-
Notifications
You must be signed in to change notification settings - Fork 144
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
[Pangenomics] Protein clusters -> gene clusters? #644
Comments
|
Hi Meren,
These are great thoughts! I have a question for clarification--maybe it's
helpful to others...
Is a gene cluster simply a group of homologous sequences (according to set
parameters), which could belong to a single genome or be shared across
multiple genomes? Followup question: if so, does the pangenomic analysis
distinguish between one genome with 25 genes belonging to gene cluster X,
and another genome with only 1 gene belonging to gene cluster X? Or will
these look identical in the display?
…-Luke
On Thu, Nov 16, 2017 at 1:58 PM, A. Murat Eren ***@***.***> wrote:
I propose to change all instances of 'protein clusters' in our pangenomic
workflow with 'gene clusters'.
But briefly my thinking goes like this:
In the pangenomic workflow we take translated DNA sequences from open
reading frames that are often identified by prodigal, and then organize
them in such a way that sequences that show a certain degree of homology
form distinct 'clusters'. We currently call these resulting units as
'protein clusters', and investigate their distribution across contributing
genomes to infer relationships between them.
To me, a more accurate definition of the result of this process is *'clusters
of translated DNA sequences from open reading frames'*, which is not very
helpful. But it is rather straightforward and safe to suggest 'DNA
sequences from open reading frames' represent 'genes', regardless of
whether they ever become 'proteins' or not. 'Gene' is a much more modest
term than 'protein' when one considers the fact that most of the open
reading frames will correspond to genes by definition, but the actual
number of them that ever turn into proteins will be less than that. So I
think 'gene clusters' is a more appropriate term to describe what we are
actually generating.
Clearly gene clusters can be generated from DNA or AA sequences, and the
term 'protein clusters' clarifies that it was the AA sequences that were
used to generate them. But this can be clarified in any methods or results
section:
We used translated DNA sequences of gene calls to identify 'gene clusters'
in our pangenome.
Literature note
The first 'pangenome' paper from Tettelin et al only uses paralog and
homolog clusters to describe gene clusters, and never protein clusters:
https://www.ncbi.nlm.nih.gov/pubmed/16172379
PGAP: pan-genomes analysis pipeline, from Zhao et al., uses gene clusters,
and never protein clusters:
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3268234/
Concerns
The term gene cluster has a very well-defined and established meaning
elsewhere in life sciences:
https://en.wikipedia.org/wiki/Gene_cluster
Can there be a more specific and meaningful term for what we generate in
pangenomes?
(another note: for instance the NCBI use the term 'protein cluster' to
describe "collection of related protein sequences consists of proteins
derived from the annotations of whole genomes, organelles and plasmids
<https://www.ncbi.nlm.nih.gov/proteinclusters>" (the grouping is informed
by the protein function, so it is different than what pangenomic workflows
do).
Any input is most welcome.
Best,
—
You are receiving this because you are subscribed to this thread.
Reply to this email directly, view it on GitHub
<#644>, or mute the thread
<https://github.com/notifications/unsubscribe-auth/AY6jPNspcmKOG7zIIYtDTasas2d5jA5sks5s3KHbgaJpZM4QhIL7>
.
--
Dr. Luke McKay
Postdoctoral Research Scientist
Department of Land Resources and Environmental Sciences (815 LJH)
Center for Biofilm Engineering (313 Barnard Hall)
Montana State University
|
|
Hi Meren et al., Here are some points that I hope you will find helpful:
In any case, it would be good to provide the user with paralog information and how you dealt with it. Hervé. |
|
Hi Meren, How would the term "orthogroup" fit into this conversation? I like it: https://genomebiology.biomedcentral.com/articles/10.1186/s13059-015-0721-2 Best, |
|
Dear Luke, Thank you very much for the input.
Yes. A group of homologous sequences based on sequence identity, and/or network decomposition parameters for MCL-like algorithms. These clusters could have been generated by tools such as MMseqs2, too.
