analysis. I also checked the sf plot for a=1e5 and observed that main does better towards the start of the curves. More precisely values ranging from 10**-2 to 1 |
|
@mdhaber I tried adding conditions for the sf but the problem with isf is that for large values of a like 1e4 or 1e5 main and pr are both the same whereas mpmath is different def plot_isf():
a = 1e5
q = np.logspace(-5, 1, 200)
mp_values = np.array([mpmath_sf(_x, a) for _x in q], np.float64)
plt.semilogx(q, kappa3.isf(q, a), label="main", ls="dashed")
plt.semilogx(q, _isf(q, a), label="pr", ls="dashdot")
plt.semilogx(q, mp_values, label="mpmath", ls="dotted")
plt.legend()
plt.title("kappa3 inverse survival function")
plt.show()
So using the main for such values would not be useful. |
|
Right. In So I only added the branching condition in I'll merge when CI comes back green. Please double-check my commit and open a PR if there's something to be fixed. |
Thank you for resolving this. |






Reference issue
Towards: gh-17832
What does this implement/fix?
Additional information