-
Notifications
You must be signed in to change notification settings - Fork 3
How to use plugin
After installation, first run Plugins-> BigTrace X.X.X->Open 3D image command to open an image file.
It is assumed you are going to use a 3D dataset.
The plugin supports 3D composite multicolor images/stacks + timelapse stored as TIFF, BioFormats readable (.lif/.czi etc) or HDF5/XML BDV files.
In addition, there is an experimental support of on-the-fly deskew of Zeiss LLS7 czi files.
Images that do not fit into the computer's memory (RAM) will be "lazy-loaded", i.e. the data shown on the 3D canvas will be read from the disk "on demand". It can be slow and not very responsive, therefore it is recommended to Convert large dataset to BDV format, before opening them in BigTrace.
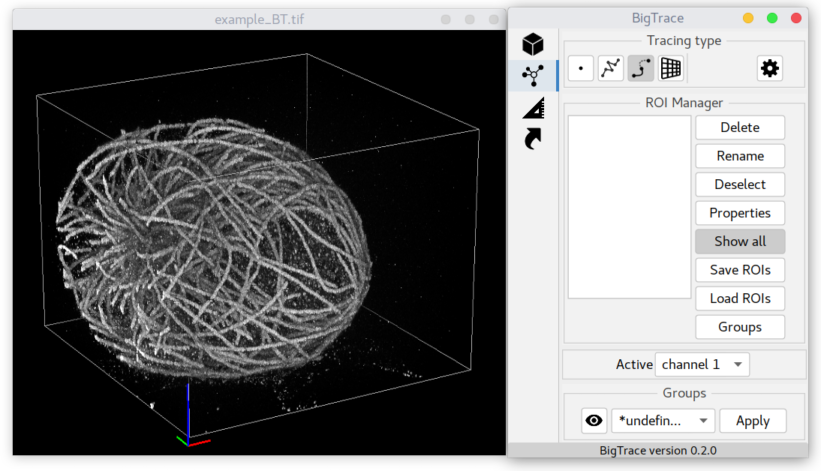
After choosing a file, the plugin should load the dataset and show two windows:
- BigVolumeViewer showing 3D volume rendering of the loaded image and
- a control panel of BigTrace.
At this point, to get started, please refer to:
- Display and navigation page describing dataset browsing;
- Creating ROIs page that covers basics of working with 3D ROIs.
- Panel Measure gives details on different measurements and volume extraction.
There is an excellent video tutorial made by Johanna M. Dela Cruz, see below.
It does not include some recently added features, so please refer to this documentation.
Developed in Cell Biology group of Utrecht University.
Check out Updates history. The plugin and this wiki are under constant development.
E-mail for any questions, feedback, errors or suggestion
or tag @ekatrukha at image.sc forum.