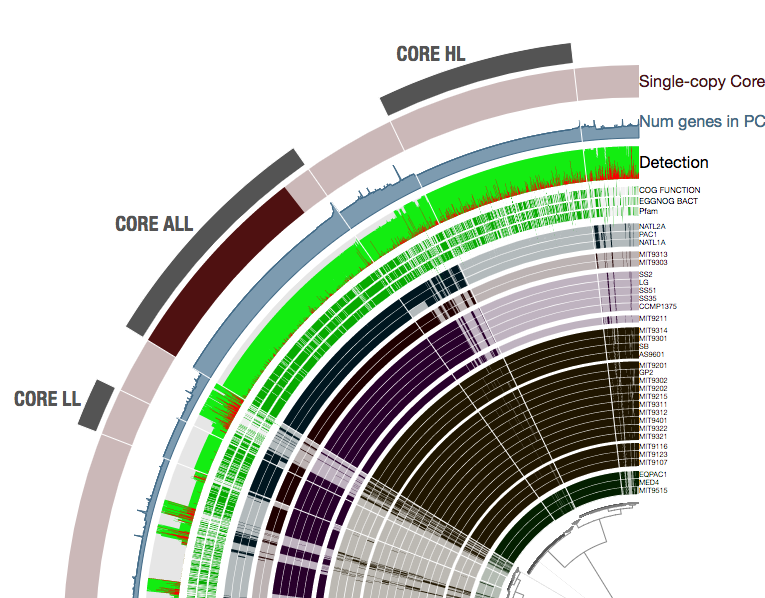
Yes, good point. We do track paralogs. At a given time we know how many genes from each genome is contributed to a given gene cluster. This allows us to identify and mark the single-copy gene clusters, and show how many genes in each gene cluster: And show paralogs when the user wants to inspect a given gene cluster: I hope these make sense. |
|
Hi Hervé, Thanks for taking the time :)
Yes, these are based on sequence similarity with defined parameters. It will also be useful to extend our efforts to use functional information, but currently we haven't done anything in that front.
Yes. Clarifying the terminology the first time we mention gene clusters will be essential to avoid any misunderstandings.
I see your point. In some cases we do get clusters with very large number of genes in them simply because they all have a conserved region in otherwise dissimilar sequences. There are many things that can be done to divide them into synthetic orthologs. Trimming the conserved region from the alignment and re-clustering the remaining sequences can be one of them to re-organize those paralogs better.
We do our best to not make this information hard to find, but what we are doing in the interface could certainly be improved. Your comment made me realize that we can add another layer to show 'extent of paralogy' in each cluster (i.e., the maximum number of genes coming from the same genome for every cluster). I entered an issue, and we will implement that. I guess no one so far has any real problems with "gene clusters" as far as how they are generated is clearly described. |
|
Hi Meren et al., |
|
Hello, the term protein clusters is not appropriate as you mentioned. Also, gene clusters is the term overlapping the other scientific domains. So, like our pan genome pipeline which refers to these as GENE FAMILIES would be appropriate as per my belief. |
I think orthology explains only a subset of the clusters we are generating, and ignores paralogy, hence it is not maybe the best option. To quote people who have thought about these a lot, Gabaldón & Koonin has a nice paper that starts like this: "Orthologues and paralogues are types of homologous genes that are related by speciation or duplication, respectively". So orthogroup does not explain what our gene clusters contain. Homology does, for instance, but "homologous groups" would have been even less helpful since the term "group" or "cluster" already implies homology.
