Calculating half mass radius of dark matter halo simulations. This module analyzes data generated by the Rockstar Halo Finder.
Dark Matter Halos are currently characterized by the Navarro-Frenk-White Profile. This profile depends on the change in deinsity gradient which is difficult to calculate precisely for simulations.
This project attempts to recharacterize halos using their half-mass-radius. That is, the ellipsoidal radius containing half the particles in the simlation. The first problem is to fit the halo particles to an ellipsoid so the correct radius can be calculated.
This package requires numpy, matplotlib, and python 2.7.x. Clone this repository. In console browse to the repository and run:
$ python setup.py installOptionally to test code features:
$ python testing.pyWhich will run various functions on test data in data\.
There are several examples present in testing.py. One primary interface is the Halos class. It performs calculations on BGC2 files that contain data for multiple halos.
import halos
H = halos.Halos(PATH_TO_BGC2_FILE) # Single string or list of paths. Wildcards allowed.
H.read_data() # Instantiates a Halo object for each halo in file
H.filter(minimum=10) # Leave out halos with < 10 particles
H.center_halos() # (1) Translate all halos around center points
H.get_covariance_matrices() # (2)
H.get_eigenvectors() # (3)
H.convert_bases() # (4)
H.get_radii() # (5)
H.get_half_mass_radii() # (6) 1-6 are intended to be run in orderThe fundamental object computed on is the Halo. The above class Halos in its function calls simply iterates over Halo objects in Halos().halos. A Halo can either be instantiated from an ascii data file or by passing coordinates:
# Generate halo from coordinates
x = [1,2,3,4,5]
y = [5,4,6,3,1]
z = [3,4,6,7,8]
part_ids = [0,1,2,3,4]
id = 'sample_halo'
center = (0,0,0)
h = halos.helpers.create_halo(id, center, x, y, z, partids)
# Run computations
h.center_halo()
h.get_covariance_matrix()
h.get_eigenvectors()
h.convert_basis()
h.get_radii()
h.get_half_mass_radius()For higher order fitting:
h.higher_order_fit(ORDER) # 2,3,4...Finally, the halo can be visualized with an ellipsoidal fit:
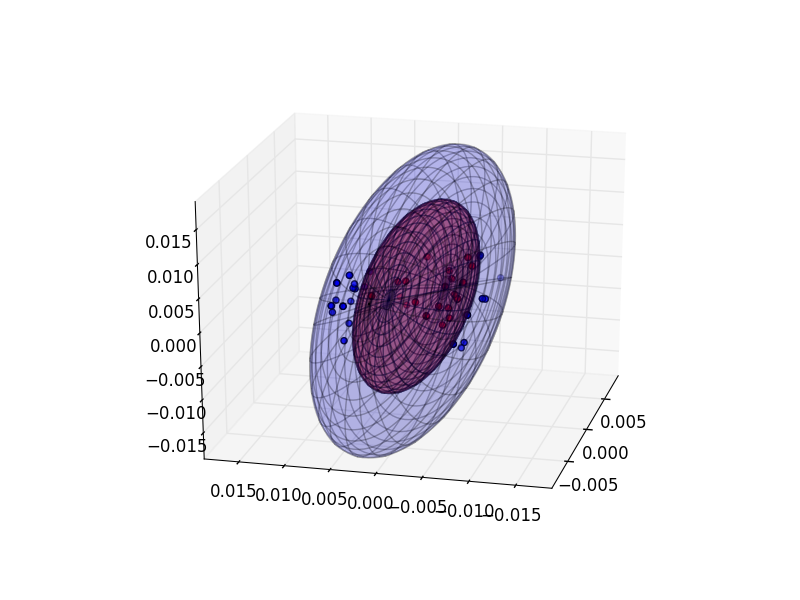
# default mode='cleave', also mode='eval'; default transform=True
h.visualize(ellipsoids=True, mode='cleave', transform=False) cleave uses absolute maximum projections of particles along each principal axis to draw ellipsoids. eval uses halo eigenvalues
to calculate ellipsoid dimensions. transform=True rotates inner halo to represent measured distribution of inner particles only,
otherwise the orientation of the inner halo is assumed to be the same as the rest of the particles.
To get stats about the halo on the console:
h.report()There are several helper functions in halos.gendata module for generating random particle distributions. Please look at source code documentation for more details.
This project is under active development and function behavior may change significantly in the future.
###Multi-core processing
The halos.multicore.Multicore class can be used to achieve data parellelism. Use of the multicore class involves two steps:
- Creating a custom class to execute particular instructions. Create and inherited class and override
parallel_processandpost_processingfunctions.
import halos.multicore as mc
from halos import helpers
class MyMulticore(mc.Multicore):
def parallel_process(self, halo): # each halo in the process passed to this function
_ = helpers.do_all(halo) # add any processing to be done
return halo.half_mass_radius # the result is appended to a list and passed to post_processing()
def post_processing(self, H, results): # H:Halos instance containing all halos in a process
num_halos = len(results)
average_radius = sum(results) / num_halos
return (average_radius, num_halos) # this result is returned to the main process- Using the class interface to start computing
num_processes = 8
m = MyMulticore(num_processes)
m.get_data_from_files(PATH_TO_BGC2_FILES) # supports wildcards. Or use get_data_from_class(Halos)
m.balance_load() # distributes halos across processes per a cost function
m.begin() # spawns processes and begins computing
res = m.get_results() # waits until all processes are finished. Returns result list
# any processing with resultsThe multicore module has additional documentation and features that can be found in the source code.
bgc2.py taken from Daniel Sissom's repository. bgc2.py modified to allow for different file read modes.