OME is a consortium of universities, research labs, industry and developers producing open-source software and format standards for microscopy data. If you are working with bioimaging data like these:
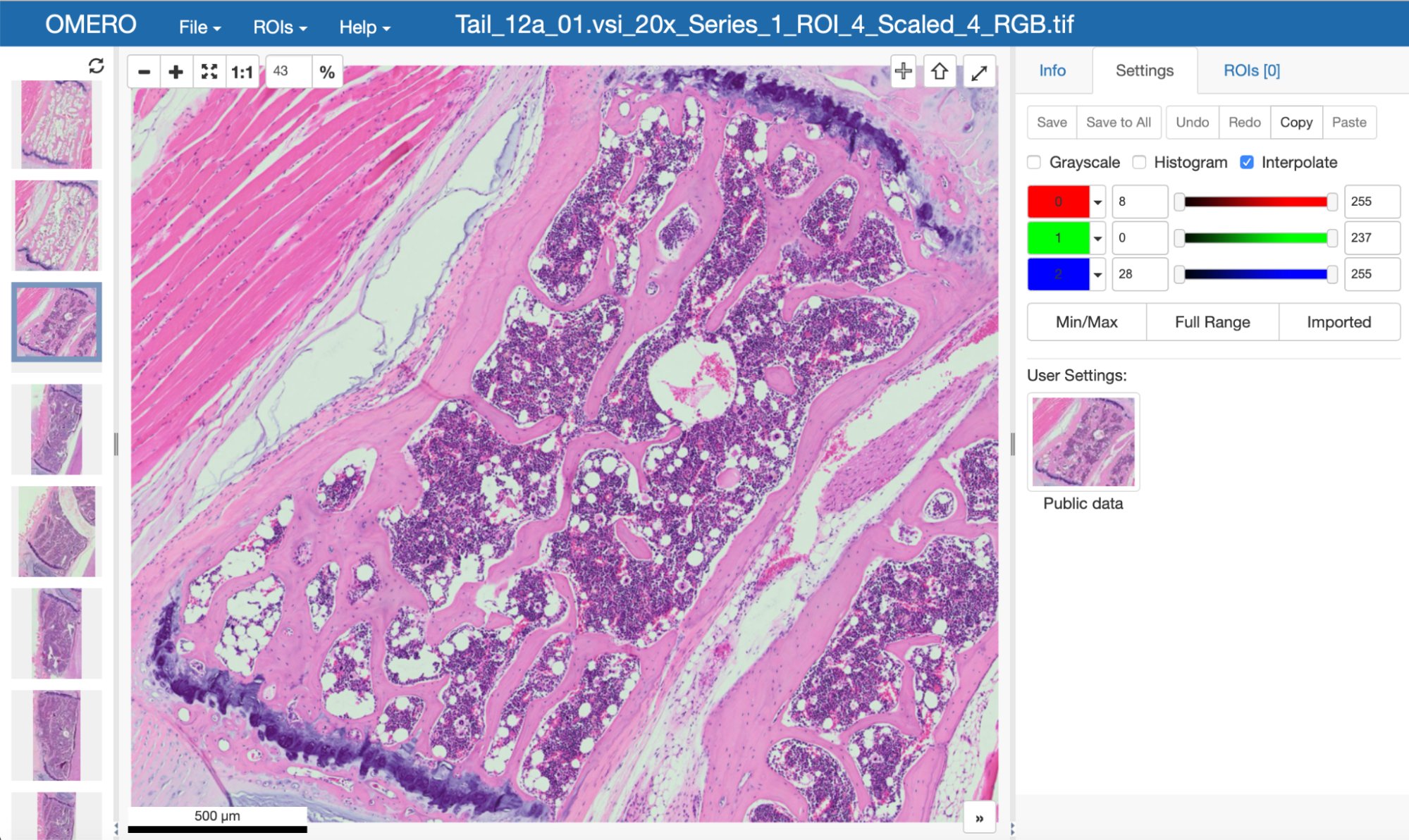 |
 |
 |
then you may already be using something from OME! (You can find out more about these images on https://idr.openmicroscopy.org.)
- OME Model
At the core of everything we do is the specification of an open data model, a vocabulary, that lets software components and users exchange bioimaging (meta)data.
- Bio-Formats
is a library translate hundreds of bioimaging formats into the common model.
- OMERO
is a data management platform that uses Bio-Formats to serve all supported formats to your users, through:
- the OMERO.web
client,
- the OMERO.insight
Java client, or
- the OMERO.cli written using the OMERO.py
API.
- The are also ansible roles and docker images available to help you manage your OMERO server.
- the OMERO.web
- OME-NGFF, next-generation file formats. See:
- ngff
for the specification
- ome-zarr-py
for the Python implementation
- ngff
Find more ways to get started under https://www.openmicroscopy.org/explore!