You make valid points, and I especially tend to agree when you suggest most of these predicted genes are translated to proteins, so why bother. But I think there is a large consensus that 'protein clusters' is not working well. Maybe 'gene clusters' is not working well either, but it seems that is a step towards a bit of a better direction. @9aren,
I'm afraid 'gene families' is not any better than 'gene clusters' with respect to existing literature. Gene families are also well defined elsewhere in life sciences, and mean "a set of several similar genes, formed by duplication of a single original gene, and generally with similar biochemical functions [within a single organism]". Additionally, 'clusters' is a more neutral term represent what they really are in a more direct way than 'families' in my opinion. So if both 'gene clusters' and 'gene families' is equally contaminated with previous literature, I think 'gene clusters' is a simpler term. Thank you for all the input. |
|
I think the big problem is the fact that even when we say 'gene' we make assumptions about the nature of these predicted sequences and are given back to us by a software that solely operates on sequencing data. The "cluster" part is OK. The rest is problematic, in my opinion. "Gene clusters" simplifies this to a degree while minimizing the amount of assumptions we make. But for instance, Prodigal often identifies two putative genes within the 16S rRNA gene (so it doesn't identify 16S rRNA gene as a gene, but often makes two short gene calls from within that gene when I carefully inspect results). Maybe 'predicted gene clusters' is much more appropriate, and will fit in most cases, but I am not sure if it would get enough support. I guess in our small world in my group, we will continue Gene Clusters if no one else suggests anything else. PS: Thanks, science, for this Saturday morning depression :/ |
True, but if we want to solve to issue of using an established term (i.e. "gene clusters"), then "clusters of homologous genes" is, in my opinion, an appropriate description. Even though, as you say, cluster implies homology, but the term homology is not used in a general manner here; homologous genes are well-defined in our field, and we use NCBI-BLAST (or diamond or a different tool) to try to identify homology, and then use MCL (or other algorithms) to cluster them into clusters of homologous genes. |
|
@meren I agree, "protein cluster" seems to be an issue for a number of folks (way more than I expected), and the issues raised are all valid. If we are looking for a denomination with the least assumption possible, then "ORF cluster" might be it, although it doesn't cover 16S, and it's kind of ugly. Everything else can be "attacked", especially calling predicted ORFs "proteins" or "genes", or calling automatically defined clusters of predicted ORFs "homologs". |
|
Hi, Could someone tell me why "sequence cluster" is a bad option? A quick google search brings me to this:
"Sequence cluster" is a very neutral terminology, which might be a good thing for the matter at hand. |
|
I think the most important thing here is for the community to find a term that we can agree is the least problematic and stick to it-- kind of like the way we circled around the terms "bin" and "population genome" and settled on "metagenome assembled genome" (MAG). I am wary of anything using the word "family" because this has a very specific meaning in the literature (i.e. see Pfam), and I agree that anything using the word "protein" makes the assumption that the sequence in question will be translated into a protein. Too many words (i.e. "clusters of homologous genes") gets unwieldy. "Gene cluster" works, and I like Tom's "sequence cluster" suggestion. |
|
I'm comfortable with either "gene clusters" or "protein clusters" as long as it's defined. I also would vote for "homologous gene/protein clusters" or similar (i.e., HPCs, or HGCs). My only problem with "sequence cluster" is that it doesn't indicate the unit being clustered, so it's a less intuitive term. But I agree with rikander that the main thing is to just find a term that's not already used, define it, and stick with it. |
It reminds me of OTUs, and makes me think that maybe we don't need a yet-another-acronym-where-every-letter-brings-its-own-baggage. Also, the study (in which this term is defined) is not related to pangenomes since these units the authors generated from open reading frames are not associated with any genomic context. The study proposes that OPFs are analogous to 16S OTUs to make comparisons among communities.. Historically important study, but I think the term it defined does not have any relevance here. |
