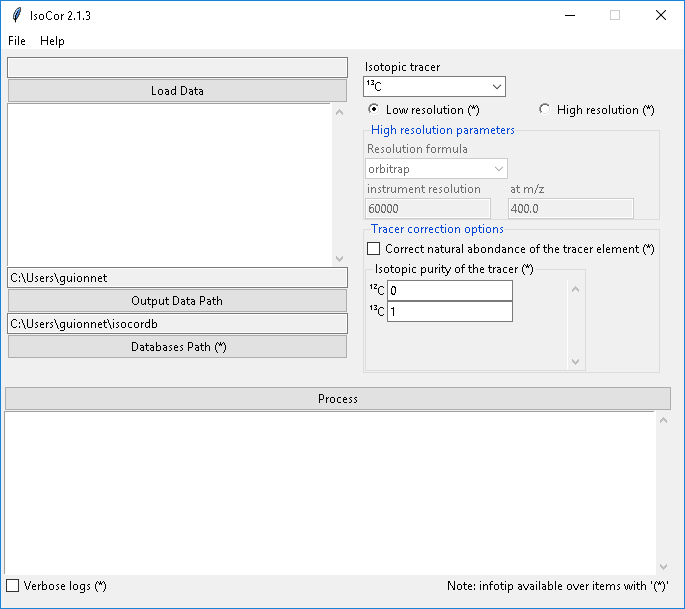
IsoCor is a scientific software dedicated to the correction of mass spectrometry (MS) data for naturally occuring isotopes. IsoCor corrects raw MS data (mass fractions) for naturally-occurring isotopes of all elements and purity of the isotopic tracer. The output of IsoCor is the isotopologue distribution of the molecule (i.e. the relative fractions of molecular entities differing only in the number of isotopic substitutions of the tracer). IsoCor also calculates the mean enrichment (i.e. the mean isotopic content in the molecule) in metabolites.
It is one of the routine tools that we use at the MetaSys team and MetaToul platform in isotopic studies of metabolic systems.
The code is open-source, and available under a GPLv3 license. Additional information can be found in IsoCor publication.
Detailed documentation can be found online at Read the Docs (https://isocor.readthedocs.io/). Check out the Tutorials to use the best correction option!
- correction of naturally occuring isotopes, both for non-tracer and tracer elements,
- correction of tracer purity,
- shipped as a library with both a graphical and command line interface,
- mass-spectrometer and resolution agnostic,
- can be applied to singly- and multiply-charged ions
- can be used with any tracer element (having two or more isotopes)
- account for the contribution of derivatization steps (if any),
- generate isotopic InChIs of tracer isotopologues,
- open-source, free and easy to install everywhere where Python 3 and pip run,
- biologist-friendly.
IsoCor requires Python 3.7 or higher and run on all platforms. Please check the documentation for complete installation and usage instructions.
Use pip to install IsoCor from PyPi:
$ pip install isocorThen, start the graphical interface with:
$ isocorIsoCor is also available directly from command-line and as a Python library.
If you have an idea on how we could improve IsoCor please submit a new issue to our GitHub issue tracker.
Contributions are very welcome! ❤️
Please work on your own fork, follow PEP8 style guide, and make sure you pass all the tests before a pull request.
In development mode, do a pip install -e /path/to/IsoCor to install
locally the development version.
Isotope correction is a complex task and we use unit tests to make sure that critical features are not compromised during development.
You can run all tests by calling pytest in the shell at project's root directory.
Build the HTML documentation with:
$ cd doc
$ make htmlThe PDF documentation can be built locally by replacing html by latexpdf
in the command above. You will need a recent latex installation.
Millard P., Delépine B., Guionnet M., Heuillet M., Bellvert F. and Letisse F. IsoCor: isotope correction for high-resolution MS labeling experiments. Bioinformatics, 2019, doi: 10.1093/bioinformatics/btz209
Baudoin Delépine, Matthieu Guionnet, Pierre Millard
📧 Pierre Millard, millard@insa-toulouse.fr