CIFTools implements reading and writing of CIF files (specification) as well as their efficiently encoded counterpart, called BinaryCIF. The idea is to have a robust, type-safe implementation for the handling of CIF files which does not care about the origin of the data: both conventional text-based and binary files should be handled the same way.
CIFTools is distributed by maven. To get started, append your pom.xml by:
<dependency>
<groupId>org.rcsb</groupId>
<artifactId>ciftools-java</artifactId>
<version>7.0.1</version>
</dependency>Requires Java 11.
class Demo {
public static void main(String[] args) {
String pdbId = "1acj";
boolean parseBinary = true;
// CIF and BinaryCIF are stored in the same data structure
// to access the data, it does not matter where and in which format the data came from
// all relevant IO operations are exposed by the CifIO class
CifFile cifFile;
if (parseBinary) {
// parse binary CIF from RCSB PDB
cifFile = CifIO.readFromURL(new URL("https://models.rcsb.org/" + pdbId + ".bcif"));
} else {
// parse CIF from RCSB PDB
cifFile = CifIO.readFromURL(new URL("https://files.rcsb.org/download/" + pdbId + ".cif"));
}
// fine-grained options are available in the CifOptions class
// access can be generic or using a specified schema - currently supports MMCIF and CIF_CORE
// you can even use a custom dictionary
MmCifFile mmCifFile = cifFile.as(StandardSchemata.MMCIF);
// get first block of CIF
MmCifBlock data = mmCifFile.getFirstBlock();
// get category with name '_atom_site' from first block - access is type-safe, all categories
// are inferred from the CIF schema
AtomSite atomSite = data.getAtomSite();
FloatColumn cartnX = atomSite.getCartnX();
// obtain entry id
String entryId = data.getEntry().getId().get(0);
System.out.println(entryId);
// calculate the average x-coordinate - #values() returns as DoubleStream as defined by the
// schema for column 'Cartn_x'
OptionalDouble averageCartnX = cartnX.values().average();
averageCartnX.ifPresent(System.out::println);
// print the last residue sequence id - this time #values() returns an IntStream
OptionalInt lastLabelSeqId = atomSite.getLabelSeqId().values().max();
lastLabelSeqId.ifPresent(System.out::println);
// print record type - or #values() may be text
Optional<String> groupPdb = data.getAtomSite().getGroupPDB().values().findFirst();
groupPdb.ifPresent(System.out::println);
}
}No difference exists in the API between text-based and binary CIF files. CIF files organize data in blocks, which contain
categories (e.g. AtomSite), which contain columns (e.g. CartnX), which contain values of a particular type (e.g.
double values representing x-coordinates of atoms). The correct names and types for all defined categories and column
from the CIF dictionary are provided.
Just as in Mol* implementation, all parsing and decoding is done as lazily as possible. This makes it cheap to acquire the data structure and hardly wastes any time on preparing information you will never access. In contrast to MMTF, all data can be accessed if needed.
class Demo {
public static void main(String[] args) {
// all builder functionality is exposed by the CifBuilder class
// again access can be generic or following a given schema
MmCifFile cifFile = CifBuilder.enterFile(StandardSchemata.MMCIF)
// create a block
.enterBlock("1EXP")
// create a category with name 'entry'
.enterEntry()
// set value of column 'id'
.enterId()
// to '1EXP'
.add("1EXP")
// leave current column
.leaveColumn()
// and category
.leaveCategory()
// create atom site category
.enterAtomSite()
// and specify some x-coordinates
.enterCartnX()
.add(1.0, -2.4, 4.5)
// values can be unknown or not specified
.markNextUnknown()
.add(-3.14, 5.0)
.leaveColumn()
// after leaving, the builder is in AtomSite again and provides column names
.enterCartnY()
.add(0.0, -1.0, 2.72)
.markNextNotPresent()
.add(42, 100)
.leaveColumn()
// leaving the builder will release the CifFile instance
.leaveCategory()
.leaveBlock()
.leaveFile();
// the created CifFile instance behaves like a parsed file and can be processed or written as needed
System.out.println(new String(CifIO.writeText(cifFile)));
System.out.println(cifFile.getFirstBlock().getEntry().getId().get(0));
cifFile.getFirstBlock()
.getAtomSite()
.getCartnX()
.values()
.forEach(System.out::println);
}
}A step-wise builder is provided for the creation of CifFile instances. If a schema is provided, the builder is aware
of category and column names and the corresponding type described by a column (e.g. the add function called above is
not overloaded, but rather will only accept String values while in entry.id and only double values in
atom_site.Cartn_x.
class Demo {
public static void main(String[] args) {
String id = "AF-Q76EI6-F1-model_v4";
CifFile cifFile = CifIO.readFromURL(new URL("https://alphafold.ebi.ac.uk/files/" + id + ".cif"));
MmCifFile mmCifFile = cifFile.as(StandardSchemata.MMCIF);
// access to properties from the model-extension is provided
// print average per-residue confidence score provided by AlphaFold
System.out.println(mmCifFile.getFirstBlock()
.getMaQaMetricLocal()
.getMetricValue()
.values()
.average()
.orElseThrow());
// convert to BinaryCIF representation
byte[] output = CifIO.writeBinary(mmCifFile);
}
}Computed structure models, e.g. from AlphaFold, are supported. Access to categories and columns defined by the mmCIF model extension is provided. This includes e.g. quality/confidence scores of the prediction. Structure data can be converted to BinaryCIF files for more efficient storage & parsing of millions of files.
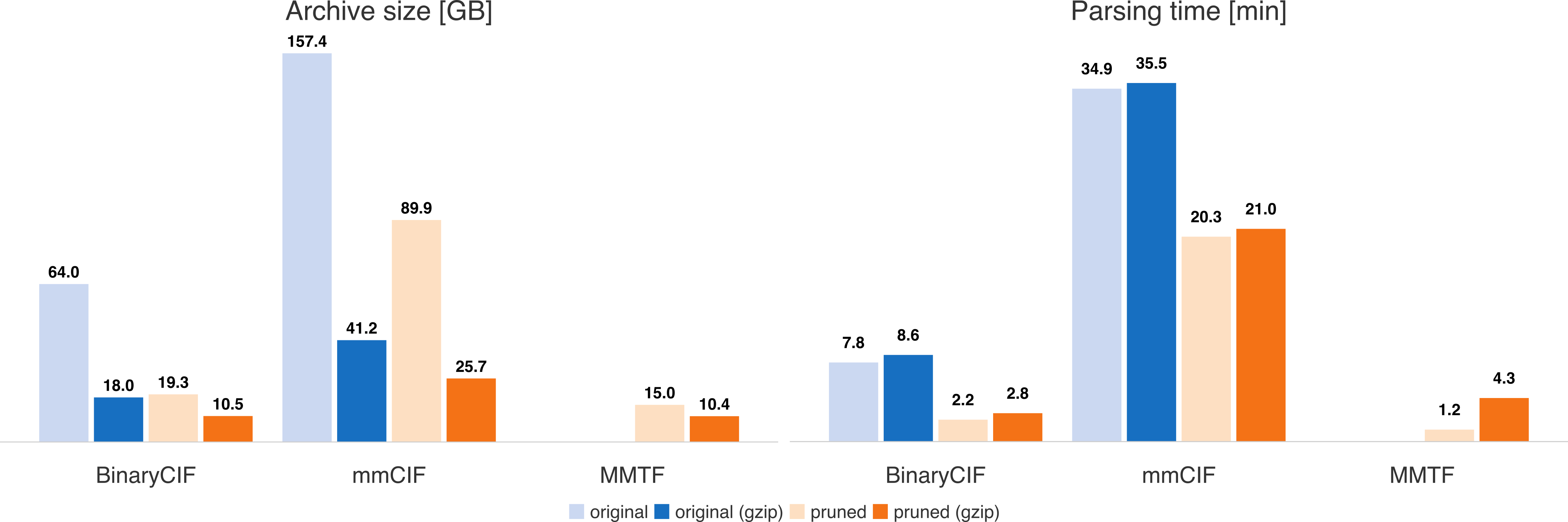
The implementation can read the full PDB archive (154,015 files) in little over 2 minutes. This is achieved by lazy decoding and parsing - all columns are decoded the first time when they are actually requested. Thus, the parsing overhead is kept minimal. Ciftools-java combines the compression and read performance of MMTF and the convenience of the CIF format.
Handling gzipped files slows down parsing in most cases. The reduced files are either native MMTF files or contain a similar selection of CIF categories (i.e. they provide primarily atomic coordinates).
- molstar/ciftools a TypeScript/JavaScript implementation
- molstar/BinaryCIF BinaryCIF format specification
- rcsb/py-mmcif Python mmCIF Core Access Library
The implementation is based on a number of other projects, namely:
- CIFtools.js by David Sehnal
- Mol* by Alexander Rose and David Sehnal
- MMTF by RCSB
- Sehnal D, Bittrich S, Velankar S, Koča J, Svobodová R, Burley SK, Rose AS (2020) BinaryCIF and CIFTools—Lightweight, efficient and extensible macromolecular data management. PLoS Comput Biol 16(10): e1008247. https://doi.org/10.1371/journal.pcbi.1008247