I really like this and I will tell you why :) We are using homology (currently it is at the sequence-level, but in the future it can be at the functional-level, or a combination of both as people tried before). We are doing it things that are very close to genes (because the information often comes from gene callers, etc). And we group them into clusters. All checks out. And your suggestion addresses a subtle point: we are truly clustering homologous genes. Here is a very basic example to elaborate: Genes Now I will tell you why I don't like it :/ While it sets a very accurate level of specificity, it doesn't remove the need to find something better to replace the word 'gene', and it doesn't add something to bring the predicted nature of this information into the mix. So the additional value to go from 'gene clusters' to 'clusters of homologous genes' is not justified in the expense of adding two more words (as @rikander mentioned above). Why does this matter? Because if we call these 'gene clusters' (and define them clearly every time we first mention them), we don't have to use an acronym. We can say gene clusters every time (I've tried it while writing this paper, and it reads rather well). If we call them "clusters of homologous genes", then we can't spell it out every time (because it is 4 words and will impact the flow too often), and we will have to use an acronym, such as CHGs. I think we are all tired of acronyms (because, you know, we are not immunologists), and in my opinion it would be great if we end up don't introduce a new acronym. My 1½ cents. |
|
Well, I think this is a very interesting discussion and that there might not be a single consensus solution in there. why? because science. Which suggests that different 'term' would work if it comes with a clear definition along (like @rbeinart 'I'm comfortable with either "gene clusters" or "protein clusters" as long as it's defined'). Including that these are 'predicted genes' and might contain errors, ORF translated into AA and that homologs are grouped together into clusters based on sequence comparison (not function here). I vote for a detailed blog post instead of a new acronym. |
|
Thanks for the input, @jreveillaud. I will do my best to summarize all these points in a blog post for future references. While I agree that it is OK to not have a single consensus, sadly, at least in the context of anvi'o, we have to pick a single term due to practical reasons (which will impact interfaces, tutorials, and the code behind). I am aware that the discussion we are having (and its result) will have no binding power outside of the context of the anvi'o codebase. But we (the anvi'o developers) would still be much more comfortable if we present people (who wish to do pangenomic analyses using this tool) with terms with which the community does not disagree :) |
protein clusters -> gene clusters. closes #644.

I propose to change all instances of 'protein clusters' in our pangenomic workflow with 'gene clusters'.
Summary of the discussions so far
I will keep this section up-to-date, the original issue starts at the "Why bother" section below.
Although there is a large diversity of opinions, it doesn't seem anyone has a major concern with "gene clusters" yet (as far as how they are generated is clearly described):
I feel that our major problem was the need for a term that communicates the fact that we are working with sequences that are coming from predicted open reading frames. Coding sequence, protein, gene, family, orthology, paralogy, all bring in assumptions that do no apply to what goes in our clusters in a pangenome.
Why bother?
In the pangenomic workflow we take translated DNA sequences from open reading frames that are often identified by a gene prediction software, and then organize them in such a way that sequences that show a certain degree of homology form distinct 'clusters'. We currently call these resulting units as 'protein clusters', and investigate their distribution across contributing genomes to infer relationships between them.
To me, a more accurate definition of the result of this process is 'clusters of translated DNA sequences from predicted open reading frames', which is not very helpful. But it is rather straightforward and safe to suggest 'DNA sequences from open reading frames' represent 'genes', regardless of whether they ever become 'proteins' or not. 'Gene' is a much more modest term than 'protein' when one considers the fact that most of the open reading frames will correspond to genes by definition, but the actual number of them that ever turn into proteins will be less than that. So I think 'gene clusters' is a more appropriate term to describe what we are actually generating.
Clearly gene clusters can be generated from DNA or AA sequences, and the term 'protein clusters' clarifies that it was the AA sequences that were used to generate them. But this can be clarified in any methods or results section:
Literature note
The first 'pangenome' paper from Tettelin et al. only uses 'paralog clusters' and 'homolog clusters' to describe gene clusters, and never protein clusters:
https://www.ncbi.nlm.nih.gov/pubmed/16172379
PGAP: pan-genomes analysis pipeline, from Zhao et al., also uses gene clusters, and never protein clusters:
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3268234/
A Concern
The term 'gene cluster' has a very well-defined and established meaning elsewhere in life sciences:
https://en.wikipedia.org/wiki/Gene_cluster
Can there be a more specific and meaningful term for what we generate in pangenomes?
Or should we not care since the context will make it obvious the gene clusters in pangenomes have nothing to with gene clusters in eukaryotic genomes like Hox genes?
For instance the NCBI uses the term 'protein cluster' to describe "collection of related protein sequences consists of proteins derived from the annotations of whole genomes, organelles and plasmids" (the grouping is informed by the protein function, so it is different than what pangenomic workflows do).
Any input is most welcome.
Best,
The text was updated successfully, but these errors were encountered: